-Search query
-Search result
Showing all 39 items for (author: caffrey & m)
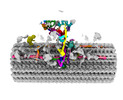
EMDB-41284: 
Combined linker domain of N-DRC and associated proteins Tetrahymena
Method: single particle / : Ghanaeian AG, Majhi SM, McCaffrey CM, Nami BN, Black CB, Yang SK, Legal TL, Papoulas OP, Janowska MJ, Valente-Paterno MV, Marcotte EM, Wloga DW, Bui KH

PDB-8tid: 
Combined linker domain of N-DRC and associated proteins Tetrahymena
Method: single particle / : Ghanaeian AG, Majhi SM, McCaffrey CM, Nami BN, Black CB, Yang SK, Legal TL, Papoulas OP, Janowska MJ, Valente-Paterno MV, Marcotte EM, Wloga DW, Bui KH

EMDB-41270: 
Focused refinement of the N-DRC from the Tetrahymena WT subtomo
Method: subtomogram averaging / : Ghanaeian AG, Majhi SM, McCaffrey CM, Nami BN, Black CB, Yang SK, Legal TL, Papoulas OP, Janowska MJ, Valente-Paterno MV, Marcotte EM, Wloga DW, Bui KH

EMDB-15786: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15787: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15788: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE (C387S mutant)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15789: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15790: 
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15791: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0k: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0l: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0m: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE (C387S mutant)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0n: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0o: 
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0p: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-24518: 
Cryo-EM structure of human p97-R155H mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

EMDB-24519: 
Cryo-EM structure of human p97-R155H mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

EMDB-24522: 
Cryo-EM structure of human p97-R191Q mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

EMDB-24523: 
Cryo-EM structure of human p97-R191Q mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

EMDB-24524: 
Cryo-EM structure of human p97-A232E mutant bound to ADP
Method: single particle / : Caffrey B, Zhu X

EMDB-24525: 
Cryo-EM structure of human p97-A232E mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X

EMDB-24526: 
Cryo-EM structure of human p97-E470D mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X

EMDB-24528: 
Cryo-EM structure of human p97-E470D mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X

EMDB-24529: 
Cryo-EM structure of human p97-D592N mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X

EMDB-24530: 
Cryo-EM structure of human p97-D592N mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X

EMDB-24531: 
Cryo-EM structure of human p97 bound to CB-5083 and ADP.
Method: single particle / : Caffrey B, Zhu X

EMDB-24532: 
Cryo-EM structure of human p97 bound to CB-5083 and ATPgS.
Method: single particle / : Caffrey B, Zhu X

PDB-7rl6: 
Cryo-EM structure of human p97-R155H mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rl7: 
Cryo-EM structure of human p97-R155H mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rl9: 
Cryo-EM structure of human p97-R191Q mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rla: 
Cryo-EM structure of human p97-R191Q mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rlb: 
Cryo-EM structure of human p97-A232E mutant bound to ADP
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rlc: 
Cryo-EM structure of human p97-A232E mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rld: 
Cryo-EM structure of human p97-E470D mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rlf: 
Cryo-EM structure of human p97-E470D mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rlg: 
Cryo-EM structure of human p97-D592N mutant bound to ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rlh: 
Cryo-EM structure of human p97-D592N mutant bound to ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rli: 
Cryo-EM structure of human p97 bound to CB-5083 and ADP.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S

PDB-7rlj: 
Cryo-EM structure of human p97 bound to CB-5083 and ATPgS.
Method: single particle / : Caffrey B, Zhu X, Berezuk A, Tuttle K, Chittori S, Subramaniam S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
