6IS1
 
 | |
6BR7
 
 | |
7OD9
 
 | | Crystal structure of activated CheY fused to the C-terminal domain of CheF | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, C-terminal domain of CheF from Methanococcus maripaludis, MAGNESIUM ION, ... | | Authors: | Altegoer, F, Weiland, P, Bange, G. | | Deposit date: | 2021-04-29 | | Release date: | 2022-04-27 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural insights into the mechanism of archaellar rotational switching.
Nat Commun, 13, 2022
|
|
1D5W
 
 | | PHOSPHORYLATED FIXJ RECEIVER DOMAIN | | Descriptor: | SULFATE ION, TRANSCRIPTIONAL REGULATORY PROTEIN FIXJ | | Authors: | Birck, C, Mourey, L, Gouet, P, Fabry, B, Schumacher, J, Rousseau, P, Kahn, D, Samama, J.P. | | Deposit date: | 1999-10-12 | | Release date: | 2000-10-11 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Conformational changes induced by phosphorylation of the FixJ receiver domain.
Structure Fold.Des., 7, 1999
|
|
4ML3
 
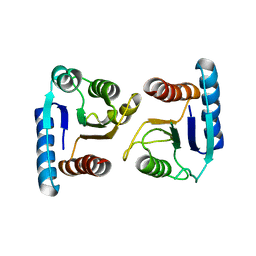 | | X-ray structure of ComE D58A REC domain from Streptococcus pneumoniae | | Descriptor: | Response regulator | | Authors: | Boudes, M, Sanchez, D, Durand, D, Graille, M, van Tilbeurgh, H, Quevillon-Cheruel, S. | | Deposit date: | 2013-09-06 | | Release date: | 2014-02-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Structural insights into the dimerization of the response regulator ComE from Streptococcus pneumoniae.
Nucleic Acids Res., 42, 2014
|
|
1CEY
 
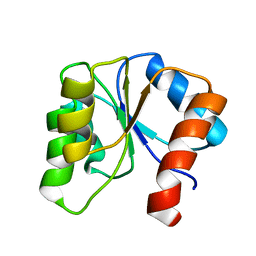 | | ASSIGNMENTS, SECONDARY STRUCTURE, GLOBAL FOLD, AND DYNAMICS OF CHEMOTAXIS Y PROTEIN USING THREE-AND FOUR-DIMENSIONAL HETERONUCLEAR (13C,15N) NMR SPECTROSCOPY | | Descriptor: | CHEY | | Authors: | Moy, F.J, Lowry, D.F, Matsumura, P, Dahlquist, F.W, Krywko, J.E, Domaille, P.J. | | Deposit date: | 1994-11-23 | | Release date: | 1995-02-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Assignments, secondary structure, global fold, and dynamics of chemotaxis Y protein using three- and four-dimensional heteronuclear (13C,15N) NMR spectroscopy.
Biochemistry, 33, 1994
|
|
6SY9
 
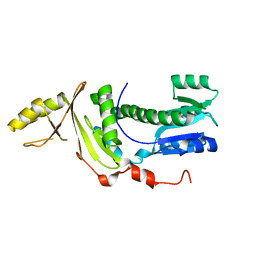 | | Structure of the Legionella pneumophila response regulator LqsR | | Descriptor: | Response regulator | | Authors: | Hochstrasser, R, Hutter, C.A.J, Arnold, F.M, Baerlocher, K, Seeger, M.A, Hilbi, H. | | Deposit date: | 2019-09-27 | | Release date: | 2020-02-05 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structure of the Legionella response regulator LqsR reveals amino acids critical for phosphorylation and dimerization.
Mol.Microbiol., 113, 2020
|
|
6SWB
 
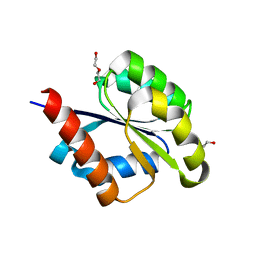 | | The REC domain of AraT, a response regulator from Geobacillus stearothermophilus | | Descriptor: | DI(HYDROXYETHYL)ETHER, Two-component response regulator | | Authors: | Lansky, S, Lavid, N, Shoham, Y, Shoham, G. | | Deposit date: | 2019-09-20 | | Release date: | 2020-10-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.259 Å) | | Cite: | The REC domain of AraT, a response regulator from Geobacillus stearothermophilus
To Be Published
|
|
6Z4E
 
