1R6L
 
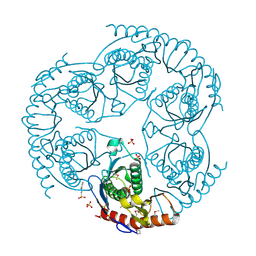 | | Crystal Structure Of The tRNA Processing Enzyme Rnase pH From Pseudomonas Aeruginosa | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, Ribonuclease PH, SULFATE ION | | Authors: | Choi, J.M, Park, E.Y, Kim, J.H, Chang, S.K, Cho, Y. | | Deposit date: | 2003-10-15 | | Release date: | 2004-02-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Probing the functional importance of the hexameric ring structure of RNase PH
J.BIOL.CHEM., 279, 2004
|
|
3LZT
 
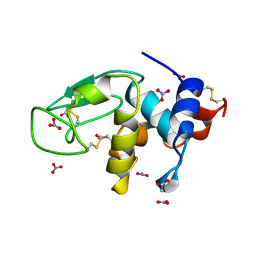 | | REFINEMENT OF TRICLINIC LYSOZYME AT ATOMIC RESOLUTION | | Descriptor: | ACETATE ION, LYSOZYME, NITRATE ION | | Authors: | Walsh, M.A, Schneider, T, Sieker, L.C, Dauter, Z, Lamzin, V, Wilson, K.S. | | Deposit date: | 1997-03-23 | | Release date: | 1998-03-25 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (0.925 Å) | | Cite: | Refinement of triclinic hen egg-white lysozyme at atomic resolution.
Acta Crystallogr.,Sect.D, 54, 1998
|
|
3M1K
 
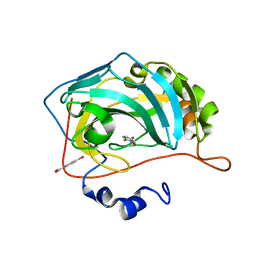 | | Carbonic Anhydrase in complex with fragment | | Descriptor: | (4-CARBOXYPHENYL)(CHLORO)MERCURY, 1-hydroxy-2-sulfanylpyridinium, Carbonic anhydrase 2, ... | | Authors: | Schulze Wischeler, J, Heine, A, Klebe, G. | | Deposit date: | 2010-03-05 | | Release date: | 2010-11-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Bidentate Zinc chelators for alpha-carbonic anhydrases that produce a trigonal bipyramidal coordination geometry.
Chemmedchem, 5, 2010
|
|
3M04
 | | Carbonic Anhydrase II in complex with novel sulfonamide inhibitor | | Descriptor: | (4-CARBOXYPHENYL)(CHLORO)MERCURY, Carbonic anhydrase 2, ZINC ION, ... | | Authors: | Schulze Wischeler, J, Heine, A, Klebe, G, Sandner, N.U. | | Deposit date: | 2010-03-02 | | Release date: | 2011-05-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural investigation and inhibitor studies on Carbonic Anhydrase II
To be Published
|
|
3M2K
 
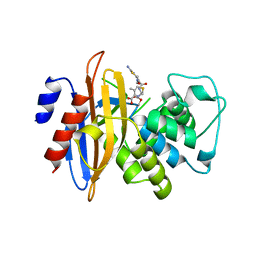 | | Crystal Structure of fluorescein-labeled Class A -beta lactamase PenP in complex with cefotaxime | | Descriptor: | Beta-lactamase, CEFOTAXIME, C3' cleaved, ... | | Authors: | Zhao, Y.X, Leung, Y.C, Wong, W.T. | | Deposit date: | 2010-03-07 | | Release date: | 2011-03-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural studies of the mechanism for biosensing antibiotics in a fluorescein-labeled beta-lactamase.
BMC Struct. Biol., 11, 2011
|
|
3M28
 
 | |
3M41
 
 | | Crystal structure of the mutant V182A of orotidine 5'-monophosphate decarboxylase from Methanobacterium thermoautotrophicum | | Descriptor: | GLYCEROL, Orotidine 5'-phosphate decarboxylase | | Authors: | Fedorov, A.A, Fedorov, E.V, Wood, B.M, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2010-03-10 | | Release date: | 2010-06-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Conformational changes in orotidine 5'-monophosphate decarboxylase: "remote" residues that stabilize the active conformation.
Biochemistry, 49, 2010
|
|
3M2E
 
