4GLL
 
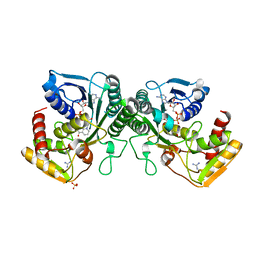 | | Crystal structure of human UDP-xylose synthase. | | Descriptor: | GUANIDINE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, SULFATE ION, ... | | Authors: | Wood, Z.A, Polizzi, S.J. | | Deposit date: | 2012-08-14 | | Release date: | 2012-09-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Human UDP-alpha-d-xylose Synthase and Escherichia coli ArnA Conserve a Conformational Shunt That Controls Whether Xylose or 4-Keto-Xylose Is Produced.
Biochemistry, 51, 2012
|
|
4LK3
 
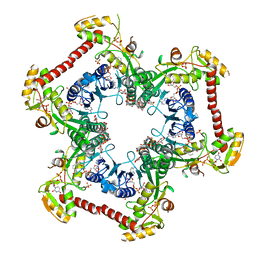 | | Crystal structure of Human UDP-xylose synthase R236A substitution | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, PYROPHOSPHATE 2-, SULFATE ION, ... | | Authors: | Walsh Jr, R.M, Polizzi, S.J, Wood, Z.A. | | Deposit date: | 2013-07-05 | | Release date: | 2013-08-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Man o' war mutation in UDP-alpha-D-xylose synthase favors the abortive catalytic cycle and uncovers a latent potential for hexamer formation.
Biochemistry, 54, 2015
|
|
4M55
 
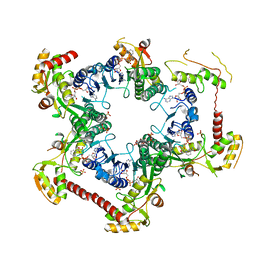 | | Crystal structure of Human UDP-xylose synthase R236H substitution | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, PYROPHOSPHATE 2-, SULFATE ION, ... | | Authors: | Walsh Jr, R.M, Polizzi, S.J, Wood, Z.A. | | Deposit date: | 2013-08-08 | | Release date: | 2013-08-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Man o' war mutation in UDP-alpha-D-xylose synthase favors the abortive catalytic cycle and uncovers a latent potential for hexamer formation.
Biochemistry, 54, 2015
|
|
2YFU
 
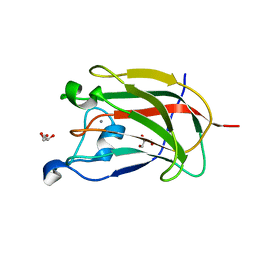 | | CBM62 FROM CLOSTRIDIUM THERMOCELLUM XYL5A | | Descriptor: | CALCIUM ION, CARBOHYDRATE BINDING FAMILY 6, GLYCEROL | | Authors: | Montanier, C.Y, Correia, M.A.S, Flint, J.E, Zhu, Y, Basle, A, Mckee, L.S, Prates, J.A.M, Polizzi, S.J, Coutinho, P.M, Henrissat, B, Fontes, C.M.G.A, Gilbert, H.J. | | Deposit date: | 2011-04-08 | | Release date: | 2011-05-18 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | A Novel, Noncatalytic Carbohydrate-Binding Module Displays Specificity for Galactose-Containing Polysaccharides Through Calcium-Mediated Oligomerization.
J.Biol.Chem., 286, 2011
|
|
2YG0
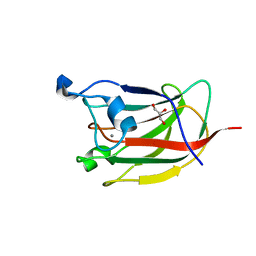 
 | | CBM62 FROM CLOSTRIDIUM THERMOCELLUM XYL5A | | Descriptor: | CALCIUM ION, CARBOHYDRATE BINDING FAMILY 6, GLYCEROL, ... | | Authors: | Montanier, C.Y, Correia, M.A.S, Flint, J.E, Zhu, Y, Basle, A, Mckee, L.S, Prates, J.A.M, Polizzi, S.J, Coutinho, P.M, Henrissat, B, Fontes, C.M.G.A, Gilbert, H.J. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A Novel, Noncatalytic Carbohydrate-Binding Module Displays Specificity for Galactose-Containing Polysaccharides Through Calcium-Mediated Oligomerization.
J.Biol.Chem., 286, 2011
|
|
2YFZ
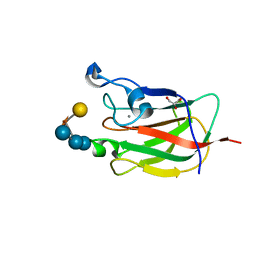 
 | | CBM62 FROM CLOSTRIDIUM THERMOCELLUM XYL5A | | Descriptor: | CALCIUM ION, CARBOHYDRATE BINDING FAMILY 6, GLYCEROL, ... | | Authors: | Montanier, C.Y, Correia, M.A.S, Flint, J.E, Zhu, Y, Basle, A, Mckee, L.S, Prates, J.A.M, Polizzi, S.J, Coutinho, P.M, Henrissat, B, Fontes, C.M.G.A, Gilbert, H.J. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A Novel, Noncatalytic Carbohydrate-Binding Module Displays Specificity for Galactose-Containing Polysaccharides Through Calcium-Mediated Oligomerization.
J.Biol.Chem., 286, 2011
|
|
2YB7
 
 | | CBM62 in complex with 6-alpha-D-Galactosyl-mannotriose | | Descriptor: | CALCIUM ION, CARBOHYDRATE BINDING FAMILY 6, GLYCEROL, ... | | Authors: | Montanier, C.Y, Correia, M.A.S, Flint, J.E, Zhu, Y, McKee, L.S, Prates, J.A.M, Polizzi, S.J, Coutinho, P.M, Henrissat, B, Fontes, C.M.G.A, Gilbert, H.J. | | Deposit date: | 2011-03-02 | | Release date: | 2011-08-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A Novel, Noncatalytic Carbohydrate-Binding Module Displays Specificity for Galactose-Containing Polysaccharides Through Calcium-Mediated Oligomerization.
J.Biol.Chem., 286, 2011
|
|
3PRJ
 
 | |
3PTZ
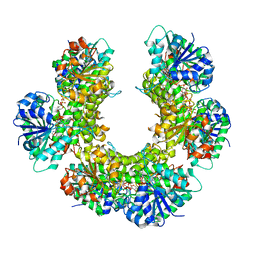 
 | |