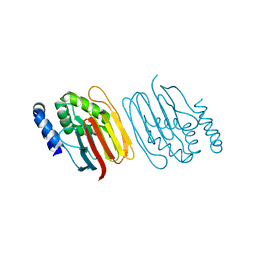 | | The structure of the C-terminal domain of RssB from E. coli | | Descriptor: | Regulator of RpoS | | Authors: | Zeth, K, Dimce, M, Terrence, D.M, Schuenemann, V, Dougan, D. | | Deposit date: | 2020-05-25 | | Release date: | 2020-07-29 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insight into the RssB-Mediated Recognition and Delivery of sigma s to the AAA+ Protease, ClpXP.
Biomolecules, 10, 2020
|
|
6TNE
 
 | |
2VUH
 
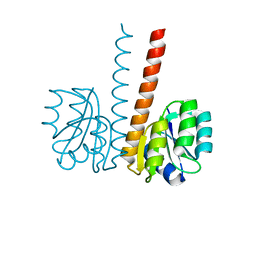 | |
1K68
 
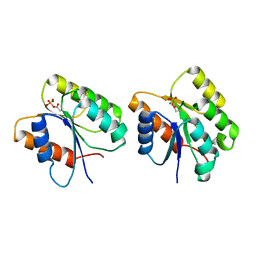 | | Crystal Structure of the Phosphorylated Cyanobacterial Phytochrome Response Regulator RcpA | | Descriptor: | MAGNESIUM ION, Phytochrome Response Regulator RcpA | | Authors: | Benda, C, Scheufler, C, Tandeau de Marsac, N, Gaertner, W. | | Deposit date: | 2001-10-15 | | Release date: | 2003-12-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structures of two cyanobacterial response regulators in apo- and phosphorylated form reveal a novel dimerization motif of phytochrome-associated response regulators
Biophys.J., 87, 2004
|
|
2VUI
 
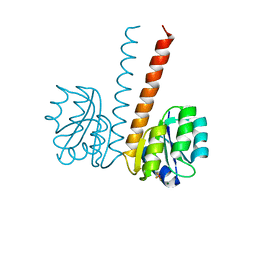 | | Crystal structure of the HupR receiver domain in inhibitory phospho- state | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, HYDROGENASE TRANSCRIPTIONAL REGULATORY PROTEIN HUPR1, MAGNESIUM ION | | Authors: | Davies, K.M, Lowe, E.D, Venien-Bryan, C, Johnson, L.N. | | Deposit date: | 2008-05-26 | | Release date: | 2008-11-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The Hupr Receiver Domain Crystal Structure in its Nonphospho and Inhibitory Phospho States.
J.Mol.Biol., 385, 2009
|
|
1K66
 
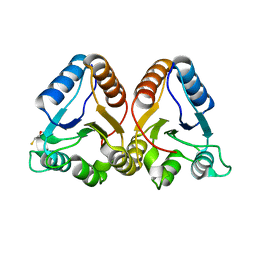 | | Crystal Structure of the Cyanobacterial Phytochrome Response Regulator, RcpB | | Descriptor: | BETA-MERCAPTOETHANOL, Phytochrome Response Regulator RcpB | | Authors: | Benda, C, Scheufler, C, Tandeau de Marsac, N, Gaertner, W. | | Deposit date: | 2001-10-15 | | Release date: | 2003-12-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structures of two cyanobacterial response regulators in apo- and phosphorylated form reveal a novel dimerization motif of phytochrome-associated response regulators
Biophys.J., 87, 2004
|
|
6OEC
 
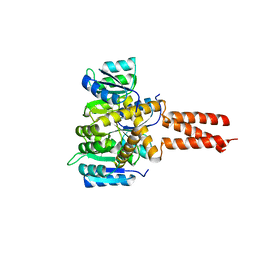 | | Yeast Spc42 Trimeric Coiled-Coil Amino Acids 181-211 fused to PDB: 3H5I | | Descriptor: | CALCIUM ION, Response regulator/sensory box protein/GGDEF domain protein,Spindle pole body component SPC42 | | Authors: | Drennan, A.C, Shivaani, K, Seeger, M.A, Andreas, M.P, Gardner, J.M, Sether, E.K.R, Jasperson, S.L, Rayment, I. | | Deposit date: | 2019-03-27 | | Release date: | 2019-04-24 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.514 Å) | | Cite: | Structure and function of Spc42 coiled-coils in yeast centrosome assembly and duplication.
Mol.Biol.Cell, 30, 2019
|
|
6ONT
 
