8WWM
 
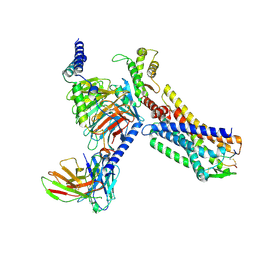 | | MCH-MCHR1-Gi complex, L2 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Gong, W, Ye, X, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-12-18 | | Method: | ELECTRON MICROSCOPY (2.81 Å) | | Cite: | Structural insights into physiological activation and antagonism of melanin-concentrating hormone receptor MCHR1.
Cell Discov, 10, 2024
|
|
8WWJ
 
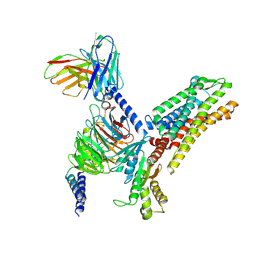 | | MCHR1-Gi complex,S2 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Ye, X, Gong, W, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (3.03 Å) | | Cite: | Structural insights into ligand recognition and activation of melanin-concentrating hormone receptor MCHR1
To Be Published
|
|
8WWK
 
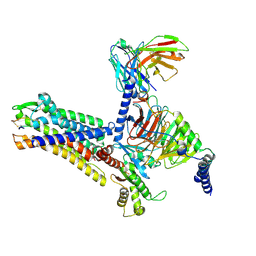 | | MCH-MCHR1-Gi complex, T1 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Gong, W, Ye, X, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-12-18 | | Method: | ELECTRON MICROSCOPY (2.61 Å) | | Cite: | Structural insights into physiological activation and antagonism of melanin-concentrating hormone receptor MCHR1.
Cell Discov, 10, 2024
|
|
8WWI
 
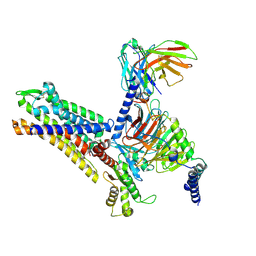 | | MCHR1-Gi complex,S3 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Gong, W, Ye, X, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (3.43 Å) | | Cite: | Structural insights into ligand recognition and activation of melanin-concentrating hormone receptor MCHR1
To Be Published
|
|
8WWN
 
 | | MCH-MCHR1-Gi complex,L1 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Gong, W, Ye, X, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-12-18 | | Method: | ELECTRON MICROSCOPY (2.65 Å) | | Cite: | Structural insights into physiological activation and antagonism of melanin-concentrating hormone receptor MCHR1.
Cell Discov, 10, 2024
|
|
8WWL
 
 | | MCH-MCHR1-Gi complex, T2 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Gong, W, Ye, X, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-12-18 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Structural insights into physiological activation and antagonism of melanin-concentrating hormone receptor MCHR1.
Cell Discov, 10, 2024
|
|
8WWH
 
 | | MCHR1-Gi complex,S1 state | | Descriptor: | Antibody fragment ScFv16, Fusion protein 1,Melanin-concentrating hormone receptor 1,Fusion protein 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Gong, W, Ye, X, Liu, G. | | Deposit date: | 2023-10-25 | | Release date: | 2024-11-13 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Structural insights into ligand recognition and activation of melanin-concentrating hormone receptor MCHR1
To Be Published
|
|
3U8X
 
 | | Crystal Structure of a chimera containing the N-terminal domain (residues 8-29) of drosophila Ciboulot and the C-terminal domain (residues 18-44) of bovine Thymosin-beta4, bound to G-actin-ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Renault, L, Husson, C, Carlier, M.F, Didry, D. | | Deposit date: | 2011-10-17 | | Release date: | 2012-01-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | How a single residue in individual beta-thymosin/WH2 domains controls their functions in actin assembly
Embo J., 31, 2012
|
|
3TYO
 
 | | Structure of neuronal nitric oxide synthase heme domain in complex with 6-(((3S,4S)-4-(2-((furan-2-ylmethyl)amino)ethoxy)pyrrolidin-3-yl)methyl)-4-methylpyridin-2-amine | | Descriptor: | 5,6,7,8-TETRAHYDROBIOPTERIN, 6-{[(3S,4S)-4-{2-[(furan-2-ylmethyl)amino]ethoxy}pyrrolidin-3-yl]methyl}-4-methylpyridin-2-amine, ACETATE ION, ... | | Authors: | Li, H, Poulos, T.L. | | Deposit date: | 2011-09-26 | | Release date: | 2012-03-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.927 Å) | | Cite: | Intramolecular hydrogen bonding: A potential strategy for more bioavailable inhibitors of neuronal nitric oxide synthase.
Bioorg.Med.Chem., 20, 2012
|
|
7P33
 
