2ZYE
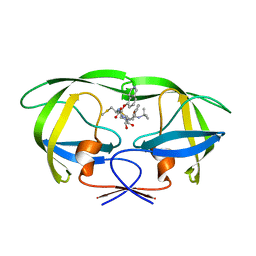 
 | | Structure of HIV-1 Protease in Complex with Potent Inhibitor KNI-272 Determined by Neutron Crystallography | | Descriptor: | (4R)-N-tert-butyl-3-[(2S,3S)-2-hydroxy-3-({N-[(isoquinolin-5-yloxy)acetyl]-S-methyl-L-cysteinyl}amino)-4-phenylbutanoyl]-1,3-thiazolidine-4-carboxamide, protease | | Authors: | Adachi, M, Ohhara, T, Tamada, T, Okazaki, N, Kuroki, R. | | Deposit date: | 2009-01-20 | | Release date: | 2009-03-24 | | Last modified: | 2024-05-29 | | Method: | NEUTRON DIFFRACTION (1.9 Å) | | Cite: | Structure of HIV-1 protease in complex with potent inhibitor KNI-272 determined by high-resolution X-ray and neutron crystallography.
Proc.Natl.Acad.Sci.USA, 2009
|
|
362D
 
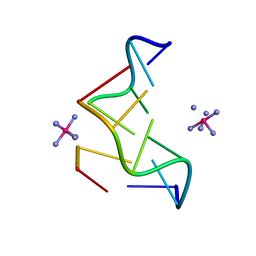 | | THE STRUCTURE OF D(TGCGCA)2 AND A COMPARISON TO OTHER Z-DNA HEXAMERS | | Descriptor: | COBALT HEXAMMINE(III), DNA (5'-D(*TP*GP*CP*GP*CP*A)-3') | | Authors: | Harper, N.A, Brannigan, J.A, Buck, M, Lewis, R.J, Moore, M.H, Schneider, B. | | Deposit date: | 1997-08-20 | | Release date: | 1997-11-20 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structure of d(TGCGCA)2 and a comparison to other DNA hexamers.
Acta Crystallogr.,Sect.D, 54, 1998
|
|
3AAP
 
 | |
3ARO
 
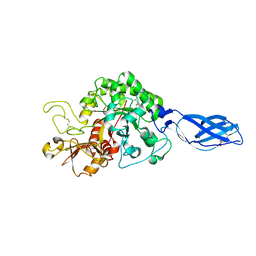 | | Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - apo structure | | Descriptor: | Chitinase A, GLYCEROL | | Authors: | Pantoom, S, Vetter, I.R, Prinz, H, Suginta, W. | | Deposit date: | 2010-12-09 | | Release date: | 2011-04-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Potent family-18 chitinase inhibitors: x-ray structures, affinities, and binding mechanisms
J.Biol.Chem., 286, 2011
|
|
3ARY
 
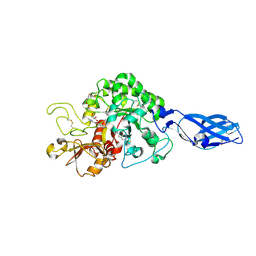 | | Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with 2-(imidazolin-2-yl)-5-isothiocyanatobenzofuran | | Descriptor: | 2-(5-isothiocyanato-1-benzofuran-2-yl)-4,5-dihydro-1H-imidazole, Chitinase A, GLYCEROL | | Authors: | Pantoom, S, Vetter, I.R, Prinz, H, Suginta, W. | | Deposit date: | 2010-12-09 | | Release date: | 2011-04-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Potent family-18 chitinase inhibitors: x-ray structures, affinities, and binding mechanisms
J.Biol.Chem., 286, 2011
|
|
3ARX
 
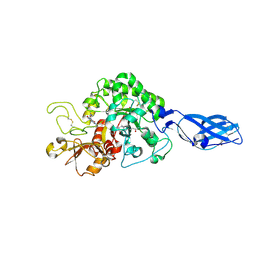 | | Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with Propentofylline | | Descriptor: | 3-methyl-1-(5-oxohexyl)-7-propyl-3,7-dihydro-1H-purine-2,6-dione, Chitinase A, GLYCEROL | | Authors: | Pantoom, S, Vetter, I.R, Prinz, H, Suginta, W. | | Deposit date: | 2010-12-09 | | Release date: | 2011-04-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | Potent family-18 chitinase inhibitors: x-ray structures, affinities, and binding mechanisms
J.Biol.Chem., 286, 2011
|
|
3ASU
 
