2JQP
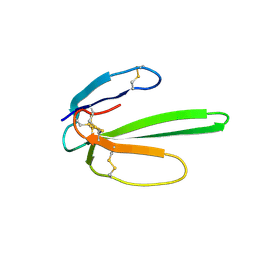 
 | | NMR structure determination of Bungatoxin from Bungarus candidus (Malayan Krait) | | Descriptor: | Weak toxin 1 | | Authors: | Vivekanandan, S, Jois, S.D, Kini, R.M, Troncone, L.R.P, de Magalhaes, L, Ujikawa, G.Y, Ramos, A.T. | | Deposit date: | 2007-06-06 | | Release date: | 2007-06-19 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | NMR solution Structure of Bungatoxin from Bungarus Candidus (Malayan Krait)venom
To be Published
|
|
2LA7
 
 | | NMR structure of the protein YP_557733.1 from Burkholderia xenovorans | | Descriptor: | Uncharacterized protein | | Authors: | Jaudzems, K, Serrano, P, Michael, G, Reto, H, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2011-03-04 | | Release date: | 2011-03-30 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the protein YP_557733.1 from Burkholderia xenovorans
To be Published
|
|
2L2U
 
 | |
2L2V
 
 | |
2MKS
 
 | |
2M9P
 
 | |
2MC8
 
 | |
2ML6
 
 | | NMR structure of protein ZP_02069618.1 from Bacteroides uniformis ATCC 8492 | | Descriptor: | Uncharacterized protein | | Authors: | Wallmann, A, Dutta, S.K, Serrano, P, Geralt, M, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2014-02-19 | | Release date: | 2014-03-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of protein ZP_02069618.1 from Bacteroides uniformis ATCC 8492.
To be Published
|
|
2M4L
 
 | |
2KP5
 
 | |
2KTS
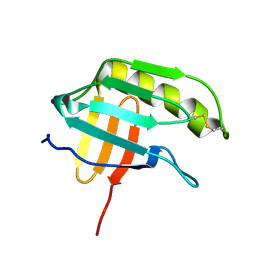 
 | | NMR structure of the protein NP_415897.1 | | Descriptor: | Heat shock protein hslJ | | Authors: | Serrano, P, Jaudzems, K, Geralt, M, Horst, R, Wuthrich, K, Wilson, I.A, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2010-02-06 | | Release date: | 2010-02-23 | | Last modified: | 2024-10-16 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the protein NP_415897.1
To be Published
|
|
2HLG
 
 | |
2JQO
 
 | | NMR solution structure of Bacillus subtilis YobA 21-120: Northeast Structural Genomics Consortium target SR547 | | Descriptor: | Hypothetical protein yobA | | Authors: | Cort, J.R, Ramelot, T.A, Chen, C.X, Jang, M, Cunningham, K, Ma, L, Xiao, R, Liu, J, Baran, M.C, Acton, T.B, Rost, B, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-06-06 | | Release date: | 2007-07-03 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of Bacillus subtilis YobA 21-120: Northeast Structural Genomics Consortium target SR547
To be Published
|
|
2LR4
 
 | |
2LIZ
 
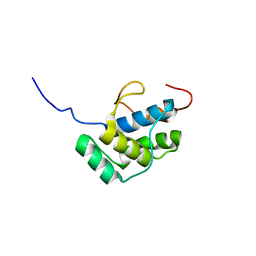 | |
2JTU
 
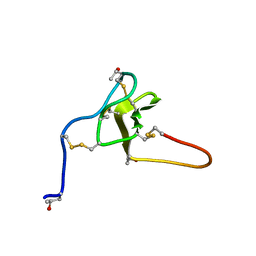 | | NMR structure of iota-RXIA(38) | | Descriptor: | I-superfamily conotoxin r11a | | Authors: | Wei, D, Norton, R. | | Deposit date: | 2007-08-06 | | Release date: | 2008-08-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | NMR structure of iota-RXIA(38)
To be Published
|
|
2JN9
 
