+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-6774 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of human respiratory complex III (cytochrome bc1 complex) | |||||||||
 マップデータ マップデータ | This map was obtained by sub-region refinement. | |||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報Complex III assembly / Complex III assembly / Complex III assembly / Complex III assembly / Complex III assembly / response to D-galactosamine / Complex III assembly / response to cobalamin / Complex III assembly / Complex III assembly ...Complex III assembly / Complex III assembly / Complex III assembly / Complex III assembly / Complex III assembly / response to D-galactosamine / Complex III assembly / response to cobalamin / Complex III assembly / Complex III assembly / Complex III assembly / response to mercury ion / Respiratory electron transport / subthalamus development / pons development / cerebellar Purkinje cell layer development / mitochondrial respiratory chain complex III assembly / pyramidal neuron development / thalamus development / response to alkaloid / Mitochondrial protein import / Mitochondrial translation termination / respiratory chain complex / respiratory chain complex III / cellular respiration / oxidative phosphorylation / response to glucagon / quinol-cytochrome-c reductase / quinol-cytochrome-c reductase activity / mitochondrial electron transport, ubiquinol to cytochrome c / response to copper ion / hypothalamus development / midbrain development / electron transport coupled proton transport / animal organ regeneration / response to hyperoxia / response to cadmium ion / Mitochondrial protein degradation / aerobic respiration / response to activity / respiratory electron transport chain / hippocampus development / generation of precursor metabolites and energy / metalloendopeptidase activity / response to calcium ion / 2 iron, 2 sulfur cluster binding / response to toxic substance / response to ethanol / response to hypoxia / electron transfer activity / oxidoreductase activity / mitochondrial inner membrane / response to xenobiotic stimulus / mitochondrial matrix / heme binding / ubiquitin protein ligase binding / protein-containing complex binding / mitochondrion / proteolysis / nucleoplasm / membrane / metal ion binding / nucleus 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.4 Å | |||||||||
 データ登録者 データ登録者 | Gu J / Wu M / Yang M | |||||||||
 引用 引用 |  ジャーナル: Cell / 年: 2017 ジャーナル: Cell / 年: 2017タイトル: Architecture of Human Mitochondrial Respiratory Megacomplex IIIIIV. 著者: Runyu Guo / Shuai Zong / Meng Wu / Jinke Gu / Maojun Yang /  要旨: The respiratory megacomplex represents the highest-order assembly of respiratory chain complexes, and it allows mitochondria to respond to energy-requiring conditions. To understand its architecture, ...The respiratory megacomplex represents the highest-order assembly of respiratory chain complexes, and it allows mitochondria to respond to energy-requiring conditions. To understand its architecture, we examined the human respiratory chain megacomplex-IIIIIV (MCIIIIIV) with 140 subunits and a subset of associated cofactors using cryo-electron microscopy. The MCIIIIIV forms a circular structure with the dimeric CIII located in the center, where it is surrounded by two copies each of CI and CIV. Two cytochrome c (Cyt.c) molecules are positioned to accept electrons on the surface of the c state CIII dimer. Analyses indicate that CII could insert into the gaps between CI and CIV to form a closed ring, which we termed the electron transport chain supercomplex. The structure not only reveals the precise assignment of individual subunits of human CI and CIII, but also enables future in-depth analysis of the electron transport chain as a whole. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_6774.map.gz emd_6774.map.gz | 22.1 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-6774-v30.xml emd-6774-v30.xml emd-6774.xml emd-6774.xml | 10 KB 10 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_6774.png emd_6774.png | 18.2 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6774 http://ftp.pdbj.org/pub/emdb/structures/EMD-6774 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6774 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6774 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  5xteMC  6771C  6772C  6773C  6775C  6776C  5xtbC  5xtcC  5xtdC  5xthC  5xtiC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_6774.map.gz / 形式: CCP4 / 大きさ: 421.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_6774.map.gz / 形式: CCP4 / 大きさ: 421.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | This map was obtained by sub-region refinement. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.083 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
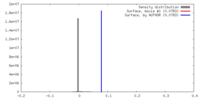
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : Human respiratory complex III
| 全体 | 名称: Human respiratory complex III |
|---|---|
| 要素 |
|
-超分子 #1: Human respiratory complex III
| 超分子 | 名称: Human respiratory complex III / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#11 / 詳細: Human cytochrome bc1 complex |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 実験値: 500 KDa |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON II (4k x 4k) 平均電子線量: 1.25 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: OTHER / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| CTF補正 | ソフトウェア - 名称: CTFFIND (ver. 3.0) |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: C1 (非対称) / 解像度のタイプ: BY AUTHOR / 解像度: 3.4 Å / 解像度の算出法: FSC 0.143 CUT-OFF / ソフトウェア - 名称: RELION (ver. 1.4) / 使用した粒子像数: 167761 |
| 初期 角度割当 | タイプ: RANDOM ASSIGNMENT / ソフトウェア - 名称: RELION (ver. 1.4) |
| 最終 角度割当 | タイプ: RANDOM ASSIGNMENT / ソフトウェア - 名称: RELION (ver. 1.4) |
 ムービー
ムービー コントローラー
コントローラー




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)