-Search query
-Search result
Showing all 37 items for (author: nazzari & af)

EMDB-29396: 
Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD
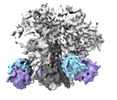
EMDB-29836: 
vFP52.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-29880: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 1)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29881: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 2)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29882: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 3)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29905: 
vFP48.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

PDB-8fr6: 
Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

PDB-8g85: 
vFP52.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

PDB-8g9w: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 1)
Method: single particle / : Changela A, Gorman J, Kwong PD

PDB-8g9x: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 2)
Method: single particle / : Changela A, Gorman J, Kwong PD

PDB-8g9y: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 3)
Method: single particle / : Changela A, Gorman J, Kwong PD

PDB-8gas: 
vFP48.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-27735: 
Cryo-EM structure of SIVmac239 SOS-2P Env trimer in complex with human bNAb PGT145
Method: single particle / : Gorman J, Kwong PD

PDB-8dvd: 
Cryo-EM structure of SIVmac239 SOS-2P Env trimer in complex with human bNAb PGT145
Method: single particle / : Gorman J, Kwong PD

EMDB-27718: 
SIV E660.CR54 SOS-2P Env Trimer with ITS92.02
Method: single particle / : Gorman J, Kwong PD

PDB-8dua: 
SIV E660.CR54 SOS-2P Env Trimer with ITS92.02
Method: single particle / : Gorman J, Kwong PD

EMDB-25062: 
In-situ structure of SIV trimer
Method: subtomogram averaging / : Gorman J, Wang CY, Mason RD, Nazzari AF, Welles H, Zhou TQ, Jr JB, Tsybovsky Y, Verardi R, Yang YP, Zhang BS, Lifson JD, Liu J, Roederer M, Kwong PD

EMDB-25063: 
in situ SIVmac239-Env trimers on the surface of AT-2-inactivated virions
Method: subtomogram averaging / : Liu J, Wang CY

EMDB-25064: 
in situ SIVmac239-Env trimers on the surface of AT-2-inactivated virions
Method: subtomogram averaging / : Liu J, Wang CY

EMDB-25065: 
In-situ structure of SIVmac239-Env trimers with ITS90.03 associated
Method: subtomogram averaging / : Liu J, Wang CY

EMDB-27631: 
SIV mac239 SOS-2P K169T Env trimer with bNAb PGT145 and jacalin
Method: single particle / : Gorman J, Kwong PD

EMDB-24128: 
Cryo-EM structure of broadly neutralizing V2-apex-targeting antibody J033 in complex with HIV-1 Env
Method: single particle / : Zhou T, Gao F

EMDB-24071: 
Cryo-EM structure of broadly neutralizing V2-apex-targeting antibody J038 in complex with HIV-1 Env
Method: single particle / : Zhou T, Gao F

EMDB-23098: 
Cryo-EM structure of the VRC316 clinical trial, vaccine-elicited, human antibody 316-310-1B11 in complex with an H2 CAN05 HA trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-23816: 
Cryo-EM structure of the VRC310 clinical trial, vaccine-elicited, human antibody 310-030-1D06 Fab in complex with an H1 NC99 HA trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-23982: 
Structure of freshly purified SARS-CoV-2 S2P spike at pH 7.4
Method: single particle / : Tsybovsky Y, Olia AS, Kwong PD

EMDB-23983: 
Structure of aged SARS-CoV-2 S2P spike at pH 7.4
Method: single particle / : Tsybovsky Y, Olia AS, Kwong PD

EMDB-23984: 
Structure of SARS-CoV-2 S2P spike at pH 7.4 refolded by low-pH treatment
Method: single particle / : Tsybovsky Y, Olia AS, Kwong PD

EMDB-23914: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-23915: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-23498: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky Y

EMDB-23499: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky T

EMDB-21961: 
Cryo-EM structure of the VRC315 clinical trial, vaccine-elicited, human antibody 1D12 in complex with an H7 SH13 HA trimer
Method: single particle / : Gorman J, Kwong PD

PDB-6wxl: 
Cryo-EM structure of the VRC315 clinical trial, vaccine-elicited, human antibody 1D12 in complex with an H7 SH13 HA trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-23016: 
Cryo-EM structure of prefusion SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab
Method: single particle / : Cerutti G, Shapiro L

EMDB-23039: 
Cryo-EM map of SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab (disrupted form)
Method: single particle / : Cerutti G, Shapiro L
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
