-Search query
-Search result
Showing all 34 items for (author: sedat & j)

EMDB-15057: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-15058: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-15059: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-15060: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-15061: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-15062: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-15063: 
Fibroblast nucleus sub-volume.
Method: electron tomography / : Sedat J, McDonald A, Elbaum M

EMDB-14924: 
Cryo-STEM tomography of fibroblast nuclear periphery
Method: electron tomography / : Wolf SG, Fass D, Elbaum M

EMDB-24416: 
Microtubule subtomogram average, randomized azimuth and restricted angular search
Method: subtomogram averaging / : Croxford M, Villa E

EMDB-24417: 
Microtubule subtomogram average from particles with a uniform starting orientation with respect to the missing wedge
Method: subtomogram averaging / : Croxford M, Villa E

EMDB-24418: 
Microtubule subtomogram average of deconvolved particles, randomized starting azimuth and restricted angular search
Method: subtomogram averaging / : Croxford M, Villa E

EMDB-24419: 
Microtubule subtomogram average of deconvolved rarticles, randomized starting azimuth relative to the missing wedge, and restricted angular search
Method: subtomogram averaging / : Croxford M, Villa E

EMDB-24433: 
Yeast Nucleus Tomogram
Method: electron tomography / : Croxford M, Villa E

EMDB-24434: 
Deconvolved Yeast Nuclear Tomogram
Method: electron tomography / : Croxford M, Villa E

EMDB-24435: 
Tomogram of HEK292 Inculsion Body
Method: electron tomography / : Croxford M, Villa E

EMDB-24436: 
Deconvolved Tomogram of HEK293 Inclusion Body
Method: electron tomography / : Croxford M, Villa E

PDB-7l6o: 
Cryo-EM structure of HIV-1 Env CH848.3.D0949.10.17chim.6R.DS.SOSIP.664
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23518: 
Cryo-EM map of DH851.3 bound to HIV-1 CH505 Env
Method: single particle / : Edwards RJ, Manne K, Acharya P

PDB-7lu9: 
Cryo-EM structure of DH851.3 bound to HIV-1 CH505 Env
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23519: 
Cryo-EM map of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Edwards RJ, Manne K, Acharya P

PDB-7lua: 
Cryo-EM structure of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23152: 
Cryo-electron microscopy reconstruction of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer
Method: single particle / : Edwards RJ, Acharya P

EMDB-23153: 
Cryo-electron microscopy local refinement of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer
Method: single particle / : Edwards RJ, Acharya P

EMDB-23149: 
Cryo-electron microscopy reconstruction of antibody DH898.1 Fab-dimer bound near the CD4 binding site of HIV Env SOSIP trimer CH848 10.17
Method: single particle / : Edwards RJ, Acharya P

EMDB-23124: 
Cryo-electron microcospy reconstruction of CH848.3.D0949.10.17chim.6R.DS.SOSIP.664 HIV Env
Method: single particle / : Edwards RJ, Acharya P

EMDB-23145: 
Cryo-electron microscopy reconstruction of locally refined antibody DH898.1 Fab-dimer
Method: single particle / : Edwards RJ, Acharya P

PDB-7l6m: 
Cryo-EM structure of DH898.1 Fab-dimer from local refinement of the Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23094: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R

EMDB-23095: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R

EMDB-23097: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement
Method: single particle / : Manne K, Henderson R

PDB-7l02: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l06: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l09: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement
Method: single particle / : Manne K, Henderson R, Acharya P
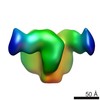
EMDB-8507: 
92BR SOSIP.664 trimer in complex with DH270.1 Fab
Method: single particle / : Fera D, Harrison SC
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
