[English] 日本語
 Yorodumi
Yorodumi- PDB-5f9o: Crystal structure of broadly neutralizing VH1-46 germline-derived... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5f9o | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of broadly neutralizing VH1-46 germline-derived CD4-binding site-directed antibody CH235.09 in complex with HIV-1 clade A/E 93TH057 gp120 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | IMMUNE SYSTEM / antibody evolution HIV-1 broadly neutralizing CD4 binding site | ||||||
| Function / homology |  Function and homology information Function and homology informationmembrane fusion involved in viral entry into host cell / viral envelope / symbiont entry into host cell / virion attachment to host cell / virion membrane Similarity search - Function | ||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.86 Å SYNCHROTRON / Resolution: 1.86 Å | ||||||
 Authors Authors | Joyce, M.G. / Mascola, J.R. / Kwong, P.D. | ||||||
 Citation Citation |  Journal: Cell / Year: 2016 Journal: Cell / Year: 2016Title: Maturation Pathway from Germline to Broad HIV-1 Neutralizer of a CD4-Mimic Antibody. Authors: Mattia Bonsignori / Tongqing Zhou / Zizhang Sheng / Lei Chen / Feng Gao / M Gordon Joyce / Gabriel Ozorowski / Gwo-Yu Chuang / Chaim A Schramm / Kevin Wiehe / S Munir Alam / Todd Bradley / ...Authors: Mattia Bonsignori / Tongqing Zhou / Zizhang Sheng / Lei Chen / Feng Gao / M Gordon Joyce / Gabriel Ozorowski / Gwo-Yu Chuang / Chaim A Schramm / Kevin Wiehe / S Munir Alam / Todd Bradley / Morgan A Gladden / Kwan-Ki Hwang / Sheelah Iyengar / Amit Kumar / Xiaozhi Lu / Kan Luo / Michael C Mangiapani / Robert J Parks / Hongshuo Song / Priyamvada Acharya / Robert T Bailer / Allen Cao / Aliaksandr Druz / Ivelin S Georgiev / Young D Kwon / Mark K Louder / Baoshan Zhang / Anqi Zheng / Brenna J Hill / Rui Kong / Cinque Soto / / James C Mullikin / Daniel C Douek / David C Montefiori / Michael A Moody / George M Shaw / Beatrice H Hahn / Garnett Kelsoe / Peter T Hraber / Bette T Korber / Scott D Boyd / Andrew Z Fire / Thomas B Kepler / Lawrence Shapiro / Andrew B Ward / John R Mascola / Hua-Xin Liao / Peter D Kwong / Barton F Haynes /  Abstract: Antibodies with ontogenies from VH1-2 or VH1-46-germline genes dominate the broadly neutralizing response against the CD4-binding site (CD4bs) on HIV-1. Here, we define with longitudinal sampling ...Antibodies with ontogenies from VH1-2 or VH1-46-germline genes dominate the broadly neutralizing response against the CD4-binding site (CD4bs) on HIV-1. Here, we define with longitudinal sampling from time-of-infection the development of a VH1-46-derived antibody lineage that matured to neutralize 90% of HIV-1 isolates. Structures of lineage antibodies CH235 (week 41 from time-of-infection, 18% breadth), CH235.9 (week 152, 77%), and CH235.12 (week 323, 90%) demonstrated the maturing epitope to focus on the conformationally invariant portion of the CD4bs. Similarities between CH235 lineage and five unrelated CD4bs lineages in epitope focusing, length-of-time to develop breadth, and extraordinary level of somatic hypermutation suggested commonalities in maturation among all CD4bs antibodies. Fortunately, the required CH235-lineage hypermutation appeared substantially guided by the intrinsic mutability of the VH1-46 gene, which closely resembled VH1-2. We integrated our CH235-lineage findings with a second broadly neutralizing lineage and HIV-1 co-evolution to suggest a vaccination strategy for inducing both lineages. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5f9o.cif.gz 5f9o.cif.gz | 334.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5f9o.ent.gz pdb5f9o.ent.gz | 267.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5f9o.json.gz 5f9o.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/f9/5f9o https://data.pdbj.org/pub/pdb/validation_reports/f9/5f9o ftp://data.pdbj.org/pub/pdb/validation_reports/f9/5f9o ftp://data.pdbj.org/pub/pdb/validation_reports/f9/5f9o | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  8078C  8079C  8080C  8081C  8082C  8083C  5f96C  5f9wC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Antibody , 2 types, 2 molecules HL
| #2: Antibody | Mass: 25218.555 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  Homo sapiens (human) Homo sapiens (human) |
|---|---|
| #3: Antibody | Mass: 23532.176 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  Homo sapiens (human) Homo sapiens (human) |
-Protein / Sugars , 2 types, 13 molecules G

| #1: Protein | Mass: 39082.320 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Human immunodeficiency virus 1 / Gene: HIV-1 Env / Production host: Human immunodeficiency virus 1 / Gene: HIV-1 Env / Production host:  Homo sapiens (human) / References: UniProt: A0A0M3KKW9 Homo sapiens (human) / References: UniProt: A0A0M3KKW9 |
|---|---|
| #4: Sugar | ChemComp-NAG / |
-Non-polymers , 3 types, 463 molecules 
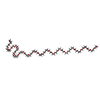



| #5: Chemical | | #6: Chemical | ChemComp-15P / | #7: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.77 Å3/Da / Density % sol: 55.52 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop Details: 9% (w/v) PEG8000, 19% (w/v) of PEG400, 0.1 M HEPES pH 7.5 PH range: 7.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1 Å / Beamline: 22-ID / Wavelength: 1 Å |
| Detector | Type: RAYONIX MX300-HS / Detector: CCD / Date: Aug 23, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8599→50 Å / Num. obs: 73935 / % possible obs: 89.7 % / Redundancy: 3.4 % / Net I/σ(I): 7.06 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.86→35.815 Å / SU ML: 0.19 / Cross valid method: FREE R-VALUE / σ(F): 1.35 / Phase error: 25.48 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.86→35.815 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 3.2729 Å / Origin y: -23.8821 Å / Origin z: -27.8733 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller












 PDBj
PDBj







