-Search query
-Search result
Showing 1 - 50 of 61 items for (author: grotjahn & da)

EMDB-56238: 
In situ cryo-ET subtomogram averaged map of Flotillin complex
Method: subtomogram averaging / : Li D, Lizarrondo J, Wilfling F

EMDB-56295: 
In situ cryo-ET tomogram of a lysosomal structure in untreated HeLa TMEM192-3xHA cell.
Method: electron tomography / : Li D, Wilfling F

EMDB-56296: 
In situ cryo-ET tomogram of lysosome damaged by LLOMe (0.5mM, 60min) in HeLa TMEM192-3xHA cell.
Method: electron tomography / : Li D, Wilfling F

EMDB-56297: 
In situ cryo-ET of lysosome damaged by LLOMe (0.5mM, 60min) encapsulated in an autophagosome in HeLa TMEM192-3xHA cell.
Method: electron tomography / : Li D, Wilfling F

EMDB-56298: 
In situ cryo-ET tomogram of lysosomes in BAPTA AM pre-treated (50uM, 30min) and LLOMe (0.5mM, 60min) treated TMEM192-3xHA HeLa cell.
Method: electron tomography / : Li D, Wilfling F

EMDB-56300: 
In situ cryo-ET tomogram of lysosomes in LLOMe (0.5mM, 60min) treated TMEM192-3xHA HeLa cell.
Method: electron tomography / : Li D, Wilfling F

EMDB-56327: 
In situ cryo-ET tomogram of lysosomal structure in untreated rat hippocampal neurons
Method: electron tomography / : Li D, Schwarz A, Wilfling F

EMDB-56329: 
In situ cryo-ET tomogram of lysosomes in E64d pre-treated (20uM, 30min) and LLOMe (0.5mM, 60min) treated TMEM192-3xHA HeLa cell.
Method: electron tomography / : Li D, Wilfling F

EMDB-56330: 
In situ cryo-ET tomogram of lysosomal structure in LLOMe-treated (0.5mM, 1h) rat hippocampal neuron.
Method: electron tomography / : Li D, Schwarz A, Wilfling F

EMDB-73977: 
Masked Classification of Prohibitin Complexes Showing the Prohibitin complex without an Additional Matrix-Facing Density (Class 2)
Method: subtomogram averaging / : Medina M, Rahmani H, Chang Y, Barad BA, Grotjahn DA

EMDB-73978: 
Masked Classification of Prohibitin Complexes Showing the Prohibitin complex with an Additional Matrix-Facing Density (Class 1)
Method: subtomogram averaging / : Medina M, Rahmani H, Chang Y, Barad BA, Grotjahn DA

EMDB-72321: 
Structure of ATP synthase monomer from mouse embryonic fibroblasts
Method: subtomogram averaging / : Medina M, Chang Y, Rahmani H, Fuentes D, Barad BA, Grotjahn DA

EMDB-48751: 
Structrure of cytoplasmic S. cerevisiae ribosome oriented for protein import on the outer mitochondrial membrane
Method: subtomogram averaging / : Chang Y, Barad BA, Rahmani H, Zid BM, Grotjahn DA

EMDB-48752: 
Structure of cytoplasmic 80S ribosome from S. cerevisiae
Method: subtomogram averaging / : Chang Y, Barad BA, Rahmani H, Zid BM, Grotjahn DA

EMDB-51640: 
Subtomogram average of immature Langat virus from cryo-electron tomograms of infected cells
Method: subtomogram averaging / : Carlson LA, Dahmane S

EMDB-51642: 
Subtomogram average of mature Langat virus
Method: subtomogram averaging / : Carlson LA, Dahmane S

EMDB-42152: 
Representative tomogram of PolyP + No DNA
Method: electron tomography / : Racki LR, Deniz AA, Park D

EMDB-42153: 
Representative tomogram of PolyP + pUC19
Method: electron tomography / : Racki LR, Deniz AA, Park D

EMDB-42154: 
Representative tomogram of PolyP + pUC19 (10x)
Method: electron tomography / : Racki LR, Deniz AA, Park D
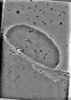
EMDB-42155: 
Representative tomogram of PolyP + 15kb DNA
Method: electron tomography / : Racki LR, Deniz AA, Park D

EMDB-42074: 
Representative tomogram of Enterococcus faecium WT Com15
Method: electron tomography / : Hang HC, Park D

EMDB-42086: 
Representative tomogram of Enterococcus faecium SagA complementation strain
Method: electron tomography / : Hang HC, Park D

EMDB-42087: 
Representative tomogram of Enterococcus faecium SagA deletion strain
Method: electron tomography / : Hang HC, Park D

EMDB-42884: 
Microtubule-TTLL6 map
Method: single particle / : Mahalingan KK, Grotjahn D, Li Y, Lander GC, Zehr EA, Roll-Mecak A

PDB-8u3z: 
Model of TTLL6 bound to microtubule from composite map
Method: single particle / : Mahalingan KK, Grotjahn D, Li Y, Lander GC, Zehr EA, Roll-Mecak A

EMDB-41022: 
Map of Tubulin Tyrosine Ligase Like 6
Method: single particle / : Mahalingan KK, Grotjahn D, Li Y, Lander GC, Zehr EA, Roll-Mecak A

EMDB-41090: 
Composite map of TTLL6 bound to unmodified human microtubule
Method: single particle / : Mahalingan KK, Grotjahn D, Li Y, Lander GC, Zehr EA, Roll-Mecak A

EMDB-41018: 
Microtubule-TTLL6 map
Method: single particle / : Mahalingan KK, Grotjahn D, Li Y, Lander GC, Zehr EA, Roll-Mecak A

PDB-8t42: 
Model of TTLL6 MTBH1-2 bound to microtubule
Method: single particle / : Mahalingan KK, Grotjahn D, Li Y, Lander GC, Zehr EA, Roll-Mecak A

EMDB-36766: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36767: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36768: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36769: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36770: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36771: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36772: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36807: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36809: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36810: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36811: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36812: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36813: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36814: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36816: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36817: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36818: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36819: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36820: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36821: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36822: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
