-Search query
-Search result
Showing 1 - 50 of 533 items for (author: brown & z)

EMDB-16825: 
Structure of human terminal uridylyltransferase 7 (hTUT7/ZCCHC6)

EMDB-17084: 
Structure of human terminal uridylyltransferase 7 (hTUT7/ZCCHC6) bound with pre-let7g miRNA and UTPalphaS
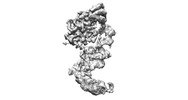
EMDB-17086: 
Human terminal uridylyltransferase 7 (TUT7/ZCCHC6) bound with pre-let7g miRNA and Lin28A - complex 1
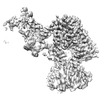
EMDB-17087: 
Human terminal uridylyltransferase 7 (TUT7/ZCCHC6) bound with pre-let7g miRNA and Lin28A - complex 2

PDB-8oef: 
Structure of human terminal uridylyltransferase 7 (hTUT7/ZCCHC6)

PDB-8opp: 
Structure of human terminal uridylyltransferase 7 (hTUT7/ZCCHC6) bound with pre-let7g miRNA and UTPalphaS

PDB-8ops: 
Human terminal uridylyltransferase 7 (TUT7/ZCCHC6) bound with pre-let7g miRNA and Lin28A - complex 1

PDB-8opt: 
Human terminal uridylyltransferase 7 (TUT7/ZCCHC6) bound with pre-let7g miRNA and Lin28A - complex 2
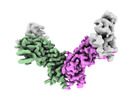
EMDB-44389: 
Cryo-EM structure of the ZBTB5 BTB domain filament
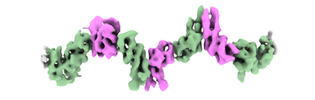
EMDB-44391: 
Cryo-EM structure of the ZBTB9 BTB domain filament

PDB-9b9r: 
Cryo-EM structure of the ZBTB5 BTB domain filament

PDB-9b9v: 
Cryo-EM structure of the ZBTB9 BTB domain filament

EMDB-43762: 
Aca2 from Pectobacterium phage ZF40 bound to RNA

PDB-8w35: 
Aca2 from Pectobacterium phage ZF40 bound to RNA
EMDB-17164: 
Structure of human terminal uridylyltransferase 4 (TUT4, ZCCHC11) in complex with pre-let7g miRNA and Lin28A

PDB-8ost: 
Structure of human terminal uridylyltransferase 4 (TUT4, ZCCHC11) in complex with pre-let7g miRNA and Lin28A

EMDB-17377: 
Structure of human SIT1 (focussed map / refinement)

EMDB-17378: 
Structure of human SIT1:ACE2 complex (open PD conformation)

EMDB-17379: 
Structure of human SIT1:ACE2 complex (closed PD conformation)

EMDB-17380: 
Structure of human SIT1 bound to L-pipecolate (focussed map / refinement)

EMDB-17381: 
Structure of human SIT1:ACE2 complex (open PD conformation) bound to L-pipecolate

EMDB-17382: 
Structure of human SIT1:ACE2 complex (closed PD conformation) bound to L-pipecolate

PDB-8p2w: 
Structure of human SIT1 (focussed map / refinement)

PDB-8p2x: 
Structure of human SIT1:ACE2 complex (open PD conformation)

PDB-8p2y: 
Structure of human SIT1:ACE2 complex (closed PD conformation)

PDB-8p2z: 
Structure of human SIT1 bound to L-pipecolate (focussed map / refinement)

PDB-8p30: 
Structure of human SIT1:ACE2 complex (open PD conformation) bound to L-pipecolate

PDB-8p31: 
Structure of human SIT1:ACE2 complex (closed PD conformation) bound to L-pipecolate

EMDB-17197: 
Human TPC2 in Complex with Antagonist (S)-SG-094

EMDB-19108: 
Human TPC2 in Complex withAntagonist (R)-SG-094

PDB-8ouo: 
Human TPC2 in Complex with Antagonist (S)-SG-094

EMDB-41152: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus SZ3 in Closed Conformation

PDB-8tc5: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus SZ3 in Closed Conformation

EMDB-41149: 
Cryo-EM Structure of Spike Glycoprotein from Bat Coronavirus WIV1 in Closed Conformation

EMDB-41150: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus 007 in Closed Conformation

PDB-8tc0: 
Cryo-EM Structure of Spike Glycoprotein from Bat Coronavirus WIV1 in Closed Conformation

PDB-8tc1: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus 007 in Closed Conformation

EMDB-43889: 
Chlamydomonas reinhardtii mastigoneme (constituent map 1)

EMDB-43890: 
Chlamydomonas reinhardtii mastigoneme (constituent map 2)

EMDB-43891: 
Chlamydomonas reinhardtii mastigoneme (constituent map 3)

EMDB-43892: 
Composite cryo-EM map of the Chlamydomonas reinhardtii mastigoneme

PDB-9b4h: 
Chlamydomonas reinhardtii mastigoneme filament

EMDB-41363: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5

PDB-8tl6: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5

EMDB-19067: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).

EMDB-19076: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).

EMDB-19077: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).

PDB-8rd8: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).

PDB-8rdv: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).

PDB-8rdw: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
