[English] 日本語
 Yorodumi
Yorodumi- EMDB-8083: B41 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235 wk41 Fab -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8083 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | B41 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV CH235 wk41 Fab | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 24.0 Å | |||||||||
 Authors Authors | Ozorowski G / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2016 Journal: Cell / Year: 2016Title: Maturation Pathway from Germline to Broad HIV-1 Neutralizer of a CD4-Mimic Antibody. Authors: Mattia Bonsignori / Tongqing Zhou / Zizhang Sheng / Lei Chen / Feng Gao / M Gordon Joyce / Gabriel Ozorowski / Gwo-Yu Chuang / Chaim A Schramm / Kevin Wiehe / S Munir Alam / Todd Bradley / ...Authors: Mattia Bonsignori / Tongqing Zhou / Zizhang Sheng / Lei Chen / Feng Gao / M Gordon Joyce / Gabriel Ozorowski / Gwo-Yu Chuang / Chaim A Schramm / Kevin Wiehe / S Munir Alam / Todd Bradley / Morgan A Gladden / Kwan-Ki Hwang / Sheelah Iyengar / Amit Kumar / Xiaozhi Lu / Kan Luo / Michael C Mangiapani / Robert J Parks / Hongshuo Song / Priyamvada Acharya / Robert T Bailer / Allen Cao / Aliaksandr Druz / Ivelin S Georgiev / Young D Kwon / Mark K Louder / Baoshan Zhang / Anqi Zheng / Brenna J Hill / Rui Kong / Cinque Soto / / James C Mullikin / Daniel C Douek / David C Montefiori / Michael A Moody / George M Shaw / Beatrice H Hahn / Garnett Kelsoe / Peter T Hraber / Bette T Korber / Scott D Boyd / Andrew Z Fire / Thomas B Kepler / Lawrence Shapiro / Andrew B Ward / John R Mascola / Hua-Xin Liao / Peter D Kwong / Barton F Haynes /  Abstract: Antibodies with ontogenies from VH1-2 or VH1-46-germline genes dominate the broadly neutralizing response against the CD4-binding site (CD4bs) on HIV-1. Here, we define with longitudinal sampling ...Antibodies with ontogenies from VH1-2 or VH1-46-germline genes dominate the broadly neutralizing response against the CD4-binding site (CD4bs) on HIV-1. Here, we define with longitudinal sampling from time-of-infection the development of a VH1-46-derived antibody lineage that matured to neutralize 90% of HIV-1 isolates. Structures of lineage antibodies CH235 (week 41 from time-of-infection, 18% breadth), CH235.9 (week 152, 77%), and CH235.12 (week 323, 90%) demonstrated the maturing epitope to focus on the conformationally invariant portion of the CD4bs. Similarities between CH235 lineage and five unrelated CD4bs lineages in epitope focusing, length-of-time to develop breadth, and extraordinary level of somatic hypermutation suggested commonalities in maturation among all CD4bs antibodies. Fortunately, the required CH235-lineage hypermutation appeared substantially guided by the intrinsic mutability of the VH1-46 gene, which closely resembled VH1-2. We integrated our CH235-lineage findings with a second broadly neutralizing lineage and HIV-1 co-evolution to suggest a vaccination strategy for inducing both lineages. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8083.map.gz emd_8083.map.gz | 11.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8083-v30.xml emd-8083-v30.xml emd-8083.xml emd-8083.xml | 16 KB 16 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8083.png emd_8083.png | 43.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8083 http://ftp.pdbj.org/pub/emdb/structures/EMD-8083 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8083 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8083 | HTTPS FTP |
-Related structure data
| Related structure data |  8078C  8079C  8080C  8081C  8082C  5f96C  5f9oC  5f9wC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8083.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8083.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
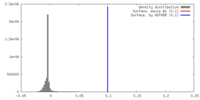
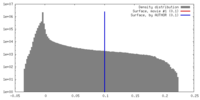
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Complex containing 2 copies of CH235 wk41 anti-HIV Fab bound to a...
| Entire | Name: Complex containing 2 copies of CH235 wk41 anti-HIV Fab bound to a trimer of HIV-1 Env B41 SOSIP.664 |
|---|---|
| Components |
|
-Supramolecule #1: Complex containing 2 copies of CH235 wk41 anti-HIV Fab bound to a...
| Supramolecule | Name: Complex containing 2 copies of CH235 wk41 anti-HIV Fab bound to a trimer of HIV-1 Env B41 SOSIP.664 type: complex / ID: 1 / Parent: 0 |
|---|---|
| Molecular weight | Theoretical: 420 KDa |
-Supramolecule #2: HIV-1 Env B41 SOSIP.664
| Supramolecule | Name: HIV-1 Env B41 SOSIP.664 / type: complex / ID: 2 / Parent: 1 Details: Solubilized and stabilized HIV-1 Env trimer from strain B41. Engineered disulfide between A501C and T605C. I559P mutation to stabilize in pre-fusion state. Truncation after D664 to increase ...Details: Solubilized and stabilized HIV-1 Env trimer from strain B41. Engineered disulfide between A501C and T605C. I559P mutation to stabilize in pre-fusion state. Truncation after D664 to increase solubility. Formed with three gp140 subunits. |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 / Strain: B41 Human immunodeficiency virus 1 / Strain: B41 |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK293F / Recombinant plasmid: pPPI4 Homo sapiens (human) / Recombinant cell: HEK293F / Recombinant plasmid: pPPI4 |
-Supramolecule #3: Anti-HIV CH235 wk41 antibody fragment antigen binding
| Supramolecule | Name: Anti-HIV CH235 wk41 antibody fragment antigen binding / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: Expi 293 / Recombinant plasmid: pcDNA3.1 Homo sapiens (human) / Recombinant cell: Expi 293 / Recombinant plasmid: pcDNA3.1 |
| Molecular weight | Theoretical: 50 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.03 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: 2% w/v uranyl formate Details: Negatively stained EM samples were prepared on carbon-coated Cu400 grids by applying sample for 10 seconds, blotting, applying 2% w/v uranyl formate for 60 seconds, and blotting again. | |||||||||
| Grid | Model: EMS / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Pretreatment - Type: PLASMA CLEANING |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Details | Collected a tilt series of -50, -40, -30, -20, -10, and 0 degrees. |
| Image recording | Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Sampling interval: 15.6 µm / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus min: 1.5 µm / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)