-Search query
-Search result
Showing all 34 items for (author: biggin & pc)

EMDB-44599: 
Cryo-EM structure of the mammalian peptide transporter PepT2 bound to cefadroxil

EMDB-44600: 
Cryo-EM structure of the mammalian peptide transporter PepT2 bound to amoxicillin

EMDB-44601: 
Cryo-EM structure of the mammalian peptide transporter PepT2 bound to cloxacillin, pose 1

EMDB-44602: 
Cryo-EM structure of the mammalian peptide transporter PepT2 bound to cloxacillin, pose 2

EMDB-35163: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 5.5

EMDB-35164: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in closed state
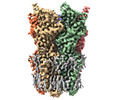
EMDB-36339: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 2.5
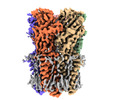
EMDB-37446: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in intermediate state
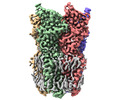
EMDB-37447: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in open state

EMDB-35161: 
Cryo-EM structure of nanodisc (asolectin) reconstituted GLIC at pH 7.5

EMDB-35162: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 7.5

EMDB-15985: 
Small molecule positive allosteric modulation of homomeric kainate receptors GluK1-3: Development of screening assays and insight into GluK3 structure

EMDB-15986: 
Ionotropic glutamate receptor K3 with glutamate and BPAM

EMDB-29019: 
Alpha1/BetaB Heteromeric Glycine Receptor in 1 mM Glycine 20 uM Ivermectin State

EMDB-26130: 
Alpha1/BetaB Heteromeric Glycine Receptor in Strychnine-Bound State
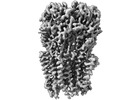
EMDB-26141: 
Alpha1/BetaB Heteromeric Glycine Receptor in Glycine-Bound State

EMDB-23700: 
Full length alpha1 Glycine receptor in presence of 32uM Tetrahydrocannabinol

EMDB-23701: 
Full length alpha1 Glycine receptor in presence of 0.1mM Glycine

EMDB-23702: 
Full length alpha1 Glycine receptor in presence of 0.1mM Glycine and 32uM Tetrahydrocannabinol

EMDB-23703: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine

EMDB-23704: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine and 32uM Tetrahydrocannabinol State 1

EMDB-23705: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine and 32uM Tetrahydrocannabinol State 2

EMDB-23706: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine and 32uM Tetrahydrocannabinol State 3

EMDB-24946: 
Glycine and glutamate bound GluN1a-GluN2B NMDA receptors in non-active 1 conformation at 2.97 Angstrom resolution

EMDB-24947: 
Phencyclidine-bound GluN1a-GluN2B NMDA receptors
EMDB-24948: 
S-(+)-ketamine bound GluN1a-GluN2B NMDA receptors at 3.69 Angstrom resolution

EMDB-24949: 
Memantine-bound GluN1a-GluN2B NMDA receptors

EMDB-13266: 
Cryo EM structure of System XC- in complex with glutamate

EMDB-13267: 
Cryo EM structure of System XC-

EMDB-12528: 
Cryo-EM structure of the mammalian peptide transporter PepT2

EMDB-12140: 
Cryo-EM structure of the outward open proton coupled folate transporter at pH 7.5

EMDB-12141: 
Cryo-EM structure of the proton coupled folate transporter at pH 6.0 bound to pemetrexed

EMDB-11064: 
Structure of canine Sec61 inhibited by mycolactone

EMDB-1661: 
The three-dimensional structure of a hepatitis C virus p7 ion channel by electron microscopy
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