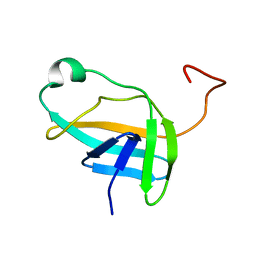 | | NMR solution structure of YkvR protein from Bacillus subtilis: NESG target SR358 | | Descriptor: | YkvR protein | | Authors: | Gurla, S.V.T, Aramini, J.M, Chi Ho, K, Cunningham, K, Ma, L.-C, Xiao, R, Baran, M.C, Acton, T.B, Liu, J, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-12-29 | | Release date: | 2007-05-01 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of YkvR protein from Bacillus subtilis: NESG target SR358
To be Published
|
|
2LR6
 
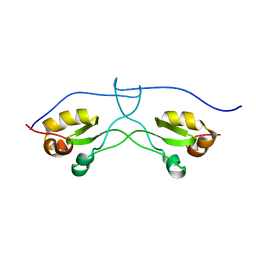 | | NMR structure of a LINE-1 type transposase domain-containing protein 1 (L1TD1) from Homo sapiens | | Descriptor: | LINE-1 type transposase domain-containing protein 1 | | Authors: | Serrano, P, Geralt, M, Mohanty, B, Wuthrich, K, Joint Center for Structural Genomics (JCSG), Partnership for Stem Cell Biology, Partnership for Stem Cell Biology (STEMCELL) | | Deposit date: | 2012-03-26 | | Release date: | 2012-07-04 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of a LINE-1 type transposase domain-containing protein 1 (L1TD1) from Homo sapiens
To be Published
|
|
2LL0
 
 | | NMR structure of the putative ATPase regulatory protein YP_916642.1 from Paracoccus denitrificans | | Descriptor: | Uncharacterized protein | | Authors: | Serrano, P, Geralt, M, Wuthrich, K, Morales-Rios, E, Zarco-Zavala, M, Garcia-Trejo, J.J, Dutta, S.K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2011-10-24 | | Release date: | 2011-12-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the putative ATPase regulatory protein YP_916642.1 from Paracoccus denitrificans
To be Published
|
|
2LRG
 
 | |
2LRA
 
 | | NMR Structure of Signal Sequence Deleted (SSD) Rv0603 Protein from Mycobacterium tuberculosis without N-terminal His-tag | | Descriptor: | POSSIBLE EXPORTED PROTEIN | | Authors: | Tripathi, S, Pulavarti, S, Yadav, R, Jain, A, Pathak, P, Meher, A, Arora, A. | | Deposit date: | 2012-03-28 | | Release date: | 2013-05-22 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of Signal Sequence Deleted (SSD) Rv0603 Protein from Mycobacterium tuberculosis without N-terminal His-tag
To be Published
|
|
1TI7
 
 | | CRYSTAL STRUCTURE OF NMRA, A NEGATIVE TRANSCRIPTIONAL REGULATOR, IN COMPLEX WITH NADP AT 1.7A RESOLUTION | | Descriptor: | CHLORIDE ION, GLYCEROL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Lamb, H.K, Leslie, K, Dodds, A.L, Nutley, M, Cooper, A, Johnson, C, Thompson, P, Stammers, D.K, Hawkins, A.R. | | Deposit date: | 2004-06-02 | | Release date: | 2004-06-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The negative transcriptional regulator NmrA discriminates between oxidized and reduced dinucleotides.
J.Biol.Chem., 278, 2003
|
|
2LGE
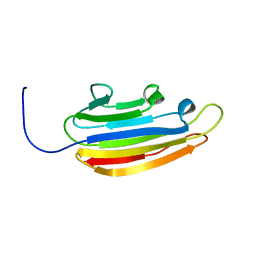 
 | |
2LG7
 
 | |
2L6O
 
 | | NMR structure of the protein YP_926445.1 from Shewanella Amazonensis | | Descriptor: | uncharacterized protein YP_926445.1 | | Authors: | Serrano, P, Geralt, M, Mohanty, B, Horst, R, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2010-11-23 | | Release date: | 2010-12-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the protein YP_926445.1 from Shewanella Amazonensis
To be Published
|
|
