3DGY
 
 | |
3DH2
 
 | |
3D5I
 
 | |
3D4A
 
 | |
3D5G
 
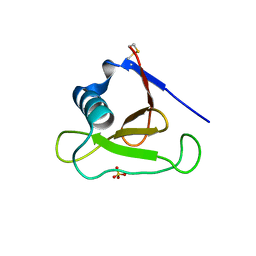 | | Structure of ribonuclease Sa2 complexes with mononucleotides: new aspects of catalytic reaction and substrate recognition | | Descriptor: | Ribonuclease, SULFATE ION | | Authors: | Bauerova-Hlinkova, V, Dvorsky, R, Povazanec, F, Sevcik, J. | | Deposit date: | 2008-05-16 | | Release date: | 2009-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of RNase Sa2 complexes with mononucleotides - new aspects of catalytic reaction and substrate recognition
Febs J., 276, 2009
|
|
3DA7
 
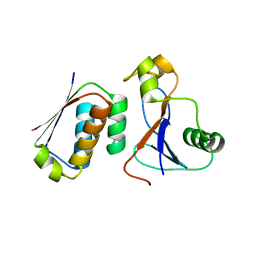 | | A conformationally strained, circular permutant of barnase | | Descriptor: | Barnase circular permutant, Barstar | | Authors: | Mitrousis, G, Butler, J, Loh, S.N, Cingolani, G. | | Deposit date: | 2008-05-28 | | Release date: | 2009-04-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and thermodynamic analysis of a conformationally strained circular permutant of barnase.
Biochemistry, 48, 2009
|
|
2ZA4
 
 | |
1ZGX
 
 | |
2F5W
 
 | | Cross-linked barnase soaked in 3 M thiourea | | Descriptor: | Ribonuclease, THIOUREA, ZINC ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-28 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F56
 
 | | Barnase cross-linked with glutaraldehyde soaked in 6M urea | | Descriptor: | Ribonuclease, UREA, ZINC ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-25 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.955 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F5M
 
 | | Cross-linked barnase soaked in bromo-ethanol | | Descriptor: | 2-BROMOETHANOL, Ribonuclease | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-26 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F4Y
 
 | | Barnase cross-linked with glutaraldehyde | | Descriptor: | Ribonuclease, ZINC ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-24 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2C4B
 
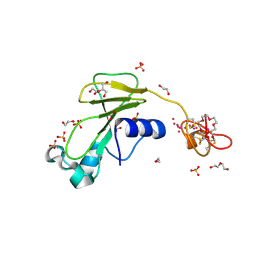 | | Inhibitor cystine knot protein McoEeTI fused to the catalytically inactive barnase mutant H102A | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, BARNASE MCOEETI FUSION, ... | | Authors: | Niemann, H.H, Schmoldt, H.U, Wentzel, A, Kolmar, H, Heinz, D.W. | | Deposit date: | 2005-10-18 | | Release date: | 2005-11-21 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Barnase Fusion as a Tool to Determine the Crystal Structure of the Small Disulfide-Rich Protein Mcoeeti.
J.Mol.Biol., 356, 2006
|
|
1TTO
 
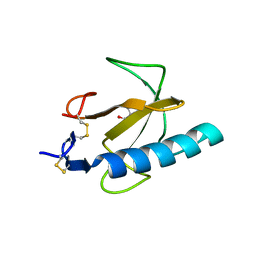 | | Crystal structure of the Rnase T1 variant R2 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, RNase T1 | | Authors: | Hahn, U, Czaja, R, Perbandt, M, Betzel, C. | | Deposit date: | 2004-06-23 | | Release date: | 2005-09-06 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Purine activity of RNase T1RV is further improved by substitution of Trp59 by tyrosine
Biochem.Biophys.Res.Commun., 336, 2005
|
|
1YNV
 
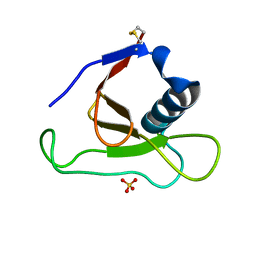 | | Asp79 makes a large, unfavorable contribution to the stability of RNase Sa | | Descriptor: | Guanyl-specific ribonuclease Sa, SULFATE ION | | Authors: | Trevino, S.R, Gokulan, K, Newsom, S, Thurlkill, R.L, Shaw, K.L, Mitkevich, V.A, Makarov, A.A, Sacchettini, J.C, Scholtz, J.M, Pace, C.N. | | Deposit date: | 2005-01-25 | | Release date: | 2005-07-19 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Asp79 Makes a Large, Unfavorable Contribution to the Stability of RNase Sa.
J.Mol.Biol., 354, 2005
|
|
1X1U
 
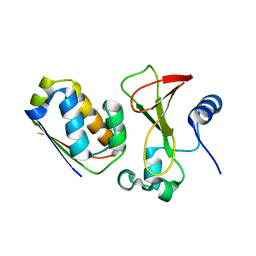 | |
1X1Y
 
 | |
1X1X
 
 | |
1X1W
 
 | |
1T2I
 
 | |
1T2H
 
 | |
1PY3
 
 | | Crystal structure of Ribonuclease Sa2 | | Descriptor: | SULFATE ION, ribonuclease | | Authors: | Sevcik, J, Dauter, Z, Wilson, K.S. | | Deposit date: | 2003-07-08 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure reveals two alternative conformations in the active site of ribonuclease Sa2.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1PYL
 
 | | Crystal structure of Ribonuclease Sa2 | | Descriptor: | SULFATE ION, ribonuclease | | Authors: | Sevcik, J, Dauter, Z, Wilson, K.S. | | Deposit date: | 2003-07-09 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.507 Å) | | Cite: | Crystal structure reveals two alternative conformations in the active site of ribonuclease Sa2.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1R4Y
 
 | | SOLUTION STRUCTURE OF THE DELETION MUTANT DELTA(7-22) OF THE CYTOTOXIC RIBONUCLEASE ALPHA-SARCIN | | Descriptor: | Ribonuclease alpha-sarcin | | Authors: | Garcia-Mayoral, M.F, Garcia-Ortega, L, Lillo, M.P, Santoro, J, Martinez Del Pozo, A, Gavilanes, J.G, Rico, M, Bruix, M. | | Deposit date: | 2003-10-09 | | Release date: | 2004-04-06 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the noncytotoxic {alpha}-sarcin mutant {Delta}(7-22): The importance of the native conformation of peripheral loops for activity.
Protein Sci., 13, 2004
|
|
1Q9E
 
 | | RNase T1 variant with adenine specificity | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Guanyl-specific ribonuclease T1 precursor | | Authors: | Czaja, R, Struhalla, M, Hoeschler, K, Saenger, W, Straeter, N, Hahn, U. | | Deposit date: | 2003-08-25 | | Release date: | 2004-03-23 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | RNase T1 Variant RV Cleaves Single-Stranded RNA after Purines Due to Specific Recognition by the Asn46 Side Chain Amide.
Biochemistry, 43, 2004
|
|
