[English] 日本語
 Yorodumi
Yorodumi- EMDB-23914: Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23914 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain | |||||||||
 Map data Map data | Local refinement | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / receptor-binding domain / antibody / IMMUNE SYSTEM-VIRAL PROTEIN complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.71 Å | |||||||||
 Authors Authors | Zhou T / Tsybovsky T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants. Authors: Lingshu Wang / Tongqing Zhou / Yi Zhang / Eun Sung Yang / Chaim A Schramm / Wei Shi / Amarendra Pegu / Olamide K Oloniniyi / Amy R Henry / Samuel Darko / Sandeep R Narpala / Christian ...Authors: Lingshu Wang / Tongqing Zhou / Yi Zhang / Eun Sung Yang / Chaim A Schramm / Wei Shi / Amarendra Pegu / Olamide K Oloniniyi / Amy R Henry / Samuel Darko / Sandeep R Narpala / Christian Hatcher / David R Martinez / Yaroslav Tsybovsky / Emily Phung / Olubukola M Abiona / Avan Antia / Evan M Cale / Lauren A Chang / Misook Choe / Kizzmekia S Corbett / Rachel L Davis / Anthony T DiPiazza / Ingelise J Gordon / Sabrina Helmold Hait / Tandile Hermanus / Prudence Kgagudi / Farida Laboune / Kwanyee Leung / Tracy Liu / Rosemarie D Mason / Alexandra F Nazzari / Laura Novik / Sarah O'Connell / Sijy O'Dell / Adam S Olia / Stephen D Schmidt / Tyler Stephens / Christopher D Stringham / Chloe Adrienna Talana / I-Ting Teng / Danielle A Wagner / Alicia T Widge / Baoshan Zhang / Mario Roederer / Julie E Ledgerwood / Tracy J Ruckwardt / Martin R Gaudinski / Penny L Moore / Nicole A Doria-Rose / Ralph S Baric / Barney S Graham / Adrian B McDermott / Daniel C Douek / Peter D Kwong / John R Mascola / Nancy J Sullivan / John Misasi /   Abstract: The emergence of highly transmissible SARS-CoV-2 variants of concern (VOCs) that are resistant to therapeutic antibodies highlights the need for continuing discovery of broadly reactive antibodies. ...The emergence of highly transmissible SARS-CoV-2 variants of concern (VOCs) that are resistant to therapeutic antibodies highlights the need for continuing discovery of broadly reactive antibodies. We identified four receptor binding domain-targeting antibodies from three early-outbreak convalescent donors with potent neutralizing activity against 23 variants, including the B.1.1.7, B.1.351, P.1, B.1.429, B.1.526, and B.1.617 VOCs. Two antibodies are ultrapotent, with subnanomolar neutralization titers [half-maximal inhibitory concentration (IC) 0.3 to 11.1 nanograms per milliliter; IC 1.5 to 34.5 nanograms per milliliter). We define the structural and functional determinants of binding for all four VOC-targeting antibodies and show that combinations of two antibodies decrease the in vitro generation of escape mutants, suggesting their potential in mitigating resistance development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23914.map.gz emd_23914.map.gz | 266 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23914-v30.xml emd-23914-v30.xml emd-23914.xml emd-23914.xml | 23.5 KB 23.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23914.png emd_23914.png | 92.2 KB | ||
| Masks |  emd_23914_msk_1.map emd_23914_msk_1.map | 536.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-23914.cif.gz emd-23914.cif.gz | 7 KB | ||
| Others |  emd_23914_additional_1.map.gz emd_23914_additional_1.map.gz emd_23914_half_map_1.map.gz emd_23914_half_map_1.map.gz emd_23914_half_map_2.map.gz emd_23914_half_map_2.map.gz | 505.9 MB 498.4 MB 498.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23914 http://ftp.pdbj.org/pub/emdb/structures/EMD-23914 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23914 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23914 | HTTPS FTP |
-Related structure data
| Related structure data |  7mlzMC  7lrsC  7lrtC  7mm0C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23914.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23914.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.873 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
