-Search query
-Search result
Showing 1 - 50 of 2,483 items for (author: chang & g)

EMDB-60915: 
Cryo-EM structure of mouse heavy-chain apoferritin resolved at 1.51 Angstroms

EMDB-39212: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)

EMDB-39213: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)

EMDB-39214: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)

EMDB-39215: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)

EMDB-39217: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH5.0 (4.36A)

PDB-8yf6: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)

PDB-8yf7: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)

PDB-8yf8: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)

PDB-8yf9: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)

EMDB-41770: 
Apo form of human ATE1

EMDB-42071: 
human ATE1 in complex with Arg-tRNA and a peptide substrate

PDB-8tzv: 
Apo form of human ATE1

PDB-8uau: 
human ATE1 in complex with Arg-tRNA and a peptide substrate

EMDB entry, No image
EMDB-51190: 
High-resolution structure of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-activator Cdh1

PDB-9gaw: 
High-resolution structure of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-activator Cdh1

EMDB-43835: 
Structure of biofilm-forming functional amyloid PSMa1 from Staphylococcus aureus

EMDB-42118: 
Toxoplasma gondii apical end with ICMAP1 conditionally knocked down

EMDB-42119: 
Toxoplasma gondii apical end with ICMAP2 conditionally knocked down

EMDB-42120: 
Toxoplasma gondii apical end with ICMAP3i conditionally knocked down

EMDB-42121: 
Toxoplasma gondii apical end with ICMAP3ii conditionally knocked down
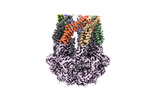
EMDB-36965: 
CryoEM structure of LonC protease hepatmer, apo state
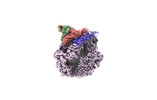
EMDB-36966: 
CryoEM structure of LonC protease open hexamer, apo state
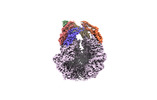
EMDB-36967: 
CryoEM of LonC open pentamer, apo state
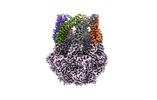
EMDB-36968: 
CryoEM structure of LonC heptamer in presence of AGS
EMDB-36969: 
CryoEM structure of LonC protease hexamer in presence of AGS
EMDB-36970: 
CryoEM structure of LonC protease open pentamer in presence of AGS
EMDB-36971: 
CryoEM structure of LonC S582A hepatmer with Lysozyme
EMDB-36972: 
CryoEM structure of LonC S582A hexamer with Lysozyme
EMDB-36973: 
CryoEM structure of LonC protease S582A open hexamer with lysozyme
EMDB-36974: 
CryoEM structure of LonC protease S582A open pentamer with lysozyme
EMDB-36975: 
CryoEM structure of LonC protease open Hexamer, AGS
EMDB-36976: 
CryoEM structure of LonC protease hepatmer with Bortezomib
EMDB-36977: 
CryoEM structure of LonC protease hexamer with Bortezomib

EMDB-36978: 
CryoEM map of LonC protease open hexamer with Bortezomib

EMDB-36979: 
CryoEM map of LonC protease heptamer (heated at 343K)

PDB-8k8v: 
CryoEM structure of LonC protease hepatmer, apo state

PDB-8k8w: 
CryoEM structure of LonC protease open hexamer, apo state

PDB-8k8x: 
CryoEM of LonC open pentamer, apo state

PDB-8k8y: 
CryoEM structure of LonC heptamer in presence of AGS

PDB-8k8z: 
CryoEM structure of LonC protease hexamer in presence of AGS

PDB-8k90: 
CryoEM structure of LonC protease open pentamer in presence of AGS

PDB-8k91: 
CryoEM structure of LonC S582A hepatmer with Lysozyme

PDB-8k92: 
CryoEM structure of LonC S582A hexamer with Lysozyme

PDB-8k93: 
CryoEM structure of LonC protease S582A open hexamer with lysozyme

PDB-8k94: 
CryoEM structure of LonC protease S582A open pentamer with lysozyme

PDB-8k95: 
CryoEM structure of LonC protease open Hexamer, AGS

PDB-8k96: 
CryoEM structure of LonC protease hepatmer with Bortezomib

PDB-8k97: 
CryoEM structure of LonC protease hexamer with Bortezomib

EMDB-39645: 
The structure of HKU1-B S protein with bsAb1
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
