1T3I
 
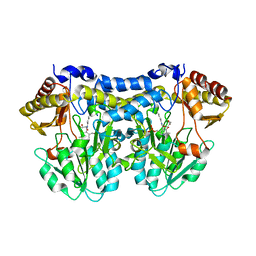 | | Structure of slr0077/SufS, the Essential Cysteine Desulfurase from Synechocystis PCC 6803 | | Descriptor: | GLYCEROL, PYRIDOXAL-5'-PHOSPHATE, Probable cysteine desulfurase, ... | | Authors: | Tirupati, B, Vey, J.L, Drennan, C.L, Bollinger Jr, J.M. | | Deposit date: | 2004-04-26 | | Release date: | 2004-09-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Kinetic and structural characterization of Slr0077/SufS, the essential cysteine desulfurase from Synechocystis sp. PCC 6803.
Biochemistry, 43, 2004
|
|
5KCN
 
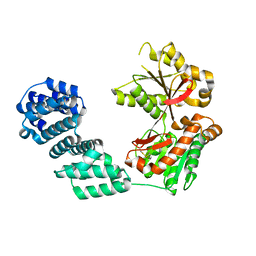 | |
4P29
 
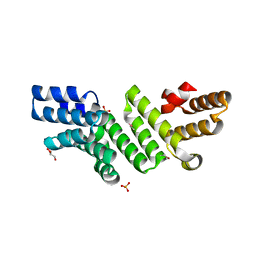 | |
5VAT
 
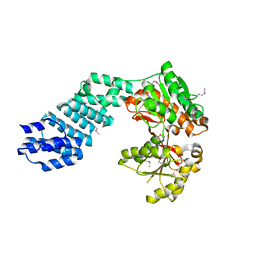 | |
5VBG
 
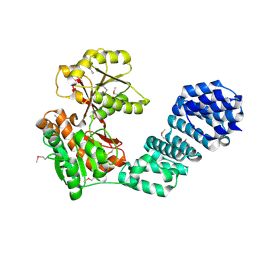 | |
3CKM
 
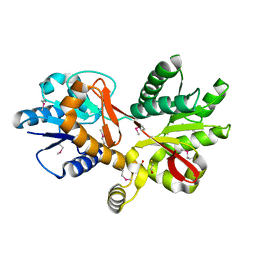 | |
6DR3
 
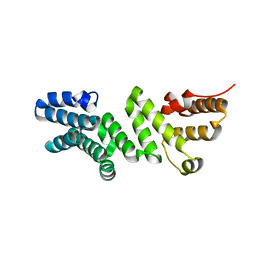 | | Crystal structure of E. coli LpoA amino terminal domain | | Descriptor: | Penicillin-binding protein activator LpoA | | Authors: | Kelley, A.C, Saper, M.A. | | Deposit date: | 2018-06-11 | | Release date: | 2019-05-08 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Crystal structures of the amino-terminal domain of LpoA from Escherichia coli and Haemophilus influenzae.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
6DCJ
 
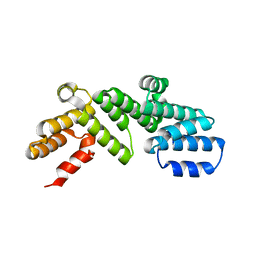 | |
