-Search query
-Search result
Showing 1 - 50 of 123 items for (author: dai & yn)

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10
Method: single particle / : Huang J, Ozorowski G, Ward AB

PDB-8sx3: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10
Method: single particle / : Huang J, Ozorowski G, Ward AB

EMDB-37671: 
Cryo EM map of SLC7A10 in the apo state
Method: single particle / : Li YN, Guo YY, Dai L, Yan RH

EMDB-37672: 
Cryo EM map of SLC7A10 with L-Alanine substrate
Method: single particle / : Li YN, Guo YY, Dai L, Yan RH

EMDB-37675: 
Cryo EM map of SLC7A10-SLC3A2 complex in the D-serine bound state
Method: single particle / : Li YN, Guo YY, Dai L, Yan RH

EMDB-43889: 
Chlamydomonas reinhardtii mastigoneme (constituent map 1)
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-43890: 
Chlamydomonas reinhardtii mastigoneme (constituent map 2)
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-43891: 
Chlamydomonas reinhardtii mastigoneme (constituent map 3)
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-43892: 
Composite cryo-EM map of the Chlamydomonas reinhardtii mastigoneme
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

PDB-9b4h: 
Chlamydomonas reinhardtii mastigoneme filament
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-35706: 
Cryo-EM structure of GIPR splice variant 1 (SV1) in complex with Gs protein
Method: single particle / : Zhao FH, Hang KN, Zhou QT, Shao LJ, Li H, Li WZ, Lin S, Dai AT, Cai XQ, Liu YY, Xu YN, Feng WB, Yang DH, Wang MW

EMDB-35707: 
Cryo-EM structure of GIPR splice variant 2 (SV2) in complex with Gs protein
Method: single particle / : Zhao FH, Hang KN, Zhou QT, Shao LJ, Li H, Li WZ, Lin S, Dai AT, Cai XQ, Liu YY, Xu YN, Feng WB, Yang DH, Wang MW

EMDB-35601: 
Cryo-EM structure of the alpha-MSH-bound human melanocortin receptor 5 (MC5R)-Gs complex
Method: single particle / : Feng WB, Zhou QT, Chen XY, Dai AT, Cai XQ, Liu X, Zhao FH, Chen Y, Ye CY, Xu YN, Cong ZT, Li H, Lin S, Yang DH, Wang MW

EMDB-35615: 
Cryo-EM structure of the gamma-MSH-bound human melanocortin receptor 3 (MC3R)-Gs complex
Method: single particle / : Feng WB, Zhou QT, Chen XY, Dai AT, Cai XQ, Liu X, Zhao FH, Chen Y, Ye CY, Xu YN, Cong ZT, Li H, Lin S

EMDB-35616: 
Cryo-EM structure of the PG-901-bound human melanocortin receptor 5 (MC5R)-Gs complex
Method: single particle / : Feng WB, Zhou QT, Chen XY, Dai AT, Cai XQ, Liu X, Zhao FH, Chen Y, Ye CY, Xu YN, Cong ZT, Li H, Lin S, Yang DH, Wang MW

EMDB-28083: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in the rigor form
Method: helical / : Doran MH, Lehman W, Rynkiewicz MJ

EMDB-28270: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin loop 4 7G mutant complex
Method: helical / : Doran MH, Lehman W, Rynkiewicz MJ

PDB-8efi: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in the rigor form
Method: helical / : Doran MH, Lehman W, Rynkiewicz MJ

PDB-8enc: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin loop 4 7G mutant complex
Method: helical / : Doran MH, Lehman W, Rynkiewicz MJ
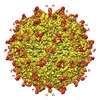
EMDB-25606: 
Zika Virus particle bound with IgM antibody DH1017 Fab fragment
Method: single particle / : Miller AS, Kuhn RJ

PDB-7t17: 
Zika Virus asymmetric unit bound with IgM antibody DH1017 Fab fragment
Method: single particle / : Miller AS, Kuhn RJ

PDB-7uti: 
ALTERNATIVE MODELING OF TROPOMYOSIN IN HUMAN CARDIAC THIN FILAMENT IN THE CALCIUM BOUND STATE
Method: single particle / : Rynkiewicz MJ, Pavadai E, Lehman W

