-Search query
-Search result
Showing all 49 items for (author: li & hx)

EMDB-80359: 
SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67
Method: single particle / : Jiang XY, Qin Q, Qian JQ, Zhu HX, Huang Q

EMDB-48668: 
Activated Leptotrichia buccalis (Lbu) CRISPR-Cas13a bound to AI-designed anti-CRISPR AIcrVIA1
Method: single particle / : Taveneau C, Knott GJ

EMDB-63216: 
Cryo-EM structure of prefusion-stabilized RSV F (DS-Cav1 strain: A2) in complex with nanobody 1G9
Method: single particle / : Wang QQ, Ke XL, Li ET, Hong DX, Li HX, Cheng ZK, Zhang JC, Jin TC, Shu B, Chiu S

EMDB-63217: 
Cryo-EM structure of prefusion-stabilized RSV F (DS-Cav1 strain: A2) in complex with nanobody 1D8
Method: single particle / : Wang QQ, Ke XL, Li ET, Hong DX, Li HX, Cheng ZK, Zhang JC, Jin TC, Shu B, Chiu S

EMDB-48093: 
NCS.1 Fab in complex with N5 NA of A/shorebird/Delaware Bay/309/2016 (DB16, H10N5) -- 4 Fabs
Method: single particle / : Borst AJ

EMDB-48101: 
NCS.1 Fab in complex with N5 NA of A/shorebird/Delaware Bay/309/2016 (DB16, H10N5) -- 3 Fabs
Method: single particle / : Borst AJ

EMDB-48102: 
NCS.1.1 Fab in complex with the sNAp of A/California/04/2009 (CA09, H1N1) -- 4 Fabs [C4 Reconstruction]
Method: single particle / : Borst AJ

EMDB-70264: 
NCS.1.1 Fab in complex with the sNAp of A/California/04/2009 (CA09, H1N1) -- 4 Fabs [C1 Reconstruction]
Method: single particle / : Borst AJ

EMDB-38802: 
ASFV RNAP M1249L C-tail occupied complex4 (MCOC4)
Method: single particle / : Zhu GL, Zhu Y, Zhu ZX, Sun F, Zheng HX

EMDB-38745: 
ASFV RNAP elongation complex
Method: single particle / : Zhu GL, Zhu Y, Zhu ZX, Sun F, Zheng HX

EMDB-38746: 
ASFV RNAP M1249L C-tail occupied complex1 (MCOC1)
Method: single particle / : Zhu GL, Zhu Y, Zhu ZX, Sun F, Zheng HX

EMDB-38757: 
ASFV RNAP core complex
Method: single particle / : Zhu GL, Zhu Y, Zhu ZX, Sun F, Zheng HX

EMDB-38760: 
ASFV RNAP M1249L C-tail occupied complex2 (MCOC2)
Method: single particle / : Zhu GL, Zhu Y, Zhu ZX, Sun F, Zheng HX

EMDB-38767: 
ASFV RNAP M1249L C-tail occupied complex3 (MCOC3)
Method: single particle / : Zhu GL, Zhu Y, Zhu ZX, Sun F, Zheng HX

EMDB-36850: 
SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67
Method: single particle / : Jiang XY, Qin Q, Qian JQ, Zhu HX, Huang Q

EMDB-39012: 
Representative tomogram of primary glioblastoma stem cell with circular inter-mitochondrial junctions.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39015: 
Representative tomogram of microglia cell with nanotunnel-like structures resembling mitochondrial fission.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39019: 
Representative tomogram of glioblastoma cell with nanotunnel-like structure and inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39021: 
Representative tomogram of normal human astrocyte with nanotunnel-like structure which is an extension of the mitochondrial outer membrane.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39023: 
Representative tomogram of primary glioblastoma differentiated cell with parallel inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39024: 
Representative tomogram of primary glioblastoma stem cell with clustered mitochondria bearing various long-short axis ratios.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-32958: 
MERS-CoV spike complex with S41 neutralizing antibody Fab Class4 (2u1d RBD with 3Fab)
Method: single particle / : Zeng JW, Zhang SY, Zhou HX, Wang XW

EMDB-32961: 
MERS-CoV spike complex with S41 neutralizing antibody Fab Class3 (2u1d RBD with 2Fab)
Method: single particle / : Zeng JW, Zhang SY, Zhou HX, Wang XW

