[English] 日本語
 Yorodumi
Yorodumi- EMDB-23940: Cryo-EM structure of the human SSU processome, state post-A1 - ra... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23940 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human SSU processome, state post-A1 - raw maps | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationmRNA N-acetyltransferase activity / preribosome / U6 snRNA 2'-O-ribose methyltransferase activity / oocyte growth / nucleologenesis / leucine zipper domain binding / snoRNA localization / granular component / rRNA acetylation involved in maturation of SSU-rRNA / rRNA cytidine N-acetyltransferase activity ...mRNA N-acetyltransferase activity / preribosome / U6 snRNA 2'-O-ribose methyltransferase activity / oocyte growth / nucleologenesis / leucine zipper domain binding / snoRNA localization / granular component / rRNA acetylation involved in maturation of SSU-rRNA / rRNA cytidine N-acetyltransferase activity / tRNA N-acetyltransferase activity / tRNA acetylation / U4atac snRNP / regulation of stem cell population maintenance / CURI complex / UTP-C complex / negative regulation of amyloid precursor protein biosynthetic process / t-UTP complex / Mpp10 complex / Pwp2p-containing subcomplex of 90S preribosome / U4atac snRNA binding / rRNA (pseudouridine) methyltransferase activity / rRNA modification / histone H2AQ104 methyltransferase activity / pre-snoRNP complex / box C/D sno(s)RNA binding / endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / dense fibrillar component / regulation of centrosome duplication / box C/D sno(s)RNA 3'-end processing / endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA methyltransferase activity / histone methyltransferase binding / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / regulation of transcription elongation by RNA polymerase II / positive regulation of rRNA processing / embryonic cleavage / tRNA export from nucleus / epigenetic programming in the zygotic pronuclei / cilium disassembly / spindle assembly involved in female meiosis / transcription elongation factor activity / rRNA base methylation / rRNA primary transcript binding / blastocyst formation / Cul4-RING E3 ubiquitin ligase complex / protein localization to nucleolus / N-acetyltransferase activity / RNA splicing, via transesterification reactions / U4 snRNA binding / negative regulation of RNA splicing / sno(s)RNA-containing ribonucleoprotein complex / box C/D methylation guide snoRNP complex / SUMOylation of RNA binding proteins / telomerase holoenzyme complex / rRNA methylation / U2-type precatalytic spliceosome / neural crest cell differentiation / box C/D snoRNP assembly / rRNA modification in the nucleus and cytosol / Formation of the ternary complex, and subsequently, the 43S complex / erythrocyte homeostasis / cytoplasmic side of rough endoplasmic reticulum membrane / U3 snoRNA binding / Cajal body / negative regulation of ubiquitin protein ligase activity / Ribosomal scanning and start codon recognition / preribosome, small subunit precursor / Translation initiation complex formation / snoRNA binding / intercellular bridge / precatalytic spliceosome / mammalian oogenesis stage / NRAGE signals death through JNK / protein acetylation / activation-induced cell death of T cells / positive regulation of transcription by RNA polymerase I / Protein hydroxylation / RNA polymerase II complex binding / regulation of mitotic cell cycle / Association of TriC/CCT with target proteins during biosynthesis / mTORC1-mediated signalling / SARS-CoV-1 modulates host translation machinery / Peptide chain elongation / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / Selenocysteine synthesis / TFIID-class transcription factor complex binding / Formation of a pool of free 40S subunits / negative regulation of telomere maintenance via telomerase / Eukaryotic Translation Termination / blastocyst development / ubiquitin ligase inhibitor activity / negative regulation of apoptotic signaling pathway / decidualization / Response of EIF2AK4 (GCN2) to amino acid deficiency / SRP-dependent cotranslational protein targeting to membrane / chromosome, centromeric region / Viral mRNA Translation / condensed nuclear chromosome / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
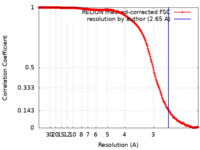
| Method | single particle reconstruction / cryo EM / Resolution: 2.65 Å | |||||||||