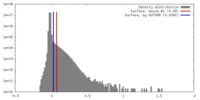
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23914_msk_1.map emd_23914_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Local refinement
| File | emd_23914_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
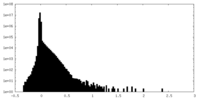
| Density Histograms |
-Half map: Local refinement
| File | emd_23914_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
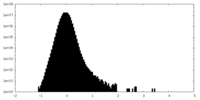
| Density Histograms |
-Half map: Local refinement
| File | emd_23914_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
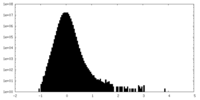
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 spike RBD in complex with neutralizing antibody B1-182.1
| Entire | Name: SARS-CoV-2 spike RBD in complex with neutralizing antibody B1-182.1 |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 spike RBD in complex with neutralizing antibody B1-182.1
| Supramolecule | Name: SARS-CoV-2 spike RBD in complex with neutralizing antibody B1-182.1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Local refinement of data collected for the complex made by mixing the SARS-CoV-2 spike protein and Fab fragment of antibody at a molar ratio of 1:3.6. Used particle substraction to obtain ...Details: Local refinement of data collected for the complex made by mixing the SARS-CoV-2 spike protein and Fab fragment of antibody at a molar ratio of 1:3.6. Used particle substraction to obtain RBD-Fab portion for refinement. |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 539.951 KDa |
-Macromolecule #1: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.100756 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NITNLCPFGE VFNATRFASV YAWNRKRISN CVADYSVLYN SASFSTFKCY GVSPTKLNDL CFTNVYADSF VIRGDEVRQI APGQTGKIA DYNYKLPDDF TGCVIAWNSN NLDSKVGGNY NYLYRLFRKS NLKPFERDIS TEIYQAGSTP CNGVEGFNCY F PLQSYGFQ ...String: NITNLCPFGE VFNATRFASV YAWNRKRISN CVADYSVLYN SASFSTFKCY GVSPTKLNDL CFTNVYADSF VIRGDEVRQI APGQTGKIA DYNYKLPDDF TGCVIAWNSN NLDSKVGGNY NYLYRLFRKS NLKPFERDIS TEIYQAGSTP CNGVEGFNCY F PLQSYGFQ PTNGVGYQPY RVVVLSFELL HAPATVCGP UniProtKB: Spike glycoprotein |
-Macromolecule #2: B1-182.1 Fab heavy chain
| Macromolecule | Name: B1-182.1 Fab heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.285312 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QMQLVQSGPE VKKPGTSVKV SCKASGFTFT SSAVQWVRQA RGQRLEWIGW IVVGSGNTNY AQKFQERVTI TRDMSTSTAY MELSSLRSE DTAVYYCAAP YCSGGSCFDG FDIWGQGTMV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA ...String: QMQLVQSGPE VKKPGTSVKV SCKASGFTFT SSAVQWVRQA RGQRLEWIGW IVVGSGNTNY AQKFQERVTI TRDMSTSTAY MELSSLRSE DTAVYYCAAP YCSGGSCFDG FDIWGQGTMV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NHKPSNTKVD KKVEPKSCDK |
-Macromolecule #3: B1-182.1 Fab light chain
| Macromolecule | Name: B1-182.1 Fab light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.536008 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EIVLTQSPGT LSLSPGERAT LSCRASQSVS SSYLAWYQQK PGQAPRLLIY GASSRATGFP DRFSGSGSGT DFTLTISRLE PEDFAVYYC QQYGNSPWTF GQGTKVEIRR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: EIVLTQSPGT LSLSPGERAT LSCRASQSVS SSYLAWYQQK PGQAPRLLIY GASSRATGFP DRFSGSGSGT DFTLTISRLE PEDFAVYYC QQYGNSPWTF GQGTKVEIRR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 100 mM HEPES, 150 mM NaCl |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Blot for 4 seconds before plugging.. |
| Details | Complex at 0.5 mg/mL concentration in the presence of 0.005% DDM |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: OTHER / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.1 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: Details: SARS-CoV-2 receptor-binding domain in complex with antibody A23-58.1 |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.71 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2.15) Details: Local refinement with particles subtracted from SARS-CoV-2 spike-antibody complex. Number images used: 208776 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)