-Search query
-Search result
Showing 1 - 50 of 729 items for (author: oh & bh)

EMDB-28966: 
CryoEM map of de novo designed oligomeric protein C4-71_6x

EMDB-28967: 
CryoEM map of de novo designed oligomeric protein C4-71_8x
EMDB-28968: 
CryoEM map of de novo designed oligomeric protein C6-71

EMDB-28969: 
CryoEM map of de novo designed oligomeric protein C6-71_6x

EMDB-28970: 
CryoEM map of de novo designed oligomeric protein C6-71_8x

EMDB-28971: 
CryoEM map of de novo designed oligomeric protein C8-71_6x

EMDB-28972: 
CryoEM map of de novo designed oligomeric protein C8-71_8x
EMDB-28973: 
CryoEM map of de novo designed oligomeric protein C4-81

EMDB-28974: 
CryoEM map of designed oligomeric protein C4-71

EMDB-17197: 
Human TPC2 in Complex with Antagonist (S)-SG-094

EMDB-19108: 
Human TPC2 in Complex withAntagonist (R)-SG-094

PDB-8ouo: 
Human TPC2 in Complex with Antagonist (S)-SG-094
EMDB-41434: 
Bottom cylinder of high-resolution phycobilisome quenched by OCP (local refinement)
EMDB-41435: 
Central rod disk in C1 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
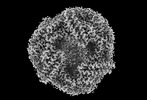
EMDB-41436: 
Central rod disk in D3 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
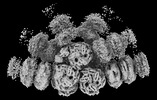
EMDB-41463: 
Synechocystis PCC 6803 Phycobilisome quenched by OCP, high resolution

EMDB-41475: 
Top cylinder bound to OCP from high-resolution phycobilisome quenched by OCP (local refinement)
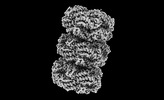
EMDB-41585: 
Rod from high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8to2: 
Bottom cylinder of high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8to5: 
Central rod disk in C1 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8tpj: 
Top cylinder bound to OCP from high-resolution phycobilisome quenched by OCP (local refinement)

PDB-8tro: 
Rod from high-resolution phycobilisome quenched by OCP (local refinement)

EMDB-40208: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15

EMDB-40209: 
Chlorophyll-binding region of de novo-designed nanocage O32-15

PDB-8glt: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
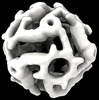
EMDB-42031: 
Computational Designed Nanocage O43_129_+8

EMDB-43318: 
Twistless helix 12 repeat ring design R12B

EMDB-17509: 
Cryo-EM structure of CAK in complex with inhibitor BS-181

EMDB-17510: 
Cryo-EM structure of CAK in complex with inhibitor BS-194

EMDB-17512: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0510-R

EMDB-17513: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0510-S

EMDB-17514: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0574

EMDB-17515: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0768

EMDB-17516: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0829

EMDB-17517: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0880 (ring-up conformation)

EMDB-17518: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0880 (ring-down conformation)

EMDB-17519: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0914

EMDB-17520: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0943

EMDB-17521: 
Cryo-EM structure of CAK in complex with inhibitor dinaciclib

EMDB-17522: 
Cryo-EM structure of CAK with averaged inhibitor density

EMDB-17523: 
Cryo-EM structure of apo CAK

EMDB-17524: 
Cryo-EM map of inhibitor-bound CAK (Krios G4 performance comparison dataset)

EMDB-17525: 
Cryo-EM map of inhibitor-bound CAK (Glacios G2 performance comparison dataset)

EMDB-17526: 
Cryo-EM map of inhibitor-bound CAK (+EF performance comparison dataset)

EMDB-17527: 
Cryo-EM map of inhibitor-bound CAK (-EF performance comparison dataset)

EMDB-17536: 
Cryo-EM structure of CDK7 subunit of CAK in complex with inhibitor LDC4297

EMDB-17754: 
Cryo-EM structure of CAK in complex with inhibitor CT7030

PDB-8p6w: 
Cryo-EM structure of CAK in complex with inhibitor BS-181

PDB-8p6x: 
Cryo-EM structure of CAK in complex with inhibitor BS-194
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
