2IRO
 
 | | The NMR Structures of (rGCUGAGGCU)2 and (rGCGGAUGCU)2 | | Descriptor: | 5'-R(P*GP*CP*GP*GP*AP*UP*GP*CP*U)-3' | | Authors: | Tolbert, B.S. | | Deposit date: | 2006-10-16 | | Release date: | 2007-02-20 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of (rGCUGAGGCU)(2) and (rGCGGAUGCU)(2): Probing the Structural Features That Shape the Thermodynamic Stability of GA Pairs(,).
Biochemistry, 46, 2007
|
|
2IRN
 
 | | The NMR Structures of (rGCUGAGGCU)2 and (rGCGGAUGCU)2 | | Descriptor: | 5'-R(P*GP*CP*UP*GP*AP*GP*GP*CP*U)-3' | | Authors: | Tolbert, B.S. | | Deposit date: | 2006-10-15 | | Release date: | 2007-02-20 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of (rGCUGAGGCU)(2) and (rGCGGAUGCU)(2): Probing the Structural Features That Shape the Thermodynamic Stability of GA Pairs(,).
Biochemistry, 46, 2007
|
|
2KYD
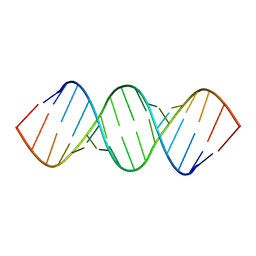 
 | | RDC and RCSA refinement of an A-form RNA: Improvements in Major Groove Width | | Descriptor: | RNA (5'-R(*CP*UP*AP*GP*UP*UP*AP*GP*CP*UP*AP*AP*CP*UP*AP*G)-3') | | Authors: | Tolbert, B.S, Summers, M.F, Miyazaki, Y, Barton, S, Kinde, B, Stark, P, Singh, R, Bax, A, Case, D. | | Deposit date: | 2010-05-24 | | Release date: | 2010-07-07 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Major groove width variations in RNA structures determined by NMR and impact of 13C residual chemical shift anisotropy and 1H-13C residual dipolar coupling on refinement.
J.Biomol.Nmr, 47, 2010
|
|
7V06
 
 | |
2N4L
 
 | | Solution Structure of the HIV-1 Intron Splicing Silencer and its Interactions with the UP1 Domain of hnRNP A1 | | Descriptor: | RNA (53-MER) | | Authors: | Tolbert, B.S, Jain, N, Morgan, C.E, Rife, B.D, Salemi, M. | | Deposit date: | 2015-06-23 | | Release date: | 2015-12-02 | | Last modified: | 2016-03-16 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the HIV-1 Intron Splicing Silencer and Its Interactions with the UP1 Domain of Heterogeneous Nuclear Ribonucleoprotein (hnRNP) A1.
J.Biol.Chem., 291, 2016
|
|
6XB7
 
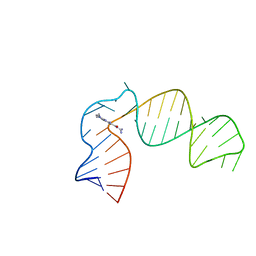 | | IRES-targeting Small Molecule Inhibits Enterovirus 71 Replication via Allosteric Stabilization of a Ternary Complex | | Descriptor: | 3-amino-N-(diaminomethylidene)-5-(dimethylamino)-6-(phenylethynyl)pyrazine-2-carboxamide, RNA (41-MER) | | Authors: | Tolbert, B.S, Davila, J, Sugarman, A.L, Chiu, L.Y. | | Deposit date: | 2020-06-05 | | Release date: | 2020-09-02 | | Last modified: | 2020-10-07 | | Method: | SOLUTION NMR | | Cite: | IRES-targeting small molecule inhibits enterovirus 71 replication via allosteric stabilization of a ternary complex.
Nat Commun, 11, 2020
|
|
5V16
 
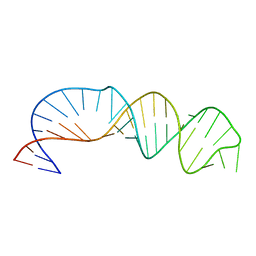 | |
5V17
 
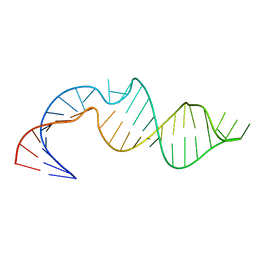 | |
2L1F
 
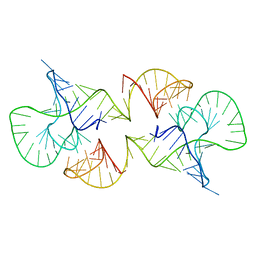 | | Structure of a conserved retroviral RNA packaging element by NMR spectroscopy and cryo-electron tomography | | Descriptor: | RNA (65-MER), RNA (66-MER) | | Authors: | Summers, M.F, Irobalieva, R.N, Tolbert, B, Smalls-Manty, A, Iyalla, K, Loeliger, K, D'Souza, V, Khant, H, Schmid, M, Garcia, E, Telesnitsky, A, Chiu, W, Miyazaki, Y. | | Deposit date: | 2010-07-28 | | Release date: | 2010-10-27 | | Last modified: | 2020-02-05 | | Method: | SOLUTION NMR | | Cite: | Structure of a conserved retroviral RNA packaging element by NMR spectroscopy and cryo-electron tomography.
J.Mol.Biol., 404, 2010
|
|
4YOE
 
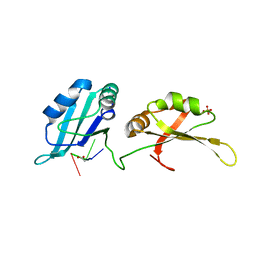 | | Structure of UP1 bound to RNA 5'-AGU-3' | | Descriptor: | ACETATE ION, Heterogeneous nuclear ribonucleoprotein A1, RNA AGU, ... | | Authors: | Meagher, J.L, Stuckey, J.A. | | Deposit date: | 2015-03-11 | | Release date: | 2015-06-10 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | The First Crystal Structure of the UP1 Domain of hnRNP A1 Bound to RNA Reveals a New Look for an Old RNA Binding Protein.
J.Mol.Biol., 427, 2015
|
|
2LDL
 
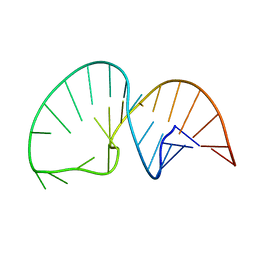 | | Solution NMR Structure of the HIV-1 Exon Splicing Silencer 3 | | Descriptor: | RNA (27-MER) | | Authors: | Mishler, C, Levengood, J.D, Johnson, C.A, Rajan, P, Znosko, B.M. | | Deposit date: | 2011-05-27 | | Release date: | 2011-12-28 | | Last modified: | 2012-02-08 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the HIV-1 Exon Splicing Silencer 3.
J.Mol.Biol., 415, 2012
|
|
2KY0
 
 | |
2KY1
 
 | |
2KXZ
 
 | |
2KY2
 
 | |
6DHS
 
 | | Structure of hnRNP H qRRM1,2 | | Descriptor: | Heterogeneous nuclear ribonucleoprotein H | | Authors: | Meagher, J.L, Stuckey, J.A. | | Deposit date: | 2018-05-21 | | Release date: | 2018-09-12 | | Last modified: | 2019-12-18 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Differential Conformational Dynamics Encoded by the Inter-qRRM linker of hnRNP H.
J. Am. Chem. Soc., 2018
|
|
6DG1
 
 | |