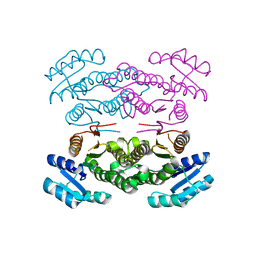 | |
3ARP
 
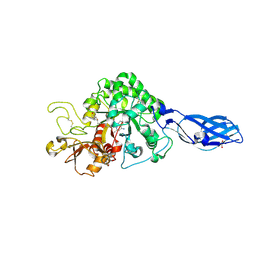 | | Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with DEQUALINIUM | | Descriptor: | Chitinase A, DEQUALINIUM, GLYCEROL | | Authors: | Pantoom, S, Vetter, I.R, Prinz, H, Suginta, W. | | Deposit date: | 2010-12-09 | | Release date: | 2011-04-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Potent family-18 chitinase inhibitors: x-ray structures, affinities, and binding mechanisms
J.Biol.Chem., 286, 2011
|
|
3ARW
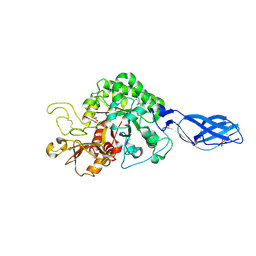 
 | | Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with chelerythrine | | Descriptor: | 1,2-dimethoxy-12-methyl[1,3]benzodioxolo[5,6-c]phenanthridin-12-ium, Chitinase A, GLYCEROL | | Authors: | Pantoom, S, Vetter, I.R, Prinz, H, Suginta, W. | | Deposit date: | 2010-12-09 | | Release date: | 2011-04-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Potent family-18 chitinase inhibitors: x-ray structures, affinities, and binding mechanisms
J.Biol.Chem., 286, 2011
|
|
3AXS
 
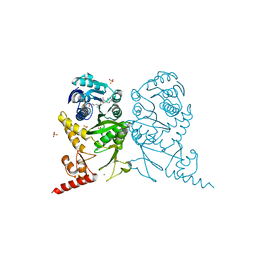 | | Complex structure of tRNA methyltransferase Trm1 from Aquifex aeolicus with sinefungin | | Descriptor: | Probable N(2),N(2)-dimethylguanosine tRNA methyltransferase Trm1, SINEFUNGIN, SULFATE ION, ... | | Authors: | Ihsanawati, Sengoku, T, Yokoyama, S, Bessho, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-04-13 | | Release date: | 2011-08-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.162 Å) | | Cite: | Substrate tRNA recognition mechanism of a multisite-specific tRNA methyltransferase, Aquifex aeolicus Trm1, based on the X-ray crystal structure
J.Biol.Chem., 286, 2011
|
|
3AXT
 
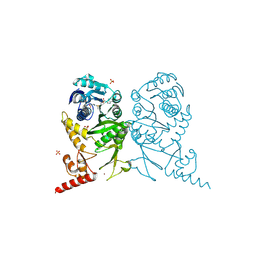 | | Complex structure of tRNA methyltransferase Trm1 from Aquifex aeolicus with S-adenosyl-L-Methionine | | Descriptor: | Probable N(2),N(2)-dimethylguanosine tRNA methyltransferase Trm1, S-ADENOSYLMETHIONINE, SULFATE ION, ... | | Authors: | Ihsanawati, Sengoku, T, Yokoyama, S, Bessho, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-04-14 | | Release date: | 2011-08-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.491 Å) | | Cite: | Substrate tRNA recognition mechanism of a multisite-specific tRNA methyltransferase, Aquifex aeolicus Trm1, based on the X-ray crystal structure
J.Biol.Chem., 286, 2011
|
|
3AY9
 
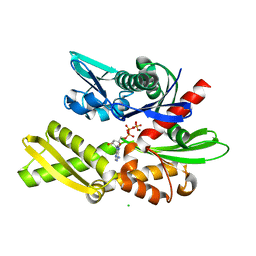 | | Crystal structure of human Hsp70 NBD in the ADP-, Mg ion-, and K ion-bound state | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Arakawa, A, Handa, N, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-05-03 | | Release date: | 2011-12-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Biochemical and structural studies on the high affinity of Hsp70 for ADP.
Protein Sci., 20, 2011
|
|
3AZC
 
