-検索条件
-検索結果
検索 (著者・登録者: kunpeng & l)の結果63件中、1から50件目までを表示しています
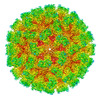

EMDB-43222: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18

EMDB-43292: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated

EMDB-43293: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment

PDB-8vgr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18

PDB-8vjr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated

PDB-8vjs: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment

EMDB-41008: 
Cryo-EM structure of the DHA bound FFA4-Gq complex

EMDB-41010: 
Cryo-EM structure of the Butyrate bound FFA2-Gq complex

EMDB-41013: 
Cryo-EM structure of the DHA bound FFA1-Gq complex

PDB-8t3q: 
Cryo-EM structure of the DHA bound FFA4-Gq complex

PDB-8t3s: 
Cryo-EM structure of the Butyrate bound FFA2-Gq complex

PDB-8t3v: 
Cryo-EM structure of the DHA bound FFA1-Gq complex

EMDB-41007: 
Cryo-EM structure of the TUG-891 bound FFA4-Gq complex

EMDB-41014: 
Cryo-EM structure of the DHA bound FFA1-Gq complex(mask on receptor)

PDB-8t3o: 
Cryo-EM structure of the TUG-891 bound FFA4-Gq complex

EMDB-40450: 
Cryo-EM structure of CMKLR1 signaling complex

PDB-8sg1: 
Cryo-EM structure of CMKLR1 signaling complex

EMDB-28087: 
CryoEM structure of GSDMB in complex with shigella IpaH7.8

EMDB-28583: 
CryoEM structure of GSDMB pore without transmembrane beta-barrel

EMDB-28584: 
CryoEM structure of the GSDMB pore

PDB-8efp: 
CryoEM structure of GSDMB in complex with shigella IpaH7.8

PDB-8et1: 
CryoEM structure of GSDMB pore without transmembrane beta-barrel

PDB-8et2: 
CryoEM structure of the GSDMB pore

EMDB-25824: 
aRML prion fibril
PDB-7td6: 
aRML prion fibril

EMDB-22675: 
Structure of VCP dodecamer purified from H1299 cells

EMDB-22676: 
Structure of apo VCP dodecamer generated from bacterially recombinant VCP/p97

EMDB-22678: 
Structure of apo VCP hexamer generated from bacterially recombinant VCP/p97

PDB-7k56: 
Structure of VCP dodecamer purified from H1299 cells

PDB-7k57: 
Structure of apo VCP dodecamer generated from bacterially recombinant VCP/p97

PDB-7k59: 
Structure of apo VCP hexamer generated from bacterially recombinant VCP/p97

EMDB-23927: 
Cryo-EM structure of affinity captured human p97 hexamer

EMDB-23928: 
Cryo-EM structure of affinity captured human p97 double-hexamer

EMDB-21695: 
Full phage G capsid cryoEM structure at 6.1 Angstrom resolution

EMDB-21702: 
Empty phage G cryoEM capsid structure at 9 Angstrom resolution

PDB-6wkk: 
Phage G gp27 major capsid proteins and gp26 decoration proteins

EMDB-0983: 
Luminal ring of the Xenopus laevis nuclear pore complex

EMDB-0984: 
Luminal ring bumper-7 conformation of the Xenopus laevis nuclear pore complex

EMDB-0985: 
Luminal ring bumper-6 conformation of the Xenopus laevis nuclear pore complex

EMDB-0986: 
Cytoplasmic ring of the Xenopus laevis nuclear pore complex

EMDB-0997: 
Inner ring of the Xenopus laevis nuclear pore complex

EMDB-0998: 
Nuclear ring of the Xenopus laevis nuclear pore complex

EMDB-20227: 
Dataset II: Sub-3 Angstrom Apoferritin Structure Determined With Full Range of Phase Shifts Using A Single Position Of Volta Phase Plate

EMDB-20225: 
Dataset I: Sub-3 Angstrom Apoferritin Structure Determined With Full Range of Phase Shifts Using A Single Position Of Volta Phase Plate

EMDB-20228: 
Dataset III: Sub-3 Angstrom Apoferritin Structure Determined With Full Range of Phase Shifts Using A Single Position Of Volta Phase Plate

EMDB-20229: 
Dataset IV: Sub-3 Angstrom Apoferritin Structure Determined With Full Range of Phase Shifts Using A Single Position Of Volta Phase Plate

EMDB-9673: 
CryoEM structure of Mud Crab Dicistrovirus

EMDB-9754: 
Cryo-EM structure of Mud crab tombus-like virus at 3.3 Angstroms resolution

EMDB-9755: 
Structure of heated Mud crab dicistrovirus

EMDB-9756: 
Empty particle structure of heated mud crab dicistrovirus
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
