1MHN
 
 | |
8R1O
 
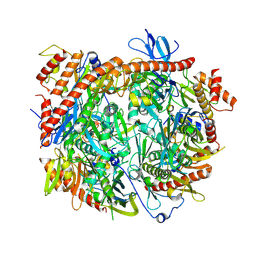 | | Structure of C. thermophilum RNA exosome core | | Descriptor: | Exoribonuclease phosphorolytic domain-containing protein, Exoribonuclease-like protein, Exosome complex component MTR3, ... | | Authors: | Lazzaretti, D, Liebau, J, Pilsl, M, Sprangers, R. | | Deposit date: | 2023-11-02 | | Release date: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.19 Å) | | Cite: | Beyond static structures: quantitative dynamics in the eukaryotic RNA exosome complex
Biorxiv, 2024
|
|
7PVM
 
 | |
1QMC
 
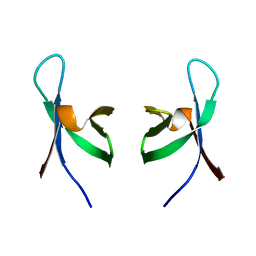 | | C-terminal DNA-binding domain of HIV-1 integrase, NMR, 42 structures | | Descriptor: | HIV-1 INTEGRASE | | Authors: | Eijkelenboom, A.P.A.M, Sprangers, R, Hard, K, Puras Lutzke, R.A, Plasterk, R.H.A, Boelens, R, Kaptein, R. | | Deposit date: | 1999-09-27 | | Release date: | 1999-12-14 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Refined Solution Structure of the C-Terminal DNA-Binding Domain of Human Immunovirus-1 Integrase.
Proteins: Struct.,Funct., Genet., 36, 1999
|
|
7OPK
 
 | |
1Y7O
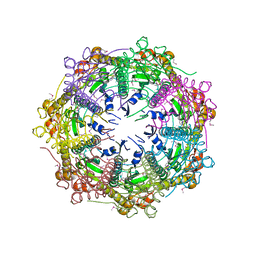 
 | | The structure of Streptococcus pneumoniae A153P ClpP | | Descriptor: | ATP-dependent Clp protease proteolytic subunit, CALCIUM ION | | Authors: | Kimber, M.S, Gribun, A, Ching, R, Sprangers, R, Fiebig, K.M, Houry, W.A. | | Deposit date: | 2004-12-09 | | Release date: | 2005-02-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | The ClpP double ring tetradecameric protease exhibits plastic ring-ring interactions, and the N termini of its subunits form flexible loops that are essential for ClpXP and ClpAP complex formation.
J.Biol.Chem., 280, 2005
|
|
5JP4
 
 | |
5N2V
 
 | |
8PNK
 
 | |
8PEL
 
 | |
5BV3
 
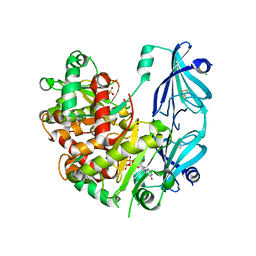 | | Yeast Scavenger Decapping Enzyme in complex with m7GDP | | Descriptor: | 7N-METHYL-8-HYDROGUANOSINE-5'-DIPHOSPHATE, CHLORIDE ION, SULFATE ION, ... | | Authors: | Neu, A, Neu, U, Sprangers, R. | | Deposit date: | 2015-06-04 | | Release date: | 2015-08-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | An excess of catalytically required motions inhibits the scavenger decapping enzyme.
Nat.Chem.Biol., 11, 2015
|
|
4Q2S
 
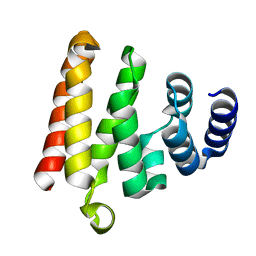 | |
7BBB
 
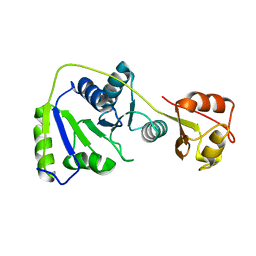 | |
6TRQ
 
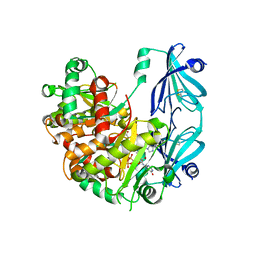 | | S.c. Scavenger Decapping Enzyme DcpS in complex with the capped RNA dinucleotide m7G-GU | | Descriptor: | 7N-METHYL-8-HYDROGUANOSINE-5'-DIPHOSPHATE, L-GUANOSINE-5'-MONOPHOSPHATE, PHOSPHONATE, ... | | Authors: | Fuchs, A.-L, Neu, A, Sprangers, R. | | Deposit date: | 2019-12-19 | | Release date: | 2020-07-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.944 Å) | | Cite: | Molecular basis of the selective processing of short mRNA substrates by the DcpS mRNA decapping enzyme.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6GBS
 
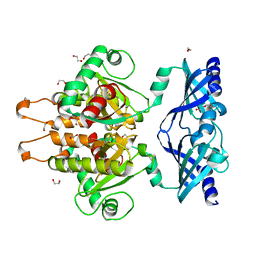 | | Crystal Structure of the C. themophilum Scavenger Decapping Enzyme DcpS apo form | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, Putative mRNA decapping protein | | Authors: | Fuchs, A.-L, Neu, A, Sprangers, R. | | Deposit date: | 2018-04-16 | | Release date: | 2019-04-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.946 Å) | | Cite: | Molecular basis of the selective processing of short mRNA substrates by the DcpS mRNA decapping enzyme.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
1K1G
 