 | |
3M4D
 
 | |
3M48
 
 | | GCN4 Leucine Zipper Peptide Mutant | | Descriptor: | General control protein GCN4, SODIUM ION | | Authors: | Du, S, Kettering, R.D, Alvarado, J.J, Tortajada, A, Yeh, J.I. | | Deposit date: | 2010-03-10 | | Release date: | 2011-02-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.451 Å) | | Cite: | GCN4 Leucine Zipper Peptide Mutant
To be Published
|
|
1REE
 
 | | ENDO-1,4-BETA-XYLANASE II COMPLEX WITH 3,4-EPOXYBUTYL-BETA-D-XYLOSIDE | | Descriptor: | (3S)-3-hydroxybutyl beta-D-xylopyranoside, BENZOIC ACID, ENDO-1,4-BETA-XYLANASE II | | Authors: | Rouvinen, J, Havukainen, R, Torronen, A. | | Deposit date: | 1995-12-21 | | Release date: | 1997-01-11 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Covalent binding of three epoxyalkyl xylosides to the active site of endo-1,4-xylanase II from Trichoderma reesei.
Biochemistry, 35, 1996
|
|
3MAE
 
 | | CRYSTAL STRUCTURE OF PROBABLE DIHYDROLIPOAMIDE ACETYLTRANSFERASE FROM LISTERIA MONOCYTOGENES 4b F2365 | | Descriptor: | 2-oxoisovalerate dehydrogenase E2 component, dihydrolipoamide acetyltransferase, CHLORIDE ION, ... | | Authors: | Patskovsky, Y, Toro, R, Gilmore, M, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2010-03-23 | | Release date: | 2010-04-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | CRYSTAL STRUCTURE OF A CATALYTIC DOMAIN OF DIHYDROLIPOAMIDE ACETYLTRANSFERASE FROM LISTERIA MONOCYTOGENES 4b F2365
To be Published
|
|
1REI
 
 | | THE MOLECULAR STRUCTURE OF A DIMER COMPOSED OF THE VARIABLE PORTIONS OF THE BENCE-JONES PROTEIN REI REFINED AT 2.0 ANGSTROMS RESOLUTION | | Descriptor: | BENCE-JONES PROTEIN REI (LIGHT CHAIN) | | Authors: | Epp, O, Lattman, E.E, Colman, P, Fehlhammer, H, Bode, W, Schiffer, M, Huber, R, Palm, W. | | Deposit date: | 1976-03-17 | | Release date: | 1976-05-19 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The molecular structure of a dimer composed of the variable portions of the Bence-Jones protein REI refined at 2.0-A resolution.
Biochemistry, 14, 1975
|
|
1RFS
 
 | | RIESKE SOLUBLE FRAGMENT FROM SPINACH | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, RIESKE PROTEIN | | Authors: | Carrell, C.J, Zhang, H, Cramer, W.A, Smith, J.L. | | Deposit date: | 1997-08-14 | | Release date: | 1998-01-28 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Biological identity and diversity in photosynthesis and respiration: structure of the lumen-side domain of the chloroplast Rieske protein.
Structure, 5, 1997
|
|
3M8J
 
 | |
3M8Q
 
 | | HIV-1 RT with AMINOPYRIMIDINE NNRTI | | Descriptor: | 3,5-dimethyl-4-{[2-({1-[4-(methylsulfonyl)benzyl]piperidin-4-yl}amino)pyrimidin-4-yl]oxy}benzonitrile, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Harris, S.F, Villasenor, A. | | Deposit date: | 2010-03-18 | | Release date: | 2010-05-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Discovery of piperidin-4-yl-aminopyrimidines as HIV-1 reverse transcriptase inhibitors. N-benzyl derivatives with broad potency against resistant mutant viruses.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
1RN1
 
 | | THREE-DIMENSIONAL STRUCTURE OF GLN 25-RIBONUCLEASE T1 AT 1.84 ANGSTROMS RESOLUTION: STRUCTURAL VARIATIONS AT THE BASE RECOGNITION AND CATALYTIC SITES | | Descriptor: | RIBONUCLEASE T1 ISOZYME, SULFATE ION | | Authors: | Arni, R.K, Pal, G.P, Ravichandran, K.G, Tulinsky, A, Walz Junior, F.G, Metcalf, P. | | Deposit date: | 1991-11-22 | | Release date: | 1994-01-31 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Three-dimensional structure of Gln25-ribonuclease T1 at 1.84-A resolution: structural variations at the base recognition and catalytic sites.
Biochemistry, 31, 1992
|
|
3M9B
 
