1WFF
 
 | | Solution structure of the zf-AN1 domain from mouse RIKEN cDNA 2810002D23 protein | | Descriptor: | RIKEN cDNA 2810002D23 protein, ZINC ION | | Authors: | Tomizawa, T, Kigawa, T, Izumi, K, Yoshida, M, Yamazaki, T, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-26 | | Release date: | 2004-11-26 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the zf-AN1 domain from mouse RIKEN cDNA 2810002D23 protein
To be Published
|
|
1WD2
 
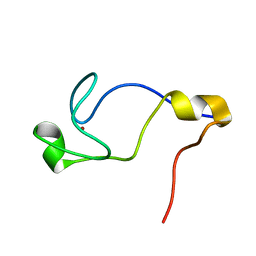 | | Solution Structure of the C-terminal RING from a RING-IBR-RING (TRIAD) motif | | Descriptor: | Ariadne-1 protein homolog, ZINC ION | | Authors: | Capili, A.D, Edghill, E.L, Wu, K, Borden, K.L.B. | | Deposit date: | 2004-05-11 | | Release date: | 2004-07-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of the C-terminal RING Finger from a RING-IBR-RING/TRIAD Motif Reveals a Novel Zinc-binding Domain Distinct from a RING
J.Mol.Biol., 340, 2004
|
|
1WI1
 
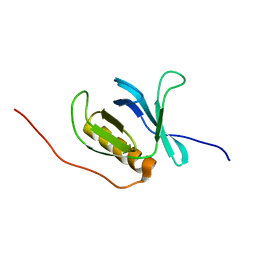 | | Solution structure of the PH domain of human calcium-dependent activator protein for secretion (CAPS) | | Descriptor: | calcium-dependent activator protein for secretion, CAPS | | Authors: | Yoneyama, M, Izumi, K, Yoshida, M, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-28 | | Release date: | 2004-11-28 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the PH domain of human calcium-dependent activator protein for secretion (CAPS)
To be Published
|
|
1WG1
 
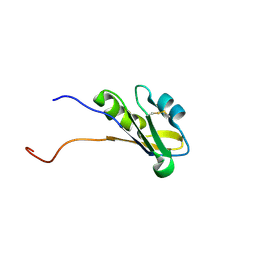 | | Solution structure of RNA binding domain in BAB13405(homolog EXC-7) | | Descriptor: | KIAA1579 protein | | Authors: | Tsuda, K, Muto, Y, Nagata, T, Suzuki, S, Someya, T, Kigawa, T, Terada, T, Shirouzu, M, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-27 | | Release date: | 2004-11-27 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RNA binding domain in BAB13405(homolog EXC-7)
To be Published
|
|
1WHD
 | | Solution structure of the PDZ domain of RGS3 | | Descriptor: | regulator of G-protein signaling 3 | | Authors: | Nakanishi, T, Nemoto, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-28 | | Release date: | 2004-11-28 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the PDZ domain of RGS3
To be Published
|
|
2K0G
 
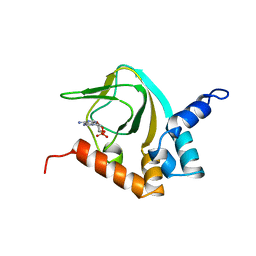 | |
1YMZ
 | | CC45, An Artificial WW Domain Designed Using Statistical Coupling Analysis | | Descriptor: | CC45 | | Authors: | Socolich, M, Lockless, S.W, Russ, W.P, Lee, H, Gardner, K.H, Ranganathan, R. | | Deposit date: | 2005-01-22 | | Release date: | 2005-09-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Evolutionary information for specifying a protein fold.
Nature, 437, 2005
|
|
1YHD
 
 | | The solution structure of YGGX from Escherichia Coli | | Descriptor: | UPF0269 protein yggX | | Authors: | Osborne, M.J, Siddiqui, N, Landgraf, D, Pomposiello, P.J, Gehring, K. | | Deposit date: | 2005-01-07 | | Release date: | 2005-01-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the oxidative stress-related protein YggX from Escherichia coli.
Protein Sci., 14, 2005
|
|
2GPQ
 | |
2KQZ
 
 | |
2KR0
 
 | |
1ZIT
 
 | |
2KZ7
 
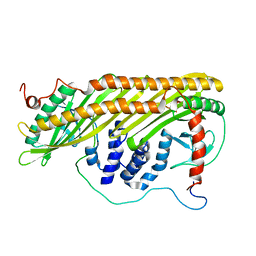 | |
1ZHC
 
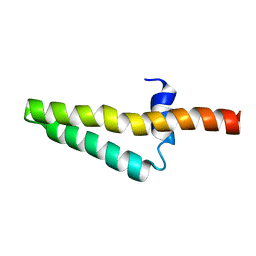 | |
1ZXF
 
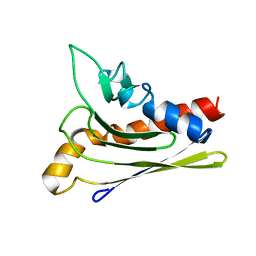 | | Solution structure of a self-sacrificing resistance protein, CalC from Micromonospora echinospora | | Descriptor: | CalC | | Authors: | Singh, S, Hager, M.H, Zhang, C, Griffith, B.R, Lee, M.S, Hallenga, K, Markley, J.L, Thorson, J.S, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2005-06-08 | | Release date: | 2005-12-13 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural insight into the self-sacrifice mechanism of enediyne resistance.
Acs Chem.Biol., 1, 2006
|
|
1Z8M
 
