1QSF
 
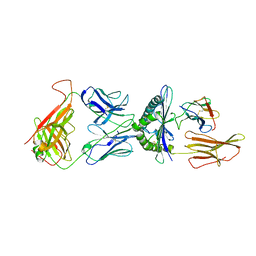 | | STRUCTURE OF A6-TCR BOUND TO HLA-A2 COMPLEXED WITH ALTERED HTLV-1 TAX PEPTIDE Y8A | | Descriptor: | BETA-2 MICROGLOBULIN, HUMAN T-CELL RECEPTOR, MHC CLASS I HLA-A, ... | | Authors: | Ding, Y.H, Baker, B.M, Garboczi, D.N, Biddison, W.E, Wiley, D.C. | | Deposit date: | 1999-06-21 | | Release date: | 1999-12-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Four A6-TCR/peptide/HLA-A2 structures that generate very different T cell signals are nearly identical.
Immunity, 11, 1999
|
|
1W7B
 
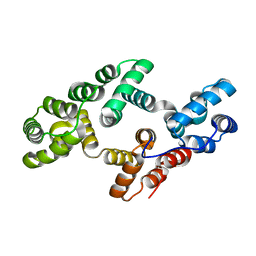 | |
1SV9
 
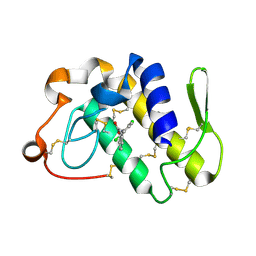 | | Crystal structure of the complex formed between groupII phospholipase A2 and anti-inflammatory agent 2-[(2,6-Dichlorophenyl)amino] benzeneacetic acid at 2.7A resolution | | Descriptor: | 2-[2,6-DICHLOROPHENYL)AMINO]BENZENEACETIC ACID, Phospholipase A2 | | Authors: | Senthil kumar, R, Singh, N, Ethayathulla, A.S, Prem kumar, R, Sharma, S, Singh, T.P. | | Deposit date: | 2004-03-29 | | Release date: | 2004-04-20 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.71 Å) | | Cite: | Crystal structure of the complex formed between group II phospholipase A2 and anti-inflammatory agent 2-[(2,6-Dichlorophenyl)amino] benzeneacetic acid at 2.7A resolution
To be Published
|
|
1DB4
 
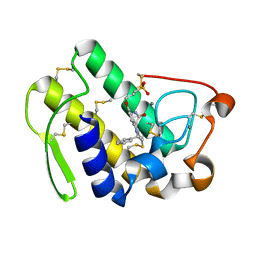 | | HUMAN S-PLA2 IN COMPLEX WITH INDOLE 8 | | Descriptor: | CALCIUM ION, PHOSPHOLIPASE A2, [3-(1-BENZYL-3-CARBAMOYLMETHYL-2-METHYL-1H-INDOL-5-YLOXY)-PROPYL-]-PHOSPHONIC ACID | | Authors: | Chirgadze, N.Y, Schevitz, R.W, Wery, J.-P. | | Deposit date: | 1999-11-02 | | Release date: | 1999-11-12 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-based design of the first potent and selective inhibitor of human non-pancreatic secretory phospholipase A2.
Nat.Struct.Biol., 2, 1995
|
|
2QU9
 
 | | Crystal structure of the complex of group II phospholipase A2 with Eugenol | | Descriptor: | 2-methoxy-4-[(1E)-prop-1-en-1-yl]phenol, Phospholipase A2 VRV-PL-VIIIa, SULFATE ION | | Authors: | Kumar, S, Vikram, G, Singh, N, Sinha, M, Sharma, S, Kaur, P, Srinivasan, A, Singh, T.P. | | Deposit date: | 2007-08-04 | | Release date: | 2007-08-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Crystal structure of the complex of group II phospholipase A2 with Eugenol
To be Published
|
|
1XJL
 