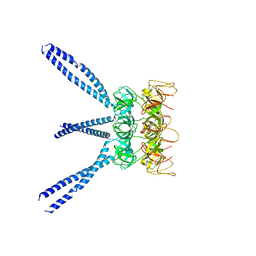 | |
3MB9
 
 | | Human Aldose Reductase mutant T113A complexed with Zopolrestat | | Descriptor: | 3,4-DIHYDRO-4-OXO-3-((5-TRIFLUOROMETHYL-2-BENZOTHIAZOLYL)METHYL)-1-PHTHALAZINE ACETIC ACID, Aldose reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Koch, C, Heine, A, Klebe, G. | | Deposit date: | 2010-03-25 | | Release date: | 2011-03-09 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Ligand-induced fit affects binding modes and provokes changes in crystal packing of aldose reductase
Biochim.Biophys.Acta, 1810, 2011
|
|
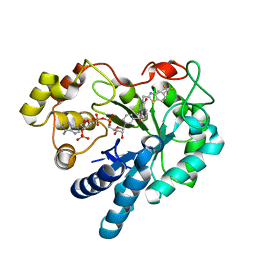
1RNW
 
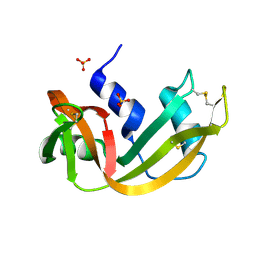 | | RECOMBINANT RIBONUCLEASE A CRYSTALLIZED FROM 80% AMMONIUM SULPHATE | | Descriptor: | RIBONUCLEASE A, SULFATE ION | | Authors: | Fedorov, A.A, Joseph-Mccarthy, D, Fedorov, E.V, Sirakova, D, Graf, I, Almo, S.C. | | Deposit date: | 1996-11-08 | | Release date: | 1997-04-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Ionic interactions in crystalline bovine pancreatic ribonuclease A.
Biochemistry, 35, 1996
|
|
1ROV
 
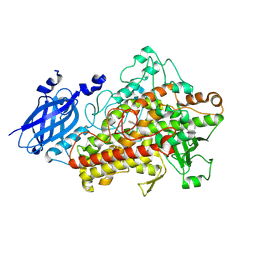 | | Lipoxygenase-3 Treated with Cumene Hydroperoxide | | Descriptor: | FE (III) ION, Seed lipoxygenase-3 | | Authors: | Vahedi-Faridi, A, Brault, P.A, Shah, P, Kim, Y.W, Dunham, W.R, Funk, M.O. | | Deposit date: | 2003-12-02 | | Release date: | 2004-03-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Interaction between non-heme iron of lipoxygenases and cumene hydroperoxide: basis for enzyme activation, inactivation, and inhibition
J.Am.Chem.Soc., 126, 2004
|
|
3MGC
 
 | | Teg12 Apo | | Descriptor: | GLYCEROL, IMIDAZOLE, N-L-ALPHA-ASPARTYL L-PHENYLALANINE 1-METHYL ESTER, ... | | Authors: | Bick, M.J, Banik, J.J, Darst, S.A, Brady, S.F. | | Deposit date: | 2010-04-05 | | Release date: | 2010-06-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Crystal structures of the glycopeptide sulfotransferase Teg12 in a complex with the teicoplanin aglycone.
Biochemistry, 49, 2010
|
|
3MC5
 
 | |
1RQB
 
 | | Propionibacterium shermanii transcarboxylase 5S subunit | | Descriptor: | COBALT (II) ION, transcarboxylase 5S subunit | | Authors: | Hall, P.R, Zheng, R, Antony, L, Pusztai-Carey, M, Carey, P.R, Yee, V.C. | | Deposit date: | 2003-12-04 | | Release date: | 2004-09-07 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Transcarboxylase 5S structures: assembly and catalytic mechanism of a multienzyme complex subunit.
Embo J., 23, 2004
|
|
1RKI
 
 | | Structure of pag5_736 from P. aerophilum with three disulphide bonds | | Descriptor: | ACETATE ION, CHLORIDE ION, HEXAETHYLENE GLYCOL, ... | | Authors: | Beeby, M, Ryttersgaard, C, Boutz, D.R, Perry, L.J, Yeates, T.O. | | Deposit date: | 2003-11-21 | | Release date: | 2005-01-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The Genomics of Disulfide Bonding and Protein Stabilization in Thermophiles.
Plos Biol., 3, 2005
|
|
