9P24
 
 | | Structure of human cardiac sodium channel Nav1.5 in intermediate open state | | Descriptor: | (3beta,14beta,17beta,25R)-3-[4-methoxy-3-(methoxymethyl)butoxy]spirost-5-en, 2-acetamido-2-deoxy-beta-D-glucopyranose, SODIUM ION, ... | | Authors: | Biswas, R, Chinthalapudi, K. | | Deposit date: | 2025-06-11 | | Release date: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (3.48 Å) | | Cite: | Structural and functional mechanisms underlying activation gate dynamics and IFM motif accessibility in human Na v 1.5.
Nat Commun, 17, 2026
|
|
377D
 
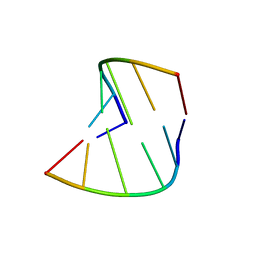 | | 5'-R(*CP*GP*UP*AP*CP*DG)-3' | | Descriptor: | RNA-DNA (5'-R(*CP*GP*UP*AP*CP*DG)-3') | | Authors: | Biswas, R, Mitra, S.N, Sundaralingam, M. | | Deposit date: | 1998-01-26 | | Release date: | 1998-08-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | 1.76 A structure of a pyrimidine start alternating A-RNA hexamer r(CGUAC)dG.
Acta Crystallogr.,Sect.D, 54, 1998
|
|
332D
 
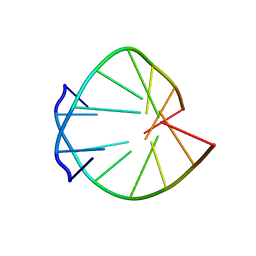 | |
315D
 
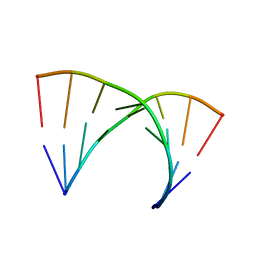 | |
4OOB
 
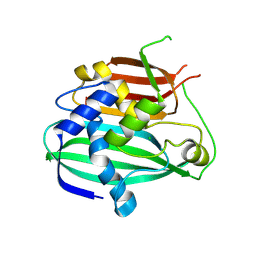 | | Crystal structure of HtdX(Rv0241c) from Mycobacterium tuberculosis | | Descriptor: | CHLORIDE ION, Uncharacterized protein HtdX | | Authors: | Biswas, R, Dutta, D, Das, A.K. | | Deposit date: | 2014-01-31 | | Release date: | 2015-02-04 | | Last modified: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure and molecular dynamics simulation of Mycobacterium tuberculosis MaoC-like dehydratase HtdX provide insights into substrate binding and membrane interactions.
Biochim Biophys Acta Proteins Proteom, 1873, 2025
|
|
4RV2
 
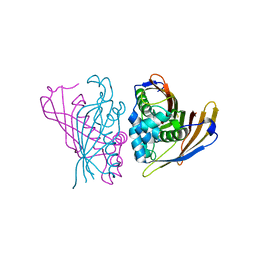 | | Crystal Structure of (3R)-hydroxyacyl-ACP dehydratase HadAB hetero-dimer from Mycobacterium smegmatis | | Descriptor: | MaoC family protein, SULFATE ION, UPF0336 protein MSMEG_1340/MSMEI_1302 | | Authors: | Biswas, R, Hazra, D, Dutta, D, Das, A.K. | | Deposit date: | 2014-11-24 | | Release date: | 2015-02-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of dehydratase component HadAB complex of mycobacterial FAS-II pathway.
Biochem.Biophys.Res.Commun., 458, 2015
|
|
3WEW
 
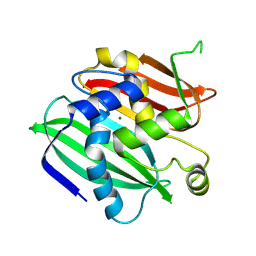 | |
8VYK
 
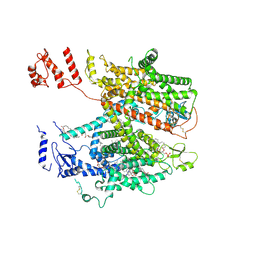 | | Structure of full-length human cardiac sodium channel - Class-II. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Biswas, R, Chinthalapudi, K. | | Deposit date: | 2024-02-08 | | Release date: | 2025-02-12 | | Last modified: | 2026-02-25 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Structural basis of human Na v 1.5 gating mechanisms.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
8VYJ
 