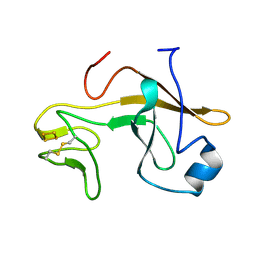 | | Crystal structure of the soluble part of cytochrome b6f complex iron-sulfur subunit from Thermosynechococcus elongatus BP-1 | | Descriptor: | Cytochrome b6-f complex iron-sulfur subunit, FE2/S2 (INORGANIC) CLUSTER | | Authors: | Veit, S, Takeda, K, Tsunoyama, Y, Roegner, M, Miki, K. | | Deposit date: | 2011-05-23 | | Release date: | 2012-05-23 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a thermophilic cyanobacterial b(6)f-type Rieske protein
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3B09
 
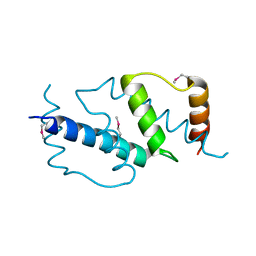 | | Crystal structure of the N-domain of FKBP22 from Shewanella sp. SIB1 | | Descriptor: | Peptidyl-prolyl cis-trans isomerase | | Authors: | Budiman, C, Angkawidjaja, C, Motoike, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-06-07 | | Release date: | 2012-04-18 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of N-domain of FKBP22 from Shewanella sp. SIB1: dimer dissociation by disruption of Val-Leu knot
Protein Sci., 20, 2011
|
|
3B5T
 
 | | Crystal Structure of Novel Immune-Type Receptor 10 Se-Met Extracellular Fragment Mutant N30D | | Descriptor: | Novel immune-type receptor 10 | | Authors: | Ostrov, D.A, Hernandez Prada, J.A, Haire, R.N, Cannon, J.P, Magis, A.T, Bailey, K.M, Litman, G.W. | | Deposit date: | 2007-10-26 | | Release date: | 2008-06-24 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | A bony fish immunological receptor of the NITR multigene family mediates allogeneic recognition.
Immunity, 29, 2008
|
|
3B0K
 
 | | Crystal structure of alpha-lactalbumin | | Descriptor: | Alpha-lactalbumin, CALCIUM ION | | Authors: | Makabe, K. | | Deposit date: | 2011-06-10 | | Release date: | 2012-06-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural insights into the stability perturbations induced by N-terminal variation in human and goat alpha-lactalbumin
Protein Eng.Des.Sel., 26, 2013
|
|
9C10
 
 | | AMG 193, a clinical stage MTA-cooperative PRMT5 inhibitor, drives anti-tumor activity preclinically and in patients with MTAP-deleted cancers | | Descriptor: | (4-amino-1,3-dihydrofuro[3,4-c][1,7]naphthyridin-8-yl){(3S)-3-[4-(trifluoromethyl)phenyl]morpholin-4-yl}methanone, 5'-DEOXY-5'-METHYLTHIOADENOSINE, DIMETHYL SULFOXIDE, ... | | Authors: | Ghimire-Rijal, S, Mukund, S. | | Deposit date: | 2024-05-28 | | Release date: | 2024-10-02 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | AMG 193, a clinical stage MTA-cooperative PRMT5 inhibitor, drives anti-tumor activity preclinically and in patients with MTAP-deleted cancers
To Be Published
|
|
1UWW
 
 | | X-ray crystal structure of a non-crystalline cellulose specific carbohydrate-binding module: CBM28. | | Descriptor: | CALCIUM ION, ENDOGLUCANASE | | Authors: | Jamal, S, Nurizzo, D, Boraston, A, Davies, G.J. | | Deposit date: | 2004-02-12 | | Release date: | 2004-05-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | X-Ray Crystal Structure of a Non-Crystalline Cellulose-Specific Carbohydrate-Binding Module: Cbm28
J.Mol.Biol., 339, 2004
|
|
1V6E
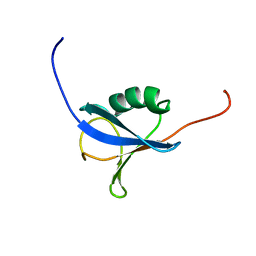 
 | | Solution Structure of a N-terminal Ubiquitin-like Domain in Mouse Tubulin-specific Chaperone B | | Descriptor: | cytoskeleton-associated protein 1 | | Authors: | Zhao, C, Kigawa, T, Saito, K, Koshiba, S, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-11-29 | | Release date: | 2004-12-14 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of a N-terminal Ubiquitin-like Domain in Mouse Tubulin-specific Chaperone B
To be Published
|
|
1V32
 
