-Search query
-Search result
Showing 1 - 50 of 226 items for (author: nguyen & ah)

EMDB-18658: 
Structure of the NCOA4 (Nuclear Receptor Coactivator 4)-FTH1 (H-Ferritin) complex

PDB-8qu9: 
Structure of the NCOA4 (Nuclear Receptor Coactivator 4)-FTH1 (H-Ferritin) complex

EMDB-18334: 
Cryo-EM structure of the inward-facing FLVCR1

EMDB-18335: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

EMDB-18336: 
Cryo-EM structure of the inward-facing FLVCR2
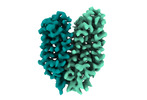
EMDB-18337: 
Cryo-EM structure of the outward-facing FLVCR2

EMDB-18339: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

EMDB-19009: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

PDB-8qcs: 
Cryo-EM structure of the inward-facing FLVCR1

PDB-8qct: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

PDB-8qcx: 
Cryo-EM structure of the inward-facing FLVCR2

PDB-8qcy: 
Cryo-EM structure of the outward-facing FLVCR2

PDB-8qd0: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

PDB-8r8t: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

EMDB-29907: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v); consensus map with only Fab 1G01 resolved

EMDB-29908: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v), locally refined map

EMDB-29909: 
Structure of human NDS.3 Fab in complex with influenza virus neuraminidase from A/Darwin/09/2021 (H3N2)

PDB-8gat: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v), based on consensus cryo-EM map with only Fab 1G01 resolved

PDB-8gau: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v)

PDB-8gav: 
Structure of human NDS.3 Fab in complex with influenza virus neuraminidase from A/Darwin/09/2021 (H3N2)
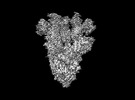
EMDB-28198: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with LLNL-199

EMDB-28199: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112

PDB-8ekd: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112

EMDB-41171: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology

EMDB-41172: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
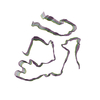
PDB-8tdn: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
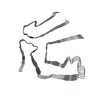
PDB-8tdo: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology

EMDB-36159: 
Cryo-EM Structure of Chikungunya Virus Nonstructural Protein 1 with m7GpppAmU

PDB-8jce: 
Cryo-EM Structure of Chikungunya Virus Nonstructural Protein 1 with m7GpppAmU

EMDB-29980: 
Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils

PDB-8gf7: 
Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils

EMDB-29783: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074

EMDB-41613: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074

PDB-8g6u: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074

PDB-8ttw: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074

EMDB-27031: 
Accurate computational design of genetically encoded 3D protein crystals

EMDB-40926: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4

PDB-8cwy: 
Accurate computational design of genetically encoded 3D protein crystals

PDB-8szz: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4

EMDB-41994: 
KCTD5/Cullin3/Gbeta1gamma2 Complex: Local Refinement of KCTD5(CTD)/Gbeta1gamma2

EMDB-41995: 
KCTD5/Cullin3/Gbeta1gamma2 Complex: Local Refinement of KCTD5(BTB)/Cullin3(NTD)

EMDB-41996: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A: Composite Map from RELION Multi-body Refinement

EMDB-41997: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A: Reference Map for Composite Map EMD-41996

EMDB-41998: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A Body 1: RELION Multi-body Refinement of KCTD5(CTD)/Gbeta1gamma2

EMDB-41999: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A Body 2: RELION Multi-body Refinemnent of KCTD5(BTB)/Cullin3(NTD)

EMDB-42000: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B: Composite Map from RELION Multi-body Refinement

EMDB-42001: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B: Reference Map for Composite Map EMD-42000

EMDB-42002: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B Body 1: RELION Multi-body Refinement of KCTD5(CTD)/Gbeta1gamma2

EMDB-42003: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B Body 2: RELION Multi-body Refinement of KCTD5(BTB)/Cullin3(NTD)

EMDB-42004: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State C: Composite Map from RELION Multi-body Refinement
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