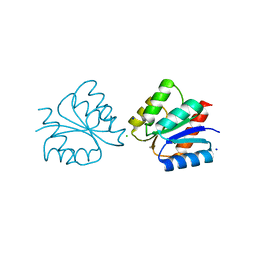 | | Structure of the Francisella response regulator 1452 receiver domain | | Descriptor: | CALCIUM ION, CHLORIDE ION, SODIUM ION, ... | | Authors: | Milton, M.E, Cavanagh, J. | | Deposit date: | 2019-04-22 | | Release date: | 2020-03-25 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.803 Å) | | Cite: | Francisella novicidaTwo-Component System Response Regulator BfpR Modulates iglC Gene Expression, Antimicrobial Peptide Resistance, and Biofilm Production.
Front Cell Infect Microbiol, 10, 2020
|
|
7W9H
 
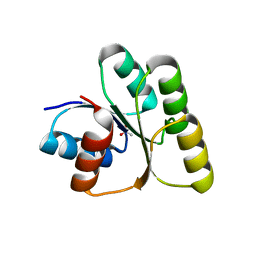 | | Crystal structure of the receiver domain of the transcription regulator FleR from Pseudomonas aeruginosa | | Descriptor: | ACETATE ION, CALCIUM ION, Response regulator protein FleR | | Authors: | Sahoo, P.K, Sheenu, n, Jain, D. | | Deposit date: | 2021-12-09 | | Release date: | 2022-12-14 | | Last modified: | 2024-01-03 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | REC domain stabilizes the active heptamer of sigma 54 -dependent transcription factor, FleR from Pseudomonas aeruginosa.
Iscience, 26, 2023
|
|
6TG7
 
 | |
6M8O
 
 | |
6C40
 
 | | CheY41PyTyrD54K from Thermotoga maritima | | Descriptor: | COPPER (II) ION, Chemotaxis protein CheY | | Authors: | Merz, G.E, Muok, A.R, Crane, B.R. | | Deposit date: | 2018-01-11 | | Release date: | 2018-10-17 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Site-Specific Incorporation of a Cu2+Spin Label into Proteins for Measuring Distances by Pulsed Dipolar Electron Spin Resonance Spectroscopy.
J Phys Chem B, 122, 2018
|
|
7LCM
 
 | | Receiver Domain of RssB bound to beryllofluoride | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, MAGNESIUM ION, Regulator of RpoS | | Authors: | Deaconescu, A.M, Schwartz, J, Son, J. | | Deposit date: | 2021-01-11 | | Release date: | 2021-04-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Phospho-dependent signaling during the general stress response by the atypical response regulator and ClpXP adaptor RssB.
Protein Sci., 30, 2021
|
|
7L9C
 
 | | Receiver Domain of RssB | | Descriptor: | Regulator of RpoS | | Authors: | Deaconescu, A.M, Son, J, Schwartz, J. | | Deposit date: | 2021-01-03 | | Release date: | 2021-04-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Phospho-dependent signaling during the general stress response by the atypical response regulator and ClpXP adaptor RssB.
Protein Sci., 30, 2021
|
|
2IYN
 
 | | The co-factor-induced pre-active conformation in PhoB | | Descriptor: | MAGNESIUM ION, PHOSPHATE REGULON TRANSCRIPTIONAL REGULATORY PROTEIN PHOB | | Authors: | Sola, M, Drew, D.L, Blanco, A.G, Gomis-Ruth, F.X, Coll, M. | | Deposit date: | 2006-07-19 | | Release date: | 2006-08-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | The Cofactor-Induced Pre-Active Conformation in Phob.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
8FK2
 
 | | The N-terminal VicR from Streptococcus mutans | | Descriptor: | Putative response regulator CovR VicR-like protein | | Authors: | Zhang, H, Wu, H. | | Deposit date: | 2022-12-20 | | Release date: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Small Molecule Attenuates Bacterial Virulence by Targeting Conserved Response Regulator.
Mbio, 14, 2023
|
|
2ID7
 
 | | 1.75 A Structure of T87I Phosphono-CheY | | Descriptor: | Chemotaxis protein cheY | | Authors: | Halkides, C.J, Haas, R.M, McAdams, K.A, Casper, E.S, Santarsiero, B.D, Mesecar, A.D. | | Deposit date: | 2006-09-14 | | Release date: | 2007-09-25 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The structures of T87I phosphono-CheY and T87I/Y106W phosphono-CheY help to explain their binding affinities to the FliM and CheZ peptides.
Arch.Biochem.Biophys., 479, 2008
|
|
