-検索条件
-検索結果
検索 (著者・登録者: ivanov & p)の結果96件中、1から50件目までを表示しています

EMDB-50672: 
A 3.3A sub-tomogram average of HIV-1 CA-SP1 from 5 tomograms in EMPIAR-10164 obtained using RELION 5

EMDB-19003: 
The structure of inactivated mature tick-borne encephalitis virus

PDB-8r8l: 
The structure of inactivated mature tick-borne encephalitis virus

EMDB-17777: 
Engineered glycolyl-CoA carboxylase (G20R variant) with bound CoA

EMDB-17778: 
Engineered glycolyl-CoA carboxylase (G20R variant) with bound CoA

PDB-8pn7: 
Engineered glycolyl-CoA carboxylase (G20R variant) with bound CoA

PDB-8pn8: 
Engineered glycolyl-CoA carboxylase (L100N variant) with bound CoA

EMDB-16872: 
Subtomogram average of long bridges of the yeast ER-mitochondria encounter structure (ERMES). The population half containing longer bridge structures was averaged.

EMDB-16873: 
Subtomogram average of bridges of the yeast ER-mitochondria encounter structure (ERMES)

EMDB-16871: 
Subtomogram average of short bridges of the yeast ER-mitochondria encounter structure (ERMES). The population half containing shorter bridge structures was averaged.
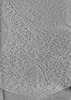

EMDB-15355: 
Electron cryo-tomography of the ER-mitochondria encounter structure ERMES

PDB-8bsh: 
COPII inner coat

EMDB-16169: 
Alpha7-nAChR extracellular ligand-binding domain (alpha7-ECD)in complex with alpha-bungarotoxin.

EMDB-16173: 
Alpha7-nAChR extracellular ligand-binding domain (alpha7-ECD)in complex with a weak neurotoxin WTX.

EMDB-16183: 
In situ structure of the Caulobacter crescentus S-layer

EMDB-16207: 
HIV-1 CA-SP1 subtomogram average with Relion4 from EMPIAR-10164 dataset, 3.2 A from 5 tomograms

EMDB-16209: 
HIV-1 CA-SP1 subtomogram average with Relion4 from EMPIAR-10164 dataset, 3.0 A from the full dataset

PDB-8bqe: 
In situ structure of the Caulobacter crescentus S-layer

EMDB-15949: 
COPII inner coat reprocessed with relion4.0

EMDB-13619: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state III (composite map)

EMDB-13620: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, state I

EMDB-13621: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state III (B1 map)

EMDB-13622: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state III (B2 map)

EMDB-13623: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state III (B3 map)

EMDB-13624: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state III (3D auto-refined map)

EMDB-13629: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state II (composite map)

EMDB-13631: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state II (B1 map)

EMDB-13635: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state II (B2 map)

EMDB-13640: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state II (3D auto-refined map)

EMDB-13644: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state I (3D auto-refined map)

EMDB-13645: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, state I (B1 map)

EMDB-13646: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, state I (B2 map)

EMDB-13647: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, state I (3D auto-refined map)

EMDB-13648: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state (3D auto-refined map)

EMDB-13649: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state A (binned 3D auto-refined map)

EMDB-13650: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state B (binned 3D auto-refined map)

EMDB-13651: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state C (binned 3D auto-refined map)

EMDB-13652: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state D (binned 3D auto-refined map)

EMDB-13653: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state E (binned 3D auto-refined map)

EMDB-13655: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, swiveled state F (binned 3D auto-refined map)

EMDB-13656: 
Structure of double-stranded DNA-bound MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATP (3D auto-refined map)

EMDB-13657: 
Structure of double-stranded DNA-bound MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATP (B1 map)

EMDB-13658: 
Structure of double-stranded DNA-bound MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATP (B2 map)

EMDB-13659: 
Structure of double-stranded DNA-bound MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATP (B3 map)

PDB-7pt6: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ATPgS, state III

PDB-7pt7: 
Structure of MCM2-7 DH complexed with Cdc7-Dbf4 in the presence of ADP:BeF3, state I

EMDB-14132: 
Bovine complex I in lipid nanodisc, Active-Q10

EMDB-14133: 
Bovine complex I in lipid nanodisc, Active-apo
EMDB-14134: 
Bovine complex I in lipid nanodisc, Deactive-ligand (composite)
EMDB-14139: 
Bovine complex I in lipid nanodisc, Deactive-apo
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
