3KH7
 
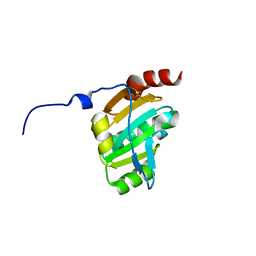 | | Crystal structure of the periplasmic soluble domain of reduced CcmG from Pseudomonas aeruginosa | | Descriptor: | Thiol:disulfide interchange protein dsbE | | Authors: | Di Matteo, A, Calosci, N, Gianni, S, Jemth, P, Brunori, M, Travaglini Allocatelli, C. | | Deposit date: | 2009-10-30 | | Release date: | 2010-04-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and functional characterization of CcmG from Pseudomonas aeruginosa, a key component of the bacterial cytochrome c maturation apparatus.
Proteins, 78, 2010
|
|
3KH9
 
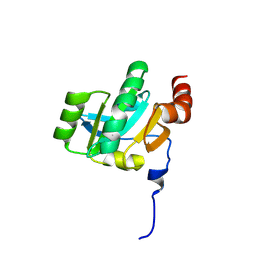 | | Crystal structure of the periplasmic soluble domain of oxidized CcmG from Pseudomonas aeruginosa | | Descriptor: | Thiol:disulfide interchange protein dsbE | | Authors: | Di Matteo, A, Calosci, N, Gianni, S, Jemth, P, Brunori, M, Travaglini Allocatelli, C. | | Deposit date: | 2009-10-30 | | Release date: | 2010-04-07 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and functional characterization of CcmG from Pseudomonas aeruginosa, a key component of the bacterial cytochrome c maturation apparatus.
Proteins, 78, 2010
|
|
2X7Z
 
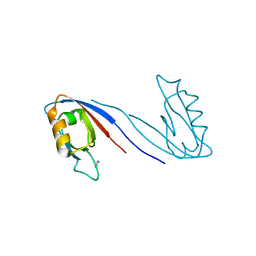 | | Crystal Structure of the SAP97 PDZ2 I342W C378A mutant protein domain | | Descriptor: | AMMONIUM ION, DISKS LARGE HOMOLOG 1, IMIDAZOLE | | Authors: | Haq, S.R, Jurgens, M.C, Chi, C.N, Elfstrom, L, Koh, C.S, Selmer, M, Gianni, S, Jemth, P. | | Deposit date: | 2010-03-04 | | Release date: | 2010-03-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Plastic Energy Landscape of Protein Folding: A Triangular Folding Mechanism with an Equilibrium Intermediate for a Small Protein Domain.
J.Biol.Chem., 285, 2010
|
|
4AMH
 
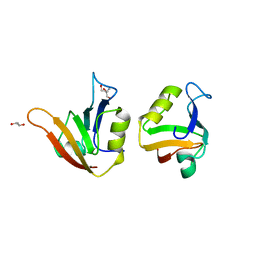 | | Influence of circular permutation on the folding pathway of a PDZ domain | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, DISKS LARGE HOMOLOG 1, GLYCEROL | | Authors: | Hultqvist, G, Punekar, A.S, Chi, C.N, Selmer, M, Gianni, S, Jemth, P. | | Deposit date: | 2012-03-10 | | Release date: | 2012-12-05 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Tolerance of Protein Folding to a Circular Permutation in a Pdz Domain
Plos One, 7, 2012
|
|
6ES7
 
 | |
6ES6
 
 | |
6ES5
 
 | |
8QNV
 
 | |
8QNJ
 
 | |
8QNT
 
 | |
7OSR
 
 | |
7OSW
 | |
1VJ6
 
 | |
6Y8X
 
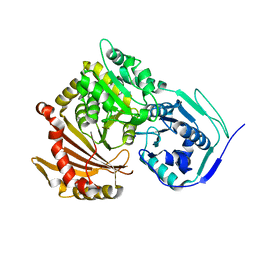 | |
2WIO
 
 | | Structure of the histidine tagged, open cytochrome P450 Eryk from S. erythraea | | Descriptor: | ERYTHROMYCIN B/D C-12 HYDROXYLASE, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Savino, C, Montemiglio, L.C, Sciara, G, Miele, A.E, Kedrew, S.G, Gianni, S, Vallone, B. | | Deposit date: | 2009-05-14 | | Release date: | 2009-07-21 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Investigating the Structural Plasticity of a Cytochrome P450: Three-Dimensional Structures of P450 Eryk and Binding to its Physiological Substrate.
J.Biol.Chem., 284, 2009
|
|
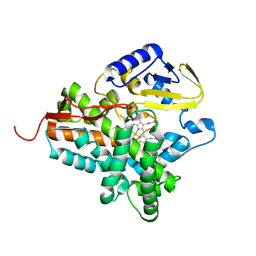
2JJN
 
 | | Structure of closed cytochrome P450 EryK | | Descriptor: | CYTOCHROME P450 113A1, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Savino, C, Sciara, G, Miele, A.E, Kendrew, S.G, Vallone, B. | | Deposit date: | 2008-04-15 | | Release date: | 2009-07-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Investigating the Structural Plasticity of a Cytochrome P450: Three-Dimensional Structures of P450 Eryk and Binding to its Physiological Substrate.
J.Biol.Chem., 284, 2009
|
|
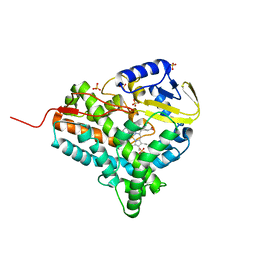
2JJO
 
