[English] 日本語
 Yorodumi
Yorodumi- EMDB-36850: SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-asse... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67 | |||||||||
 Map data Map data | SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | trivalent nanobody / viral neutralization / Omicron BA.1 / 3-RBD-up conformation / VIRAL PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 9.0 Å | |||||||||
 Authors Authors | Jiang XY / Qin Q / Qian JQ / Zhu HX / Huang Q | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: J Nanobiotechnology / Year: 2024 Journal: J Nanobiotechnology / Year: 2024Title: Computational design and engineering of self-assembling multivalent microproteins with therapeutic potential against SARS-CoV-2. Authors: Qin Qin / Xinyi Jiang / Liyun Huo / Jiaqiang Qian / Hongyuan Yu / Haixia Zhu / Wenhao Du / Yuhui Cao / Xing Zhang / Qiang Huang /  Abstract: Multivalent drugs targeting homo-oligomeric viral surface proteins, such as the SARS-CoV-2 trimeric spike (S) protein, have the potential to elicit more potent and broad-spectrum therapeutic ...Multivalent drugs targeting homo-oligomeric viral surface proteins, such as the SARS-CoV-2 trimeric spike (S) protein, have the potential to elicit more potent and broad-spectrum therapeutic responses than monovalent drugs by synergistically engaging multiple binding sites on viral targets. However, rational design and engineering of nanoscale multivalent protein drugs are still lacking. Here, we developed a computational approach to engineer self-assembling trivalent microproteins that simultaneously bind to the three receptor binding domains (RBDs) of the S protein. This approach involves four steps: structure-guided linker design, molecular simulation evaluation of self-assembly, experimental validation of self-assembly state, and functional testing. Using this approach, we first designed trivalent constructs of the microprotein miniACE2 (MP) with different trimerization scaffolds and linkers, and found that one of the constructs (MP-5ff) showed high trimerization efficiency, good conformational homogeneity, and strong antiviral neutralizing activity. With its trimerization unit (5ff), we then engineered a trivalent nanobody (Tr67) that exhibited potent and broad neutralizing activity against the dominant Omicron variants, including XBB.1 and XBB.1.5. Cryo-EM complex structure confirmed that Tr67 stably binds to all three RBDs of the Omicron S protein in a synergistic form, locking them in the "3-RBD-up" conformation that could block human receptor (ACE2) binding and potentially facilitate immune clearance. Therefore, our approach provides an effective strategy for engineering potent protein drugs against SARS-CoV-2 and other deadly coronaviruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36850.map.gz emd_36850.map.gz | 65 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36850-v30.xml emd-36850-v30.xml emd-36850.xml emd-36850.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36850_fsc.xml emd_36850_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_36850.png emd_36850.png | 84.4 KB | ||
| Filedesc metadata |  emd-36850.cif.gz emd-36850.cif.gz | 4.6 KB | ||
| Others |  emd_36850_additional_1.map.gz emd_36850_additional_1.map.gz emd_36850_half_map_1.map.gz emd_36850_half_map_1.map.gz emd_36850_half_map_2.map.gz emd_36850_half_map_2.map.gz | 77.9 MB 65.2 MB 65.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36850 http://ftp.pdbj.org/pub/emdb/structures/EMD-36850 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36850 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36850 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36850.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36850.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.57 Å | ||||||||||||||||||||||||||||||||||||
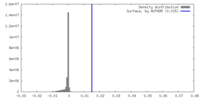
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: SARS-CoV-2 Omicron BA.1 spike protein in complex with...
| File | emd_36850_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67 | ||||||||||||
| Projections & Slices |
| ||||||||||||
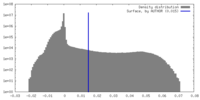
| Density Histograms |
-Half map: Half map 1
| File | emd_36850_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
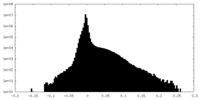
| Density Histograms |
-Half map: Half map 2
| File | emd_36850_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
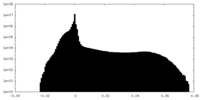
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-asse...
| Entire | Name: SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67 |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-asse...
| Supramolecule | Name: SARS-CoV-2 Omicron BA.1 spike protein in complex with a self-assembling trivalent nanobody Tr67 type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: Trivalent nanobody
| Supramolecule | Name: Trivalent nanobody / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Supramolecule #3: SARS-CoV-2 Omicron BA.1 spike protein
| Supramolecule | Name: SARS-CoV-2 Omicron BA.1 spike protein / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: 10 mM HEPES,150 mM NaCl,1% glycerol | ||||||||||||
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 296.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 43.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.8 µm / Nominal magnification: 92000 |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)