 | | Epstein-Barr virus encoded Bcl-2 homolog BHRF-1 in complex with Bid BH3 peptide | | Descriptor: | 1,2-ETHANEDIOL, Apoptosis regulator BHRF1, BH3-interacting domain death agonist p15, ... | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2021-07-07 | | Release date: | 2022-07-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.78542733 Å) | | Cite: | Crystal Structures of Epstein-Barr Virus Bcl-2 Homolog BHRF1 Bound to Bid and Puma BH3 Motif Peptides.
Viruses, 14, 2022
|
|
8X9S
 
 | | Identification, structure and agonist design of an androgen membrane receptor. | | Descriptor: | 5-ALPHA-DIHYDROTESTOSTERONE, Adhesion G-protein coupled receptor D1, Gs protein alpha subunit, ... | | Authors: | Ping, Y.Q, Yang, Z. | | Deposit date: | 2023-12-01 | | Release date: | 2025-02-12 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (3.49 Å) | | Cite: | Identification, structure, and agonist design of an androgen membrane receptor.
Cell, 188, 2025
|
|
8X9T
 
 | | Identification, structure and agonist design of an androgen membrane receptor | | Descriptor: | Adhesion G-protein coupled receptor D1, Gs protein alpha subunit, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Ping, Y.Q, Yang, Z. | | Deposit date: | 2023-12-01 | | Release date: | 2025-02-12 | | Last modified: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (2.75 Å) | | Cite: | Identification, structure, and agonist design of an androgen membrane receptor.
Cell, 188, 2025
|
|
8X9U
 
 | | Identification, structure and agonist design of an androgen membrane receptor | | Descriptor: | (5S,8R,9S,10S,13S,14S,17S)-17-hydroxy-1,10,13-trimethyl-4,5,6,7,8,9,11,12,14,15,16,17-dodecahydrocyclopenta[a]phenanthren-3-one, Adhesion G-protein coupled receptor D1, Gs protein alpha subunit, ... | | Authors: | Ping, Y.Q, Yang, Z. | | Deposit date: | 2023-12-01 | | Release date: | 2025-02-12 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | Identification, structure, and agonist design of an androgen membrane receptor.
Cell, 188, 2025
|
|
8X6L
 
 | | PRMT5:MEP50 WITH SCR-6920 AND SAM | | Descriptor: | 1-ethyl-8-[(2-methoxy-7-azaspiro[3.5]nonan-7-yl)carbonyl]-4-[2-oxidanylidene-2-[(3~{S})-1,2,3,4-tetrahydroisoquinolin-3-yl]ethyl]-2,3-dihydro-1,4-benzodiazepin-5-one, GLYCEROL, Methylosome protein WDR77, ... | | Authors: | Zhou, F, Zhou, F. | | Deposit date: | 2023-11-21 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | PRMT5:MEP50 WITH SCR-6920 AND SAM
To Be Published
|
|
3U82
 
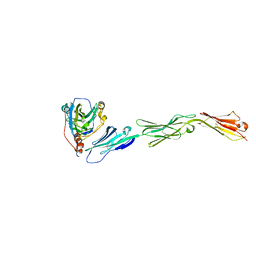 | | Binding of herpes simplex virus glycoprotein D to nectin-1 exploits host cell adhesion | | Descriptor: | Envelope glycoprotein D, Poliovirus receptor-related protein 1 | | Authors: | Zhang, N, Yan, J, Lu, G, Guo, Z, Fan, Z, Wang, J, Shi, Y, Qi, J, Gao, G.F. | | Deposit date: | 2011-10-15 | | Release date: | 2012-03-21 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3.164 Å) | | Cite: | Binding of herpes simplex virus glycoprotein D to nectin-1 exploits host cell adhesion.
Nat Commun, 2, 2011
|
|
3U96
 