 Authors Authors | Vanden Broeck A / Singh S / Klinge S | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Nucleolar maturation of the human small subunit processome. Authors: Sameer Singh / Arnaud Vanden Broeck / Linamarie Miller / Malik Chaker-Margot / Sebastian Klinge /  Abstract: The human small subunit processome mediates early maturation of the small ribosomal subunit by coupling RNA folding to subsequent RNA cleavage and processing steps. We report the high-resolution ...The human small subunit processome mediates early maturation of the small ribosomal subunit by coupling RNA folding to subsequent RNA cleavage and processing steps. We report the high-resolution cryo–electron microscopy structures of maturing human small subunit (SSU) processomes at resolutions of 2.7 to 3.9 angstroms. These structures reveal the molecular mechanisms that enable crucial progressions during SSU processome maturation. RNA folding states within these particles are communicated to and coordinated with key enzymes that drive irreversible steps such as targeted exosome-mediated RNA degradation, protein-guided site-specific endonucleolytic RNA cleavage, and tightly controlled RNA unwinding. These conserved mechanisms highlight the SSU processome’s impressive structural plasticity, which endows this 4.5-megadalton nucleolar assembly with the distinctive ability to mature the small ribosomal subunit from within. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23940.map.gz emd_23940.map.gz | 66.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23940-v30.xml emd-23940-v30.xml emd-23940.xml emd-23940.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23940_fsc.xml emd_23940_fsc.xml | 19.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_23940.png emd_23940.png | 169.4 KB | ||
| Masks |  emd_23940_msk_1.map emd_23940_msk_1.map | 669.9 MB |  Mask map Mask map | |
| Others |  emd_23940_additional_1.map.gz emd_23940_additional_1.map.gz emd_23940_half_map_1.map.gz emd_23940_half_map_1.map.gz emd_23940_half_map_2.map.gz emd_23940_half_map_2.map.gz | 627 MB 542 MB 541.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23940 http://ftp.pdbj.org/pub/emdb/structures/EMD-23940 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23940 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23940 | HTTPS FTP |
-Validation report
| Summary document |  emd_23940_validation.pdf.gz emd_23940_validation.pdf.gz | 457.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23940_full_validation.pdf.gz emd_23940_full_validation.pdf.gz | 457.1 KB | Display | |
| Data in XML |  emd_23940_validation.xml.gz emd_23940_validation.xml.gz | 27.4 KB | Display | |
| Data in CIF |  emd_23940_validation.cif.gz emd_23940_validation.cif.gz | 36.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23940 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23940 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23940 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23940 | HTTPS FTP |
-Related structure data
| Related structure data |  7mq8C  7mq9C  7mqaC  7mqjC C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10781 (Title: Nucleolar maturation of the human small subunit processome EMPIAR-10781 (Title: Nucleolar maturation of the human small subunit processomeData size: 74.6 TB Data #1: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 1 [micrographs - multiframe] Data #2: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 2 [micrographs - multiframe] Data #3: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 3 [micrographs - multiframe] Data #4: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 4 [micrographs - multiframe] Data #5: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 5 [micrographs - multiframe] Data #6: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 6 [micrographs - multiframe] Data #7: Aligned and averaged micrographs of human SSU processomes - Dataset 1 [micrographs - single frame] Data #8: Aligned and averaged micrographs of human SSU processomes - Dataset 2 [micrographs - single frame] Data #9: Aligned and averaged micrographs of human SSU processomes - Dataset 3 [micrographs - single frame] Data #10: Aligned and averaged micrographs of human SSU processomes - Dataset 4 [micrographs - single frame] Data #11: Aligned and averaged micrographs of human SSU processomes - Dataset 5 [micrographs - single frame] Data #12: Aligned and averaged micrographs of human SSU processomes - Dataset 6 [micrographs - single frame]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23940.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23940.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23940_msk_1.map emd_23940_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
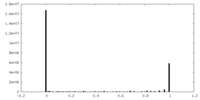
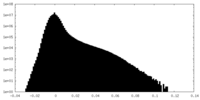
| Density Histograms |
-Additional map: #1
| File | emd_23940_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
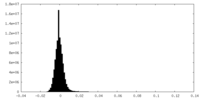
| Density Histograms |
-Half map: Half-map1
| File | emd_23940_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
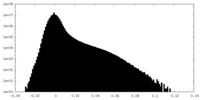
| Density Histograms |
-Half map: Half-map2
| File | emd_23940_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human SSU processome, state post-A1
| Entire | Name: Human SSU processome, state post-A1 |
|---|---|
| Components |
|
-Supramolecule #1: Human SSU processome, state post-A1
| Supramolecule | Name: Human SSU processome, state post-A1 / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 5 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 3.0 nm / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 84904 / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.7000000000000001 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|
 Movie
Movie Controller
Controller








































































 Z
Z Y
Y X
X