-Search query
-Search result
Showing 1 - 50 of 184 items for (author: nans & a)

EMDB-56951: 
Cryo-tomogram of COPI and COPII vesicles and buds from FIB-milled RPE-1 cells.
Method: electron tomography / : Downes KW, Zanetti G, Nans A

EMDB-56963: 
Ribosome subtomogram average
Method: subtomogram averaging / : Downes KW, Nans A, Zanetti G

EMDB-56606: 
Cryo-tomogram of COPI and COPII vesicles and buds from FIB-milled RPE-1 cells.
Method: electron tomography / : Downes KW, Zanetti G, Nans A

EMDB-54068: 
SIVtal integrase in complex with RNA stem-loop (focused refinement of the filament repeat unit)
Method: single particle / : Singer MR, Cherepanov P

EMDB-54071: 
CryoEM reconstruction of integrase filament at the lumen of native HIV-1 cores (box size 34.2 nm)
Method: single particle / : Cherepanov P, Singer MR, Hope J, Zhang P

PDB-9rmu: 
SIVtal integrase in complex with RNA stem-loop (focused refinement of the filament repeat unit)
Method: single particle / : Singer MR, Cherepanov P

PDB-9rmx: 
CryoEM reconstruction of integrase filament at the lumen of native HIV-1 cores (box size 34.2 nm)
Method: single particle / : Cherepanov P, Singer MR, Hope J, Zhang P

EMDB-54070: 
CryoEM reconstruction of integrase filament at the lumen of native HIV-1 cores (box size 47.3 nm)
Method: single particle / : Cherepanov P, Singer MR, Hope J, Zhang P

EMDB-55409: 
HIV-1 integrase filament at the luminal side of capsid lattice by subtomogram averaging.
Method: subtomogram averaging / : Cherepanov P, Chenavier F, Hope J, Nans A, Zhang P

EMDB-51611: 
Structure of FLuc-XBP1u+ stalled human 60S ribosome nascent chain complex
Method: single particle / : Voisin TB, Pellowe GA, Balchin D

PDB-9gul: 
Structure of FLuc-XBP1u+ stalled human 60S ribosome nascent chain complex
Method: single particle / : Voisin TB, Pellowe GA, Balchin D

EMDB-50092: 
HIV-1 envelope glycoprotein (BG505 gp140 SOSIP.664) trimer in complex with three copies of ELC07 broadly neutralizing antibody.
Method: single particle / : Hope J, Alguel Y, Nans A, Cherepanov P

PDB-9f02: 
HIV-1 envelope glycoprotein (BG505 gp140 SOSIP.664) trimer in complex with three copies of ELC07 broadly neutralizing antibody.
Method: single particle / : Hope J, Alguel Y, Nans A, Cherepanov P

EMDB-50020: 
HIV-1 envelope glycoprotein (BG505 gp140 SOSIP.664) trimer in complex with ELC07 broadly neutralizing antibody.
Method: single particle / : Hope J, Alguel Y, Nans A, Cherepanov P

PDB-9evz: 
HIV-1 envelope glycoprotein (BG505 gp140 SOSIP.664) trimer in complex with ELC07 broadly neutralizing antibody.
Method: single particle / : Hope J, Alguel Y, Nans A, Cherepanov P

PDB-8s78: 
MicroED Structure of TLR2 TIR domain-induced MyD88 TIR domain higher-order assembly
Method: electron crystallography / : Li Y, Pacoste L, Xu H, Kobe B

EMDB-46946: 
TRIF TIR Filament Cryo-EM Structure
Method: helical / : Manik MK, Xiao L, Wu H

EMDB-46977: 
CryoEM structure of the TIR domain from human TRAM
Method: helical / : Hedger A, Pospich S, Pan M, Gu W, Ve T, Raunser S, Landsberg M, Nanson JD, Kobe B

PDB-9dlg: 
CryoEM structure of the TIR domain from human TRAM
Method: helical / : Hedger A, Pospich S, Pan M, Gu W, Ve T, Raunser S, Landsberg M, Nanson JD, Kobe B

PDB-9hmf: 
Periplasmic scaffold of the Campylobacter jejuni flagellar motor (alpha carbon trace)
Method: single particle / : Drobnic T, Beeby M

EMDB-17415: 
Campylobacter jejuni flagellar motor, pflC deletion
Method: subtomogram averaging / : Drobnic T, Alzheimer M, Svensson S, Sharma CS, Beeby M

EMDB-17416: 
Campylobacter jejuni flagellar motor, pflD deletion
Method: subtomogram averaging / : Drobnic T, Henderson LD, Alzheimer M, Svensson S, Sharma CM, Beeby M

EMDB-17417: 
Campylobacter jejuni flagellar motor, truncated PflA (d16-168)
Method: subtomogram averaging / : Drobnic T, Nans A, Rosenthal PB, Beeby M

EMDB-17419: 
Campylobacter jejuni flagellar motor, FlgQ-mCherry fusion
Method: subtomogram averaging / : Drobnic T, Hendrixson DR, Beeby M

EMDB-19642: 
Campylobacter jejuni bacterial flagellar C-ring
Method: subtomogram averaging / : Beeby M

EMDB-16723: 
Wild-type Campylobacter jejuni flagellar motor, in situ
Method: single particle / : Drobnic T, Cohen EJ, Calcraft T, Singh NK, Nans A, Rosenthal PB, Beeby M

EMDB-16724: 
Periplasmic scaffold of the Campylobacter jejuni flagellar motor
Method: single particle / : Drobnic T, Cohen EJ, Singh NK, Umrekar TR, Nans A, Rosenthal PB, Beeby M

EMDB-17309: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

EMDB-17311: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

EMDB-17312: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

EMDB-17313: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

EMDB-17314: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

EMDB-17315: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

EMDB-17316: 
In situ subtomogram average of Prototype Foamy Virus Env trimer
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

EMDB-17317: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

EMDB-17318: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

EMDB-17319: 
In situ subtomogram average of the Prototype Foamy Virus capsid, wild-type Gag
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

EMDB-17320: 
In situ subtomogram average of the Prototype Foamy Virus capsid, p68 Gag
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

EMDB-17321: 
Cryotomogram of Prototype Foamy Virus particles, wild-type Gag
Method: electron tomography / : Calcraft T, Nans A, Rosenthal PB
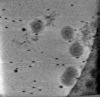
EMDB-17322: 
Cryotomogram of Prototype Foamy Virus particles, p68 Gag
Method: electron tomography / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozh: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozj: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozk: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozl: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozm: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozn: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction
Method: single particle / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozp: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

PDB-8ozq: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers
Method: subtomogram averaging / : Calcraft T, Nans A, Rosenthal PB

EMDB-17449: 
S. cerevisiae nexus-sCMGE after DNA replication initiation
Method: single particle / : Henrikus SS, Willhoft O
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
