-Search query
-Search result
Showing 1 - 50 of 104 items for (author: liu & jx)

EMDB-61426: 
The complex structure of 0086-0043 and NET determined with Cryo-EM.
Method: single particle / : Jia YJ, Gao B, Tan JX, Yan CY, Zhang W, Lan YY

EMDB-64502: 
Structure of dimeric FKS1 in complex with tRNA
Method: single particle / : Li JL, Zhu AQ, Liu JX, Dai XL, Wang X, Yan CY, Deng D

EMDB-62529: 
Cryo-EM map of carboxysomal midi-shell: T = 16 shell under C1 symmetry
Method: single particle / : Li JX, Li TP, Wang SM, Zhang YZ, Liu LN, Wang P

EMDB-62530: 
Cryo-EM map of carboxysomal midi-shell: T = 9 shell under C1 symmetry
Method: single particle / : Li JX, Li TP, Wang SM, Zhang YZ, Liu LN, Wang P

EMDB-61152: 
Cryo-EM structure of Bat SARS-like coronavirus Khosta-2 spike protein
Method: single particle / : Pan XQ, Li LJ, Liu KF, Qi JX, Gao GF

EMDB-61159: 
Cryo-EM structure of Bat SARS-like coronavirus Khosta-2 spike protein in complex with human ACE2 (local refined)
Method: single particle / : Pan XQ, Li LJ, Liu KF, Qi JX, Gao GF

EMDB-61170: 
Cryo-EM structure of Bat SARS-like coronavirus Khosta-1 spike protein
Method: single particle / : Pan XQ, Li LJ, Liu KF, Qi JX, Gao GF

EMDB-38008: 
Cryo-EM structure of H5 hemagglutinin from human-infecting H5N8 influenza virus in complex with Sialyl-Lewis X
Method: single particle / : Jin XY, Han P, Song H, Qi JX

EMDB-37532: 
Structure of hemagglutinin from Asiatic toad influenza-like virus
Method: single particle / : Zhang D, Xie YF, Chai Y, Shi Y, Liu WJ, Qi JX, Gao GF, Gao F

EMDB-37612: 
Structure of 1,3-beta-glucan synthase component FKS1
Method: single particle / : Li JL, Zhu AQ, Liu JX, Dai XL, Wang X, Yan CY, Deng D

EMDB-37614: 
Cryo-EM structure of the beta-1,3-glucan synthase FKS1-Rho1 complex
Method: single particle / : Li JL, Zhu AQ, Liu JX, Dai XL, Yan CY, Deng D, Wang X

EMDB-38773: 
Cryo-EM structure of BANAL-20-52 spike protein (6P)
Method: single particle / : Xu ZP, Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38778: 
Cryo-EM structure of CX1 spike protein (6P)
Method: single particle / : Xu ZP, Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38779: 
Cryo-EM structure of CX1 receptor binding domain in complex with human ACE2
Method: single particle / : Xu ZP, Li LJ, Gu YH, Qi JX, Gao GF

EMDB-39596: 
cryo-EM structure of carboxysomal midi-shell:icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 19)
Method: single particle / : Wang P, Li JX, Li TP, Li K, Ng PC, Wang SM, Chriscoli V, Basle A, Marles-Wright J, Zhang YZ, Liu LN

EMDB-39597: 
cryo-EM structure of carboxysomal midi-shell: icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 16)
Method: single particle / : Wang P, Li JX, LI TP, Li K, Ng PC, Wang SM, Chriscoli V, Basle A, Marles-Wright J, Zhang YZ, Liu LN

EMDB-39598: 
cryo-EM structure of carboxysomal midi-shell: icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 9)
Method: single particle / : Wang P, Li JX, Li TP, Li K, Ng PC, Wang SM, Chriscoli V, Basle A, Marles-Wright J, Zhang YZ, Liu LN

EMDB-39599: 
cryo-EM structure of carboxysomal midi-shell: assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T=9 Q=12)
Method: single particle / : Wang P, Li JX, Li TP, Li K, Ng PC, Wang SM, Chriscoli V, Basle A, Marles-Wright J, Zhang YZ, Liu LN

