-検索条件
-検索結果
検索 (著者・登録者: hess & s)の結果全32件を表示しています

EMDB-29783: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074

EMDB-41613: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074

PDB-8g6u: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074

PDB-8ttw: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074

EMDB-28378: 
Structure of the C3bB proconvertase in complex with lufaxin and factor Xa

PDB-8eok: 
Structure of the C3bB proconvertase in complex with lufaxin and factor Xa

EMDB-28279: 
Structure of the C3bB proconvertase in complex with lufaxin

PDB-8enu: 
Structure of the C3bB proconvertase in complex with lufaxin

EMDB-27596: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer in complex with 8ANC195 and 10-1074

PDB-8dok: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer in complex with 8ANC195 and 10-1074

EMDB-27103: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with Temsavir, 8ANC195, and 10-1074

PDB-8czz: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with Temsavir, 8ANC195, and 10-1074
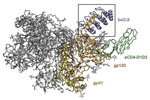
EMDB-26157: 
Cryo-EM structure of BG505 SOSIP HIV-1 Env trimer in complex with CD4 receptor (D1D2) and broadly neutralizing darpin bnD.9

PDB-7txd: 
Cryo-EM structure of BG505 SOSIP HIV-1 Env trimer in complex with CD4 receptor (D1D2) and broadly neutralizing darpin bnD.9

EMDB-33233: 
Cryo-EM structure of EDS1 and SAG101 with ATP-APDR

PDB-7xjp: 
Cryo-EM structure of EDS1 and SAG101 with ATP-APDR

EMDB-33144: 
Cryo-EM structure of EDS1 and PAD4

PDB-7xdd: 
Cryo-EM structure of EDS1 and PAD4

EMDB-12700: 
Structure of the C9orf72-SMCR8 complex

PDB-7o2w: 
Structure of the C9orf72-SMCR8 complex

EMDB-10049: 
THE STRUCTURE OF BD OXIDASE FROM ESCHERICHIA COLI

PDB-6rx4: 
THE STRUCTURE OF BD OXIDASE FROM ESCHERICHIA COLI

EMDB-4584: 
Structure and assembly of the mitochondrial membrane remodelling GTPase Mgm1
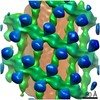
EMDB-10062: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes
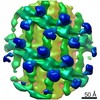
EMDB-10063: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes in the GTPgammaS bound state

EMDB-10064: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes

EMDB-10065: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes in the GTPgammaS bound state

PDB-6rzt: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes

PDB-6rzu: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes in the GTPgammaS bound state

PDB-6rzv: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes

PDB-6rzw: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes in the GTPgammaS bound state

EMDB-3401: 
Electron cryo-microscopy of CSN-SCF-N8 complex
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
