-Search query
-Search result
Showing 1 - 50 of 1,691 items for (author: sheng & d)
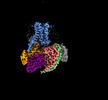
EMDB-60628: 
Carazolol-activated human beta3 adrenergic receptor
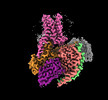
EMDB-60629: 
Epinephrine-activated human beta3 adrenergic receptor

EMDB-18661: 
Structure of the Bacteriophage PhiKZ non-virion RNA Polymerase bound to DNA and RNA

PDB-8que: 
Structure of the Bacteriophage PhiKZ non-virion RNA Polymerase bound to DNA and RNA

EMDB-36730: 
SARS-CoV-2 Spike RBD (dimer) in complex with two 2S-1244 nanobodies

EMDB-36735: 
Dimer of SARS-CoV-2 BA.2 spike and IBT-CoV144(C3 symmetry)

EMDB-36740: 
Dimer of SARS-CoV-2 BA.2 spike and IBT-CoV144(C1 symmetry)

PDB-8jys: 
SARS-CoV-2 Spike RBD (dimer) in complex with two 2S-1244 nanobodies

EMDB-38156: 
Structure of enterovirus protease in complex host factor

PDB-8x8q: 
Structure of enterovirus protease in complex host factor

EMDB-17356: 
Structure of divisome complex FtsWIQLB

PDB-8p1u: 
Structure of divisome complex FtsWIQLB

EMDB-37663: 
CryoEM structure of ZIKV rsNS1

EMDB-37670: 
ZIKV rsNS1 in complex with Fab AA12 and anti-fab nanobody

EMDB-37673: 
ZIKV rsNS1 in complex with Fab GB5 and anti-fab nanobody

EMDB-37676: 
CryoEM structure of ZIKV rsNS1 filament

EMDB-37678: 
ZIKV rsNS1 in complex with Fab EB9 and anti-fab nanobody

EMDB-43844: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP45 Class B

EMDB-43845: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP45 Class C1

EMDB-43850: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63 Class C

EMDB-43852: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63 Class E1

EMDB-17766: 
CryoEM structure of Nal1 protein, allele SPIKE, from Oryza sativa japonica group
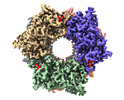
EMDB-17768: 
CryoEM structure of Nal1 protein, allele IR64, from Oryza sativa indica cultivar

PDB-8pn1: 
CryoEM structure of Nal1 protein, allele SPIKE, from Oryza sativa japonica group

PDB-8pn2: 
CryoEM structure of Nal1 protein, allele IR64, from Oryza sativa indica cultivar
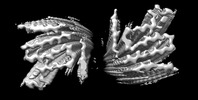
EMDB-36202: 
Cryo-EM structure of alpha-synuclein gS87 fibril

EMDB-36203: 
Cryo-EM structure of alpha-synuclein pS87 fibril

PDB-8jex: 
Cryo-EM structure of alpha-synuclein gS87 fibril

PDB-8jey: 
Cryo-EM structure of alpha-synuclein pS87 fibril
EMDB-37240: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab

EMDB-37241: 
The interface structure of Omicron RBD binding to 5817 Fab

PDB-8khc: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab

PDB-8khd: 
The interface structure of Omicron RBD binding to 5817 Fab

EMDB-43889: 
Chlamydomonas reinhardtii mastigoneme (constituent map 1)

EMDB-43890: 
Chlamydomonas reinhardtii mastigoneme (constituent map 2)

EMDB-43891: 
Chlamydomonas reinhardtii mastigoneme (constituent map 3)

EMDB-43892: 
Composite cryo-EM map of the Chlamydomonas reinhardtii mastigoneme

PDB-9b4h: 
Chlamydomonas reinhardtii mastigoneme filament

EMDB-18659: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT1

EMDB-18660: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT5

PDB-8quc: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT1

PDB-8qud: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT5

EMDB-36005: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound MK-6892

EMDB-36007: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound niacin

PDB-8j6i: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound MK-6892

PDB-8j6l: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound niacin

EMDB-37626: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein in complex with hippopotamus ACE2

EMDB-37629: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein receptor-binding domain in complex with hippopotamus ACE2

PDB-8wlo: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein in complex with hippopotamus ACE2

PDB-8wlr: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein receptor-binding domain in complex with hippopotamus ACE2
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