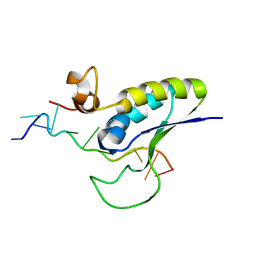 | | STRUCTURAL BASIS FOR RECOGNITION OF THE INTRON BRANCH SITE RNA BY SPLICING FACTOR 1 | | Descriptor: | 5'-R(*UP*AP*UP*AP*CP*UP*AP*AP*CP*AP*A)-3', SF1-Bo isoform | | Authors: | Liu, Z, Luyten, I, Bottomley, M.J, Messias, A.C, Houngninou-Molango, S, Sprangers, R, Zanier, K, Kramer, A, Sattler, M. | | Deposit date: | 2001-09-25 | | Release date: | 2001-11-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for recognition of the intron branch site RNA by splicing factor 1.
Science, 294, 2001
|
|
3SWN
 
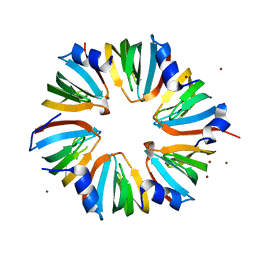 | | Structure of the LSm657 Complex: An Assembly Intermediate of the LSm1 7 and LSm2 8 Rings | | Descriptor: | U6 snRNA-associated Sm-like protein LSm5, U6 snRNA-associated Sm-like protein LSm6, U6 snRNA-associated Sm-like protein LSm7, ... | | Authors: | Mund, M, Neu, A, Ullmann, J.L, Neu, U, Sprangers, R. | | Deposit date: | 2011-07-14 | | Release date: | 2011-10-26 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the LSm657 complex: an assembly intermediate of the LSm1-7 and LSm2-8 rings.
J.Mol.Biol., 414, 2011
|
|
1G5V
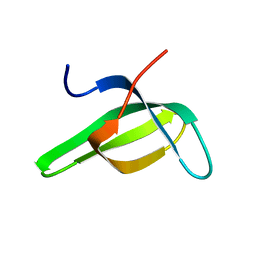 
 | | SOLUTION STRUCTURE OF THE TUDOR DOMAIN OF THE HUMAN SMN PROTEIN | | Descriptor: | SURVIVAL MOTOR NEURON PROTEIN 1 | | Authors: | Selenko, P, Sprangers, R, Stier, G, Buehler, D, Fischer, U, Sattler, M. | | Deposit date: | 2000-11-02 | | Release date: | 2001-05-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | SMN tudor domain structure and its interaction with the Sm proteins.
Nat.Struct.Biol., 8, 2001
|
|
4A54
 
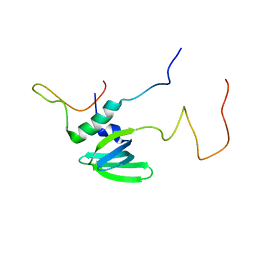 | | Structural basis of the Dcp1:Dcp2 mRNA decapping complex activation by Edc3 and Scd6 | | Descriptor: | EDC3, MRNA DECAPPING COMPLEX SUBUNIT 2 | | Authors: | Fromm, S.A, Truffault, V, Kamenz, J, Braun, J.E, Hoffmann, N.A, Izaurralde, E, Sprangers, R. | | Deposit date: | 2011-10-24 | | Release date: | 2012-02-01 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The Structural Basis of Edc3- and Scd6-Mediated Activation of the Dcp1:Dcp2 Mrna Decapping Complex.
Embo J., 31, 2011
|
|
4A53
 
 | | Structural basis of the Dcp1:Dcp2 mRNA decapping complex activation by Edc3 and Scd6 | | Descriptor: | EDC3 | | Authors: | Fromm, S.A, Truffault, V, Kamenz, J, Braun, J.E, Hoffmann, N.A, Izaurralde, E, Sprangers, R. | | Deposit date: | 2011-10-24 | | Release date: | 2012-02-01 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The Structural Basis of Edc3- and Scd6-Mediated Activation of the Dcp1:Dcp2 Mrna Decapping Complex.
Embo J., 31, 2011
|
|
2KU2
 
 | |
2KU1
 
 | |
1O0P
 
 | | Solution Structure of the third RNA Recognition Motif (RRM) of U2AF65 in complex with an N-terminal SF1 peptide | | Descriptor: | Splicing Factor SF1, Splicing factor U2AF 65 kDa subunit | | Authors: | Selenko, P, Gregorovic, G, Sprangers, R, Stier, G, Rhani, Z, Kramer, A, Sattler, M. | | Deposit date: | 2003-02-24 | | Release date: | 2003-05-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP
Mol.Cell, 11, 2003
|
|
1OPI
 
 | | SOLUTION STRUCTURE OF THE THIRD RNA RECOGNITION MOTIF (RRM) OF U2AF65 IN COMPLEX WITH AN N-TERMINAL SF1 PEPTIDE | | Descriptor: | SPLICING FACTOR SF1, SPLICING FACTOR U2AF 65 KDA SUBUNIT | | Authors: | Selenko, P, Gregorovic, G, Sprangers, R, Stier, G, Rhani, Z, Kramer, A, Sattler, M. | | Deposit date: | 2003-03-05 | | Release date: | 2004-03-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP
Mol.Cell, 11, 2003
|
|
7AC8
 
 | |
