-Search query
-Search result
Showing all 44 items for (author: cote & j)

EMDB-72652: 
Cereblon-DDB1 with Golcadomide
Method: single particle / : Watson ER, Lander GC

EMDB-72653: 
Cereblon with Golcadomide and Ikaros ZF1-2-3
Method: single particle / : Watson ER, Lander GC

PDB-9y7d: 
Cereblon with Golcadomide and Ikaros ZF1-2-3
Method: single particle / : Watson ER, Lander GC

EMDB-70451: 
SARS-COV-2-6P-MUT7 S PROTEIN-DY-III-281 complex closed conformation
Method: single particle / : Chandravanshi M, Niu L, Tolbert WD, Pazgier M

EMDB-70453: 
SARS-COV-2-6P-MUT7 S PROTEIN-DY-III-281 complex 1 RBD up conformation
Method: single particle / : Chandravanshi M, Niu L, Tolbert WD, Pazgier M

EMDB-70454: 
Apo SARS-COV-2-6P-MUT7 S PROTEIN closed conformation
Method: single particle / : Niu L, Chandravanshi M, Tolbert WD, Pazgier M

EMDB-70455: 
APO SARS-COV-2-6P-MUT7 S PROTEIN 1 RBD UP CONFORMATION
Method: single particle / : Niu L, Chandravanshi M, Tolbert WD, Pazgier M

EMDB-45206: 
Reconstituted P400 Subcomplex of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45240: 
P400 subcomplex of the native human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45252: 
ARP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c57: 
Reconstituted P400 Subcomplex of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c62: 
P400 subcomplex of the native human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c6n: 
ARP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45176: 
TRRAP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45180: 
Second BAF53a of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c47: 
TRRAP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c4b: 
Second BAF53a of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-41144: 
Cryo-EM Structure of GPR61-G protein complex stabilized by scFv16
Method: single particle / : Lees JA, Dias JM, Han S

EMDB-41145: 
Cryo-EM Structure of GPR61-
Method: single particle / : Lees JA, Dias JM, Han S

PDB-8tb0: 
Cryo-EM Structure of GPR61-G protein complex stabilized by scFv16
Method: single particle / : Lees JA, Dias JM, Han S

EMDB-27243: 
Cryo-EM map of MORF-WHs in complex with 197bp nucleosome aided by scFv
Method: single particle / : Zhou BR

EMDB-25200: 
CryoEM structure of SARS-CoV-2 HexaPro spike in complex with S2-binding CV3-25
Method: single particle / : Chen Y, Huang R, Pazgier M

EMDB-25564: 
Subtomogram averaging of SARS-CoV-2 Spike Protein bound to CV3-1
Method: subtomogram averaging / : Li W, Mothes W

EMDB-25565: 
Subtomogram averaging of SARS-CoV-2 Spike Protein bound to Fab CV3-25
Method: subtomogram averaging / : Li W, Mothes W

EMDB-25566: 
Subtomogram averaging of SARS-CoV-2 Spike Protein
Method: subtomogram averaging / : Li W, Mothes W

EMDB-10497: 
CryoEM structure of the binary DOCK2-ELMO1 complex
Method: single particle / : Chang L, Yang J

EMDB-10498: 
CryoEM structure of the ternary DOCK2-ELMO1-RAC1 complex
Method: single particle / : Chang L, Yang J

PDB-6tgb: 
CryoEM structure of the binary DOCK2-ELMO1 complex
Method: single particle / : Chang L, Yang J, Chang JH, Zhang Z, Boland A, McLaughlin SH, Abu-Thuraia A, Killoran RC, Smith MJ, Cote JF, Barford D

PDB-6tgc: 
CryoEM structure of the ternary DOCK2-ELMO1-RAC1 complex.
Method: single particle / : Chang L, Yang J, Chang JH, Zhang Z, Boland A, McLaughlin SH, Abu-Thuraia A, Killoran RC, Smith MJ, Cote JF, Barford D

EMDB-9193: 
Structure of Plasmodium falciparum CyRPA/Ripr invasion complex
Method: single particle / : Wilson W, Alan C, Zhiheng Y
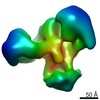
EMDB-7131: 
Chromatin-modifying complex
Method: single particle / : Setiaputra DT, Dalwadi U, Yip CK

EMDB-9192: 
Cryo-electron microscopy structure of Plasmodium falciparum Rh5/CyRPA/Ripr invasion complex
Method: single particle / : Wilson W, Zhiheng Y

PDB-6mpv: 
Cryo-electron microscopy structure of Plasmodium falciparum Rh5/CyRPA/Ripr invasion complex
Method: single particle / : Wilson W, Zhiheng Y, Cowman AF

PDB-5syc: 
Near-atomic resolution cryo-EM reconstruction of peloruside-stabilized microtubule
Method: helical / : Kellogg EH, Nogales E

PDB-5sye: 
Near-atomic resolution cryo-EM reconstruction of doubly bound Taxol- and peloruside-stabilized microtubule
Method: helical / : Kellogg EH, Nogales E

PDB-5syf: 
High-resolution cryo-EM reconstruction of Taxol-stabilized microtubule
Method: helical / : Kellogg EH, Nogales E

PDB-5syg: 
Cryo-EM reconstruction of zampanolide-bound microtubule
Method: helical / : Kellogg EH, Hejab NMA, Nogales E

EMDB-8320: 
Near-atomic resolution cryo-EM reconstruction of peloruside-stabilized microtubule
Method: helical / : Kellogg EH, Nogales E

EMDB-8321: 
Near-atomic resolution cryo-EM reconstruction of doubly bound Taxol- and peloruside-stabilized microtubule
Method: helical / : Kellogg EH, Nogales E

EMDB-8322: 
High-resolution cryo-EM reconstruction of Taxol-stabilized microtubule
Method: helical / : Kellogg EH, Nogales E

EMDB-8323: 
Cryo-EM reconstruction of zampanolide-bound microtubule
Method: helical / : Kellogg EH, Hejab NMA
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