EMDB-39601: 
cryo-EM structure of carboxysomal midi-shell: icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 13)
Method: single particle / : Wang P, Li JX, Li TP, Li K, Ng PC, Wang SM, Chriscoli V, Basle A, Marles-Wright J, Zhang YZ, Liu LN

EMDB-39208: 
Cryo-EM structure of SARS-CoV-2 prototype RBD in complex with raccoon dog ACE2 (local refinement)
Method: single particle / : Li LJ, Luo CL, Qi JX, Gao GF

EMDB-39229: 
Cryo-EM structure of SARS-CoV-2 alpha variant spike protein in complex with raccoon dog ACE2 (local refinement)
Method: single particle / : Li LJ, Luo CL, Qi JX, Gao GF

EMDB-39209: 
Cryo-EM structure of SARS-CoV-2 prototype spike protein in complex with raccoon dog ACE2
Method: single particle / : Li LJ, Luo CL, Qi JX, Gao GF

EMDB-39224: 
Cryo-EM map of SARS-CoV-2 alpha variant spike protein in complex with raccoon dog ACE2
Method: single particle / : Li LJ, Luo CL, Qi JX, Gao GF

EMDB-37725: 
Cryo-EM structure of SARS-CoV-2 XBB.1.5 receptor-binding domain (RBD) complexed with CB6 mutant,S309, and S304 antibodies
Method: single particle / : Su C, Qi JX, Gao GF

EMDB-37726: 
Cryo-EM structure of SARS-CoV-2 receptor-binding domain (RBD) complexed with CB6 mutant,S309, and S304 antibodies
Method: single particle / : Su C, Qi JX, Gao GF

EMDB-61053: 
cryo-EM structure of human cystic fibrosis transmembrane conductance regulator (CFTR) from Biortus
Method: single particle / : Cao S, Shi H, Yang YH, Li JX, Hu YF

EMDB-38208: 
Structure of radafaxine-bound state of the human Norepinephrine Transporter
Method: single particle / : Wu JX, Ji WM

EMDB-37711: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 RBD in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38460: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P), 1-RBD-up state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38463: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38476: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38488: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38495: 
SARS-CoV-2 Omicron EG.5.1 RBD in complex with human ACE2 (local refined from the spike protein)
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38496: 
SARS-CoV-2 Omicron HV.1 RBD in complex with human ACE2 (local refinement from the spike protein)
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38498: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38502: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38505: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38826: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38827: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 RBD in complex with human ACE2 (local refinement from the spike protein)
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38937: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38983: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 RBD in complex with human ACE2 and S309 Fab
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38209: 
Structural mechanism of substrate binding and inhibition of the human Norepinephrine Transporter
Method: single particle / : Ji WM, Wu JX

EMDB-38210: 
Structure of apo state of the human Norepinephrine Transporter
Method: single particle / : Wu JX, Ji WM

EMDB-36672: 
Cryo-EM structure of the N-terminal domain of Omicron BA.1 in complex with nanobody N235 and S2L20 Fab
Method: single particle / : Liu B, Liu HH, Han P, Qi JX

EMDB-36454: 
Cryo-EM structure of a Legionella effector complexed with actin and AMP
Method: single particle / : Zhou XT, Wang XF, Tan JX, Zhu YQ

EMDB-36455: 
Cryo-EM structure of a Legionella effector complexed with actin and ATP
Method: single particle / : Zhou XT, Wang XF, Tan JX, Zhu YQ

EMDB-35929: 
Cryo-EM structure of Ufd4 in complex with K29/48 triUb
Method: single particle / : Ai HS, Mao JX, Wu XW, Pan M, Liu L

EMDB-35931: 
cryo-EM structures of Ufd4 in complex with Ubc4-Ub
Method: single particle / : Ai HS, Mao JX, Wu XW, Cai HY, Pan M, Liu L
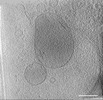
EMDB-39012: 
Representative tomogram of primary glioblastoma stem cell with circular inter-mitochondrial junctions.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39015: 
Representative tomogram of microglia cell with nanotunnel-like structures resembling mitochondrial fission.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