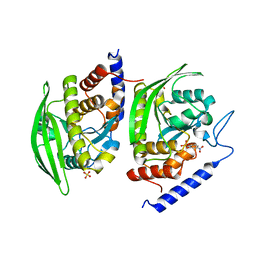 | | Crystal Structure of YopHQ357F(Catalytic Domain, Residues 163-468) in complex with pNCS | | Descriptor: | N,4-DIHYDROXY-N-OXO-3-(SULFOOXY)BENZENAMINIUM, SULFATE ION, Tyrosine-protein phosphatase yopH | | Authors: | Ho, M.C, Ke, S. | | Deposit date: | 2011-10-17 | | Release date: | 2012-08-29 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Investigation of catalytic loop structure, dynamics, and function relationship of Yersinia protein tyrosine phosphatase by temperature-jump relaxation spectroscopy and X-ray structural determination.
J.Phys.Chem.B, 116, 2012
|
|
7P9W
 
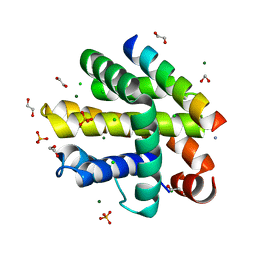 | | Epstein-Barr virus encoded apoptosis regulator BHRF1 in complex with Puma BH3 | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, AMMONIUM ION, ... | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2021-07-28 | | Release date: | 2022-08-10 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.00010061 Å) | | Cite: | Crystal Structures of Epstein-Barr Virus Bcl-2 Homolog BHRF1 Bound to Bid and Puma BH3 Motif Peptides.
Viruses, 14, 2022
|
|
7PCV
 
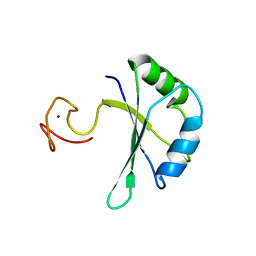 | |
8XJN
 
 | | Cloprosetnol bound Thromboxane A2 receptor-Gq Protein Complex | | Descriptor: | Antibody fragment scFv16, Cloprostenol, Engineered miniGq, ... | | Authors: | Zhang, X, Li, X, Liu, G, Gong, W. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Structural basis for ligand recognition and activation of the prostanoid receptors.
Cell Rep, 43, 2024
|
|
8XJL
 
 | | PGF2-alpha bound Prostaglandin F2-alpha receptor-Gq Protein Complex | | Descriptor: | (Z)-7-[(1R,2R,3R,5S)-3,5-bis(oxidanyl)-2-[(E,3S)-3-oxidanyloct-1-enyl]cyclopentyl]hept-5-enoic acid, Antibody fragment scFv16, Engineered miniGq, ... | | Authors: | Zhang, X, Li, X, Liu, G, Gong, W. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (2.77 Å) | | Cite: | Structural basis for ligand recognition and activation of the prostanoid receptors.
Cell Rep, 43, 2024
|
|
8XJK
 
 | | Cloprosetnol bound Prostaglandin F2-alpha receptor-Gq Protein Complex | | Descriptor: | Antibody fragment scFv16, Cloprostenol, Engineered miniGq, ... | | Authors: | Zhang, X, Li, X, Liu, G, Gong, W. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (2.63 Å) | | Cite: | Structural basis for ligand recognition and activation of the prostanoid receptors.
Cell Rep, 43, 2024
|
|
8XJO
 
 | | U46619 bound Thromboxane A2 receptor-Gq Protein Complex | | Descriptor: | (5Z)-7-{(1R,4S,5S,6R)-6-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-2-oxabicyclo[2.2.1]hept-5-yl}hept-5-enoic acid, Antibody fragment scFv16, Engineered miniGq, ... | | Authors: | Zhang, X, Li, X, Liu, G, Gong, W. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Structural basis for ligand recognition and activation of the prostanoid receptors.
Cell Rep, 43, 2024
|
|
8XJM
 
 | | Latanoprost acid bound Prostaglandin F2-alpha receptor-Gq Protein Complex | | Descriptor: | Antibody fragment scFv16, Engineered miniGq, Fusion tag,Prostaglandin F2-alpha receptor,LgBiT, ... | | Authors: | Zhang, X, Li, X, Liu, G, Gong, W. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (2.85 Å) | | Cite: | Structural basis for ligand recognition and activation of the prostanoid receptors.
Cell Rep, 43, 2024
|
|
8XOH
 
 | | Cryo-EM structure of GPR30-Gq complex structure in the presence of E2 | | Descriptor: | G-protein coupled estrogen receptor 1, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Liu, H, Xu, P, Xu, H.E. | | Deposit date: | 2024-01-01 | | Release date: | 2024-04-10 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structural and functional evidence that GPR30 is not a direct estrogen receptor.
Cell Res., 34, 2024
|
|
3SXK
 
 | |