 | | Solution structure of the SWIB/MDM2 domain of the hypothetical protein At5g08430 from Arabidopsis thaliana | | Descriptor: | hypothetical protein RAFL09-47-K03 | | Authors: | Yoneyama, M, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-10-24 | | Release date: | 2004-04-24 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the SWIB/MDM2 domain of the hypothetical protein At5g08430 from Arabidopsis thaliana
To be Published
|
|
1UUQ
 
 | | Exo-mannosidase from Cellvibrio mixtus | | Descriptor: | GLYCEROL, MANNOSYL-OLIGOSACCHARIDE GLUCOSIDASE, SULFATE ION | | Authors: | Dias, M.V.F, Vincent, F, Pell, G, Prates, J.A.M, Centeno, M.S.J, Ferreira, L.M.A, Gilbert, H.J, Davies, G.J, Fontes, C.M.G.A. | | Deposit date: | 2004-01-09 | | Release date: | 2004-04-16 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Insights Into the Molecular Determinants of Substrate Specificity in Glycoside Hydrolase Family 5 Revealed by the Crystal Structure and Kinetics of Cellvibrio Mixtus Mannosidase 5A
J.Biol.Chem., 279, 2004
|
|
1UP0
 
 | | Structure of the endoglucanase Cel6 from Mycobacterium tuberculosis in complex with cellobiose at 1.75 angstrom | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, ACETATE ION, PUTATIVE CELLULASE CEL6, ... | | Authors: | Varrot, A, Leydier, S, Pell, G, Gilbert, H.J, Davies, G.J. | | Deposit date: | 2003-09-26 | | Release date: | 2004-11-18 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Mycobacterium Tuberculosis Strains Possess Functional Cellulases.
J.Biol.Chem., 280, 2005
|
|
1V88
 
 | | Solution Structure of the Pleckstrin Homology Domain of Oxysterol-Binding Protein-Related Protein 8 (KIAA1451 Protein) | | Descriptor: | Oxysterol binding protein-related protein 8 | | Authors: | Li, H, Tomizawa, T, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-12-29 | | Release date: | 2004-06-29 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Pleckstrin Homology Domain of Oxysterol-Binding Protein-Related Protein 8 (KIAA1451 Protein)
To be Published
|
|
1UPS
 
 | | GlcNAc[alpha]1-4Gal releasing endo-[beta]-galactosidase from Clostridium perfringens | | Descriptor: | CALCIUM ION, GLCNAC-ALPHA-1,4-GAL-RELEASING ENDO-BETA-GALACTOSIDASE | | Authors: | Tempel, W, Liu, Z.-J, Horanyi, P.S, Deng, L, Lee, D, Newton, M.G, Rose, J.P, Ashida, H, Li, S.-C, Li, Y.-T, Wang, B.-C, Southeast Collaboratory for Structural Genomics (SECSG) | | Deposit date: | 2003-10-10 | | Release date: | 2004-11-25 | | Last modified: | 2019-08-21 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Three-dimensional structure of GlcNAcalpha1-4Gal releasing endo-beta-galactosidase from Clostridium perfringens.
Proteins, 59, 2005
|
|
1V1G
 
 | | Structure of the Arabidopsis thaliana SOS3 complexed with Calcium(II) ion | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCINEURIN B-LIKE PROTEIN 4, CALCIUM ION, ... | | Authors: | Sanchez-Barrena, M.J, Martinez-Ripoll, M, Zhu, J.K, Albert, A. | | Deposit date: | 2004-04-15 | | Release date: | 2005-01-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Structure of the Arabidopsis Thaliana SOS3: Molecular Mechanism of Sensing Calcium for Salt Stress Response
J.Mol.Biol., 345, 2005
|
|