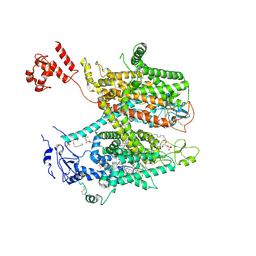 | | Structure of full-length human cardiac sodium channel - Class-I. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Biswas, R, Chinthalapudi, K. | | Deposit date: | 2024-02-08 | | Release date: | 2025-02-12 | | Last modified: | 2026-02-25 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structural basis of human Na v 1.5 gating mechanisms.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
7CZC
 
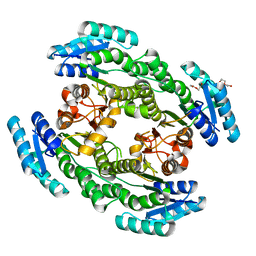 | | Crystal structure of apo-FabG from Vibrio harveyi | | Descriptor: | 3-oxoacyl-ACP reductase FabG, DI(HYDROXYETHYL)ETHER | | Authors: | Singh, B.K, Kumar, A, Paul, B, Biswas, R, Das, A.K. | | Deposit date: | 2020-09-08 | | Release date: | 2021-09-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of apo-FabG from Vibrio harveyi
To Be Published
|
|
7YL9
 
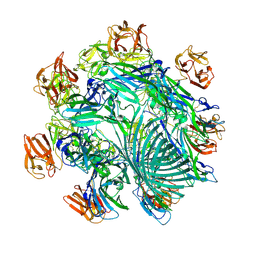 | | Cryo-EM structure of complete transmembrane channel E289A mutant Vibrio cholerae Cytolysin | | Descriptor: | Hemolysin | | Authors: | Mondal, A.K, Sengupta, N, Singh, M, Biswas, R, Lata, K, Lahiri, I, Dutta, S, Chattopadhyay, K. | | Deposit date: | 2022-07-25 | | Release date: | 2022-08-31 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (4.7 Å) | | Cite: | Cryo-EM structure of complete transmembrane channel E289A mutant Vibrio cholerae Cytolysin
J.Biol.Chem.
|
|
4OL4
 
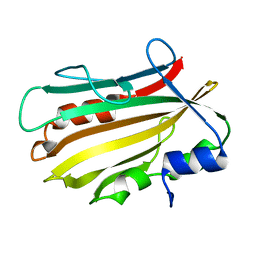 | | Crystal structure of secreted proline rich antigen MTC28 (Rv0040c) from Mycobacterium tuberculosis | | Descriptor: | Proline-rich 28 kDa antigen | | Authors: | Kundu, P, Biswas, R, Mukherjee, S, Reinhard, L, Mueller-dieckmann, J, Weiss, M.S, Das, A.K. | | Deposit date: | 2014-01-23 | | Release date: | 2015-01-28 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure-based Epitope Mapping of Mycobacterium tuberculosis Secretary Antigen MTC28
J.Biol.Chem., 291, 2016
|
|
4PWS
 
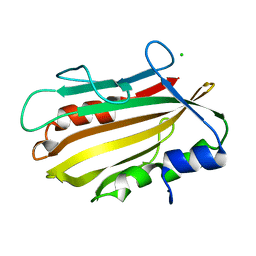 | | Crystal structure of secreted proline rich antigen MTC28 (Rv0040c) at 2.15 A with bound chloride from Mycobacterium tuberculosis | | Descriptor: | CHLORIDE ION, Proline-rich 28 kDa antigen | | Authors: | Kundu, P, Biswas, R, Mukherjee, S, Reinhard, L, Mueller-dieckmann, J, Weiss, M.S, Das, A.K. | | Deposit date: | 2014-03-21 | | Release date: | 2015-03-25 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structure-based Epitope Mapping of Mycobacterium tuberculosis Secretary Antigen MTC28
J.Biol.Chem., 291, 2016
|
|
4RUV
 