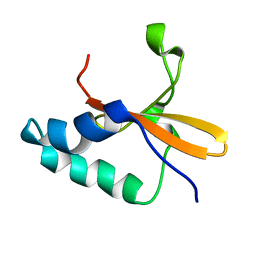 | |
1Z0R
 | | Solution Structure of the N-terminal DNA Recognition Domain of the Bacillus subtilis Transcription-State Regulator AbrB | | Descriptor: | Transition state regulatory protein abrB | | Authors: | Bobay, B.G, Andreeva, A, Mueller, G.A, Cavanagh, J, Murzin, A.G. | | Deposit date: | 2005-03-02 | | Release date: | 2005-03-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Revised structure of the AbrB N-terminal domain unifies a diverse superfamily of putative DNA-binding proteins.
Febs Lett., 579, 2005
|
|
2JXT
 
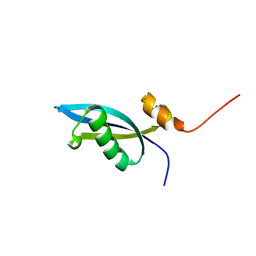 | | Solution structure of 50S ribosomal protein LX from Methanobacterium thermoautotrophicum. Northeast Structural Genomics Consortium (NESG) target TR80 | | Descriptor: | 50S ribosomal protein LX | | Authors: | Liu, G, Wang, D, Nwosu, C, Owens, L, Xiao, R, Liu, J, Baran, M.C, Swapna, G, Acton, T.B, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-11-29 | | Release date: | 2007-12-11 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution structure of 50S ribosomal protein LX from Methanobacterium thermoautotrophicum.
To be Published
|
|
2LOV
 | | AR55 solubilised in LPPG micelles | | Descriptor: | Apelin receptor | | Authors: | Langelaan, D.N, Rainey, J.K. | | Deposit date: | 2012-01-27 | | Release date: | 2013-01-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Preserved Transmembrane Segment Topology, Structure, and Dynamics in Disparate Micellar Environments.
J Phys Chem Lett, 8, 2017
|
|
1X3P
 | |
1WUZ
 
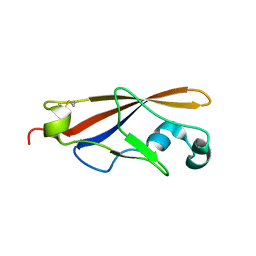 | | Structure of EC1 domain of CNR | | Descriptor: | Pcdha4 protein | | Authors: | Morishita, H, Umitsu, M, Yamaguchi, T, Murata, Y, Shibata, N, Udaka, K, Higuchi, Y, Akutsu, H, Yagi, T, Ikegami, T. | | Deposit date: | 2004-12-09 | | Release date: | 2005-12-13 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Structural diversity of the first cadherin domains revealed by the structure of CNR/Protocadherin alpha
To be Published
|
|
1WSX
 
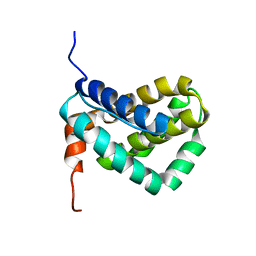 | | Solution structure of MCL-1 | | Descriptor: | myeloid cell leukemia sequence 1 | | Authors: | Day, C.L, Chen, L, Richardson, S.J, Harrison, P.J, Huang, D.C, Hinds, M.G. | | Deposit date: | 2004-11-12 | | Release date: | 2004-11-23 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Prosurvival Mcl-1 and Characterization of Its Binding by Proapoptotic BH3-only Ligands
J.Biol.Chem., 280, 2005
|
|
1X3Q
 | |
1X9L
 
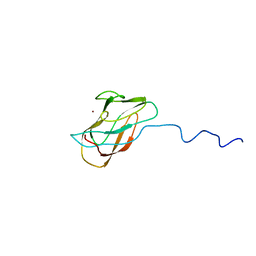 | | Solution structure of CuI-DR1885 from Deinococcus Radiodurans | | Descriptor: | COPPER (I) ION, CuI-DR1885 | | Authors: | Banci, L, Bertini, I, Ciofi-Baffoni, S, Katsari, E, Katsaros, N, Kubicek, K, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2004-08-23 | | Release date: | 2004-08-31 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A copper(I) protein possibly involved in the assembly of CuA center of bacterial cytochrome c oxidase.
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1YG0
 | |