EMDB-32962: 
MERS-CoV spike complex with S41 neutralizing antibody Fab Class2 (1u2d RBD with 2Fab)
Method: single particle / : Zeng JW, Zhang SY, Zhou HX, Wang XW

EMDB-32963: 
MERS-CoV spike complex with S41 neutralizing antibody Fab Class1 (1u2d RBD with 1Fab)
Method: single particle / : Zeng JW, Zhang SY, Zhou HX, Wang XW

EMDB-24642: 
SARS-CoV-2 Spike bound to Fab PDI 210
Method: single particle / : Pymm P, Glukhova A, Black K, Tham WH

EMDB-24643: 
SARS-CoV-2 Spike bound to Fab PDI 96
Method: single particle / : Pymm P, Glukhova A, Black K, Tham WH

EMDB-24644: 
SARS-CoV-2 Spike bound to Fab PDI 215
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24645: 
SARS-CoV-2 Spike bound to Fab WCSL 119
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24646: 
SARS-CoV-2 Spike bound to Fab WCSL 129
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24647: 
SARS-CoV-2 Spike bound to Fab PDI 93
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24648: 
SARS-CoV-2 Spike bound to Fab PDI 222
Method: single particle / : Glukhova A, Pymm P, Black K, Tham WH

EMDB-24649: 
SARS-CoV-2 receptor binding domain bound to Fab PDI 222
Method: single particle / : Pymm P, Glukhova A

EMDB-30895: 
The structure of FC08 Fab-hA.CE2-RBD complex
Method: single particle / : Cao L, Wang X

EMDB-30573: 
A proof of concept for neutralizing antibody-guided vaccine design against SARS-CoV-2
Method: single particle / : Cao L, Wang X

EMDB-30708: 
Cryo-EM structure of human TSC complex
Method: single particle / : Yang H, Yu Z, Chen X, Li J, Li N, Cheng J, Gao N, Yuan H, Ye D, Guan K, Xu Y

EMDB-30709: 
Masked core region of TSC complex
Method: single particle / : Yang H, Yu Z, Chen X, Li J, Li N, Cheng J, Gao N, Yuan H, Ye D, Guan K, Xu Y

EMDB-30710: 
Masked wing-b region of TSC complex
Method: single particle / : Yang H, Yu Z, Chen X, Li J, Li N, Cheng J, Gao N, Yuan H, Ye D, Guan K, Xu Y

EMDB-30711: 
Masked wing-a region of TSC complex
Method: single particle / : Yang H, Yu Z, Chen X, Li J, Li N, Cheng J, Gao N, Yuan H, Ye D, Guan K, Xu Y

EMDB-20735: 
Cryo-EM structure of CH235UCA bound to Man5-enriched CH505.N279K.G458Y.SOSIP.664
Method: single particle / : Acharya P

EMDB-9401: 
CH505 SOSIP.664 trimer in complex with DH522.2 Fab
Method: single particle / : Fera D, Harrison SC
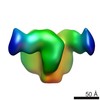
EMDB-8507: 
92BR SOSIP.664 trimer in complex with DH270.1 Fab
Method: single particle / : Fera D, Harrison SC
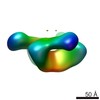
EMDB-8573: 
HIV-1 CH505 Transmitted Founder SOSIP.664 Env trimer in Complex with the DH576 Fab from the RV305 Trial
Method: single particle / : Fera D, Harrison SC
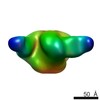
EMDB-8078: 
BG505 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235.12 wk323 Fab
Method: single particle / : Ozorowski G, Ward AB

EMDB-8079: 
B41 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235.12 wk323 Fab
Method: single particle / : Ozorowski G, Ward AB
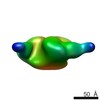
EMDB-8080: 
B41 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235.9 wk152 Fab
Method: single particle / : Ozorowski G, Ward AB

EMDB-8081: 
BG505 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235.9 wk152 Fab
Method: single particle / : Ozorowski G, Ward AB
EMDB-8082: 
BG505 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235 wk41 Fab
Method: single particle / : Ozorowski G, Ward AB
EMDB-8083: 
B41 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235 wk41 Fab
Method: single particle / : Ozorowski G, Ward AB
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