PDB-7utl: 
ALTERNATIVE MODELING OF TROPOMYOSIN IN HUMAN CARDIAC THIN FILAMENT IN THE CALCIUM FREE STATE
Method: single particle / : Rynkiewicz MJ, Pavadai E, Lehman W
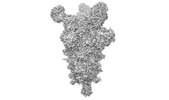
EMDB-26676: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-26677: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-26678: 
An antibody from single human VH-rearranging mouse neutralizes all SARS-CoV-2 variants through BA.5 by inhibiting membrane fusion
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-27043: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody SP1-77 Fab in complex with SARS-CoV2 2P spike
Method: single particle / : Edwards RJ, Mansouri K
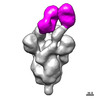
EMDB-27044: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody VHH7-7-53 Fab in complex with SARS-CoV2 2P spike
Method: single particle / : Edwards RJ, Mansouri K

EMDB-27046: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody VHH7-5-82 Fab in complex with SARS-CoV2 2P spike
Method: single particle / : Edwards RJ, Mansouri K

PDB-7upw: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

PDB-7upx: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

PDB-7upy: 
An antibody from single human VH-rearranging mouse neutralizes all SARS-CoV-2 variants through BA.5 by inhibiting membrane fusion
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-32095: 
Cryo-EM structure of human vasoactive intestinal polypeptide receptor 2 (VIP2R) in complex with PACAP27 and Gs
Method: single particle / : Xu YN, Feng WB, Zhou QT, Liang AY, Li J, Dai AT, Zhao FH, Yan JH, Chen CW, Li H, Zhao LH, Xia T, Jiang Y, Xu HE, Yang DH, Wang MW

EMDB-32401: 
Cryo-EM structure of N-terminal modified human vasoactive intestinal polypeptide receptor 2 (VIP2R) in complex with PACAP27 and Gs
Method: single particle / : Xu YN, Feng WB, Zhou QT, Liang AY, Li J, Dai AT, Zhao FH, Yan JH, Chen CW, Li H, Zhao LH, Xia T, Jiang Y, Xu HE, Yang DH, Wang MW

EMDB-24408: 
The Capsid Structure of the ChAdOx1 viral vector/chimpanzee adenovirus Y25
Method: single particle / : Baker AT, Boyd RJ

PDB-7rd1: 
The Capsid Structure of the ChAdOx1 viral vector/chimpanzee adenovirus Y25
Method: single particle / : Baker AT, Boyd RJ, Sarkar D, Vermaas JV, Williams D, Singharoy A

EMDB-11806: 
cryo-EM structure of ExbBD from Serratia Marcescens
Method: single particle / : Biou V, Adaixo R

EMDB-24642: 
SARS-CoV-2 Spike bound to Fab PDI 210
Method: single particle / : Pymm P, Glukhova A, Black K, Tham WH

EMDB-24643: 
SARS-CoV-2 Spike bound to Fab PDI 96
Method: single particle / : Pymm P, Glukhova A, Black K, Tham WH

EMDB-24644: 
SARS-CoV-2 Spike bound to Fab PDI 215
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24645: 
SARS-CoV-2 Spike bound to Fab WCSL 119
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24646: 
SARS-CoV-2 Spike bound to Fab WCSL 129
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24647: 
SARS-CoV-2 Spike bound to Fab PDI 93
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24648: 
SARS-CoV-2 Spike bound to Fab PDI 222
Method: single particle / : Glukhova A, Pymm P, Black K, Tham WH

EMDB-24649: 
SARS-CoV-2 receptor binding domain bound to Fab PDI 222
Method: single particle / : Pymm P, Glukhova A, Black KA, Tham WH

PDB-7rr0: 
SARS-CoV-2 receptor binding domain bound to Fab PDI 222
Method: single particle / : Pymm P, Glukhova A, Black KA, Tham WH

EMDB-23091: 
Reelin central fragment repeats 3-6, dimer
Method: subtomogram averaging / : Dai W, Chen M, Kuang X, Huynh K, D'Arcangelo G, Turk LS, Comoletti D

EMDB-23566: 
The SARS-CoV-2 spike protein receptor binding domain bound to neutralizing nanobodies WNb 2 and WNb 10
Method: single particle / : Pymm P, Glukhova A, Tham WH

EMDB-23567: 
Cryo-EM map of SARS-CoV-2 Spike protein bound to neutralising nanobodies WNb2 and WNb10 (C3 symmetry)
Method: single particle / : Pymm P, Tham WH, Glukhova A

EMDB-23568: 
Cryo-EM map of SARS-CoV-2 Spike protein in complex with neutralising nanobodies WNb2 and WNb10 (C1 symmetry)
Method: single particle / : Pymm P, Tham WH, Glukhova A
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