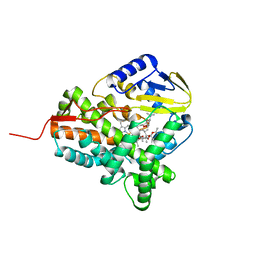 | | Structure of cytochrome P450 EryK in complex with its natural substrate erD | | Descriptor: | CYTOCHROME P450 113A1, Erythromycin D, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Savino, C, Sciara, G, Miele, A.E, Kendrew, S.G, Vallone, B. | | Deposit date: | 2008-04-15 | | Release date: | 2009-07-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Investigating the Structural Plasticity of a Cytochrome P450: Three-Dimensional Structures of P450 Eryk and Binding to its Physiological Substrate.
J.Biol.Chem., 284, 2009
|
|
6Y8Y
 
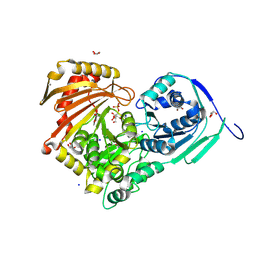 | | Structure of Baltic Herring (Clupea Harengus) Phosphoglucomutase 5 (PGM5) with bound Glucose-1-Phosphate | | Descriptor: | 1-O-phosphono-alpha-D-glucopyranose, ACETATE ION, CALCIUM ION, ... | | Authors: | Gustafsson, R, Eckhard, U, Selmer, M. | | Deposit date: | 2020-03-06 | | Release date: | 2020-12-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure and Characterization of Phosphoglucomutase 5 from Atlantic and Baltic Herring-An Inactive Enzyme with Intact Substrate Binding.
Biomolecules, 10, 2020
|
|
6Y8Z
 
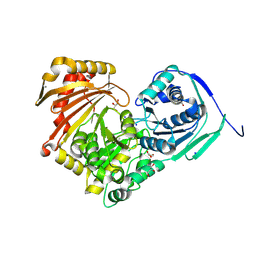 | | Structure of Baltic Herring (Clupea Harengus) Phosphoglucomutase 5 (PGM5) | | Descriptor: | ACETATE ION, CALCIUM ION, GLYCEROL, ... | | Authors: | Gustafsson, R, Eckhard, U, Selmer, M. | | Deposit date: | 2020-03-06 | | Release date: | 2020-12-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure and Characterization of Phosphoglucomutase 5 from Atlantic and Baltic Herring-An Inactive Enzyme with Intact Substrate Binding.
Biomolecules, 10, 2020
|
|
3ZRT
 | | Crystal structure of human PSD-95 PDZ1-2 | | Descriptor: | DISKS LARGE HOMOLOG 4 | | Authors: | Sorensen, P.L, Kastrup, J.S, Gajhede, M. | | Deposit date: | 2011-06-19 | | Release date: | 2012-03-21 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.398 Å) | | Cite: | A High-Affinity, Dimeric Inhibitor of Psd-95 Bivalently Interacts with Pdz1-2 and Protects Against Ischemic Brain Damage.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1UZC
 
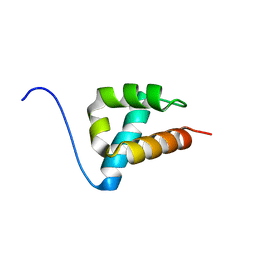 | | THE STRUCTURE OF AN FF DOMAIN FROM HUMAN HYPA/FBP11 | | Descriptor: | HYPOTHETICAL PROTEIN FLJ21157 | | Authors: | Allen, M.D, Jemth, P, Friedler, A, Schon, O, Bycroft, M. | | Deposit date: | 2004-03-09 | | Release date: | 2004-04-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Structure of an Ff Domain from Human Hypa/Fbp11.
J.Mol.Biol., 323, 2002
|
|
7BN2
 
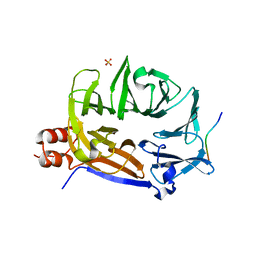 | |
7BN3
 
 | |
7BN1
 
 | |