 | | Crystal structure of thioredoxin 2 from Staphylococcus aureus NCTC8325 | | Descriptor: | Thioredoxin | | Authors: | Bose, M, Biswas, R, Roychowdhury, A, Bhattacharyya, S, Ghosh, A.K, Das, A.K. | | Deposit date: | 2014-11-22 | | Release date: | 2015-12-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Elucidation of the mechanism of disulfide exchange between staphylococcal thioredoxin2 and thioredoxin reductase2: A structural insight.
Biochimie, 160, 2019
|
|
418D
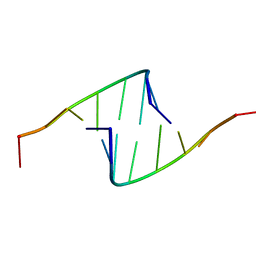 
 | |
419D
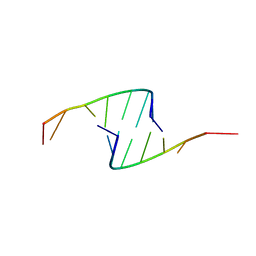 
 | |
7XNZ
 
 | | Native cystathionine beta-synthase of Mycobacterium tuberculosis. | | Descriptor: | PYRIDOXAL-5'-PHOSPHATE, Putative cystathionine beta-synthase Rv1077 | | Authors: | Bandyopadhyay, P, Pramanick, I, Biswas, R, Sabarinath, P.S, Sreedharan, S, Singh, S, Rajmani, R, Laxman, S, Dutta, S, Singh, A. | | Deposit date: | 2022-04-30 | | Release date: | 2022-05-25 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | S-Adenosylmethionine-responsive cystathionine beta-synthase modulates sulfur metabolism and redox balance in Mycobacterium tuberculosis.
Sci Adv, 8, 2022
|
|
7XOH
 
 | | Cystathionine beta-synthase of Mycobacterium tuberculosis in the presence of S-adenosylmethionine. | | Descriptor: | Probable cystathionine beta-synthase Rv1077 | | Authors: | Bandyopadhyay, P, Pramanick, I, Biswas, R, Sabarinath, P.S, Sreedharan, S, Singh, S, Rajmani, R, Laxman, S, Dutta, S, Singh, A. | | Deposit date: | 2022-05-01 | | Release date: | 2022-05-25 | | Last modified: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | S-Adenosylmethionine-responsive cystathionine beta-synthase modulates sulfur metabolism and redox balance in Mycobacterium tuberculosis.
Sci Adv, 8, 2022
|
|
7XOY
 
 | | Cystathionine beta-synthase of Mycobacterium tuberculosis in the presence of S-adenosylmethionine and serine. | | Descriptor: | Putative cystathionine beta-synthase Rv1077, [3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYL]-SERINE | | Authors: | Bandyopadhyay, P, Pramanick, I, Biswas, R, Sabarinath, P.S, Sreedharan, S, Singh, S, Rajmani, R, Laxman, S, Dutta, S, Singh, A. | | Deposit date: | 2022-05-01 | | Release date: | 2022-05-25 | | Last modified: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (4.25 Å) | | Cite: | S-Adenosylmethionine-responsive cystathionine beta-synthase modulates sulfur metabolism and redox balance in Mycobacterium tuberculosis.
Sci Adv, 8, 2022
|
|
5ZY8
 
 | | Crystal structure of C terminal truncated HadBC (3R-Hydroxyacyl-ACP Dehydratase) complex from Mycobacterium tuberculosis | | Descriptor: | 3-hydroxyacyl-ACP dehydratase, UPF0336 protein Rv0637 | | Authors: | Singh, B.K, Biswas, R, Bhattacharyya, S, Basak, A, Das, A.K. | | Deposit date: | 2018-05-23 | | Release date: | 2019-06-12 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.899 Å) | | Cite: | The C-terminal end of mycobacterial HadBC regulates AcpM interaction during the FAS-II pathway: a structural perspective.
Febs J., 2022
|
|
9LIB
 
 | |
9LJD
 
 | |
9NFU
 
 | |
9NH7
 
 | | CryoEM Structure of De Novo VHH, VHH_flu_01, bound to influenza HA, strain A/USA:Iowa/1943 H1N1. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin HA1 chain, ... | | Authors: | Borst, A.J, Weidle, C. | | Deposit date: | 2025-02-24 | | Release date: | 2025-11-12 | | Last modified: | 2026-01-07 | | Method: | ELECTRON MICROSCOPY (3.02 Å) | | Cite: | Atomically accurate de novo design of antibodies with RFdiffusion.
Nature, 649, 2026
|
|
1KVY
 
 | |