 | |
1EEY
 
 | |
1SV3
 
 | | Structure of the complex formed between Phospholipase A2 and 4-methoxybenzoic acid at 1.3A resolution. | | Descriptor: | 4-METHOXYBENZOIC ACID, Phospholipase A2, SULFATE ION | | Authors: | Singh, N, Prahathees, E, Jabeen, T, Pal, A, Ethayathulla, A.S, Prem kumar, R, Sharma, S, Singh, T.P. | | Deposit date: | 2004-03-27 | | Release date: | 2004-04-13 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal structures of the complexes of a group IIA phospholipase A2 with two natural anti-inflammatory agents, anisic acid, and atropine reveal a similar mode of binding
Proteins, 64, 2006
|
|
3PPX
 | | Crystal structure of the N1602A mutant of an engineered VWF A2 domain (N1493C and C1670S) | | Descriptor: | SODIUM ION, von Willebrand factor | | Authors: | Zhou, M, Dong, X, Zhong, C, Ding, J. | | Deposit date: | 2010-11-25 | | Release date: | 2011-05-04 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | A novel calcium-binding site of von Willebrand factor A2 domain regulates its cleavage by ADAMTS13
Blood, 117, 2011
|
|
1DB5
 
 | | HUMAN S-PLA2 IN COMPLEX WITH INDOLE 6 | | Descriptor: | 4-(1-BENZYL-3-CARBAMOYLMETHYL-2-METHYL-1H-INDOL-5-YLOXY)-BUTYRIC ACID, CALCIUM ION, PROTEIN (PHOSPHOLIPASE A2) | | Authors: | Chirgadze, N.Y, Schevitz, R.W, Wery, J.-P. | | Deposit date: | 1999-11-02 | | Release date: | 1999-11-12 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure-based design of the first potent and selective inhibitor of human non-pancreatic secretory phospholipase A2.
Nat.Struct.Biol., 2, 1995
|
|
1TG4
 
 | | Design of specific inhibitors of groupII phospholipase A2(PLA2): Crystal structure of the complex formed between russells viper PLA2 and designed peptide Phe-Leu-Ala-Tyr-Lys at 1.7A resolution | | Descriptor: | FLAYK peptide, Phospholipase A2, SULFATE ION | | Authors: | Singh, N, Somvanshi, R.K, Sharma, S, Dey, S, Perbandt, M, Betzel, C, Ethayathulla, A.S, Singh, T.P. | | Deposit date: | 2004-05-28 | | Release date: | 2004-06-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Design of specific inhibitors of groupII phospholipase A2(PLA2): Crystal structure of the complex formed between russells viper PLA2 and designed peptide Phe-Leu-Ala-Tyr-Lys at 1.7A resolution
TO BE PUBLISHED
|
|
1KP4
 
 | |
1YXL
 
 | | Crystal structure of a novel phospholipase A2 from Naja naja sagittifera at 1.5 A resolution | | Descriptor: | ACETIC ACID, CALCIUM ION, PHOSPHATE ION, ... | | Authors: | Singh, R.K, Jabeen, T, Sharma, S, Kaur, P, Singh, T.P. | | Deposit date: | 2005-02-22 | | Release date: | 2005-03-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.477 Å) | | Cite: | Crystal Structure of a novel phospholipase A2 from Naja naja sagittifera at 1.5 A resolution
To be Published
|
|
1IJL
 
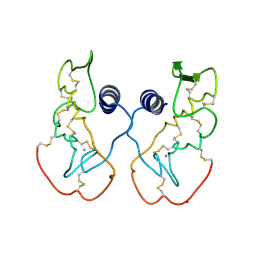 | | Crystal structure of acidic phospholipase A2 from deinagkistrodon acutus | | Descriptor: | CALCIUM ION, PHOSPHOLIPASE A2, ZINC ION | | Authors: | Gu, L, Zhang, H, Song, S, Zhou, Y, Lin, Z. | | Deposit date: | 2001-04-27 | | Release date: | 2001-12-28 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of an acidic phospholipase A2 from the venom of Deinagkistrodon acutus.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
3PPV
 | | Crystal structure of an engineered VWF A2 domain (N1493C and C1670S) | | Descriptor: | CALCIUM ION, SULFATE ION, von Willebrand factor | | Authors: | Zhou, M, Dong, X, Zhong, C, Ding, J. | | Deposit date: | 2010-11-25 | | Release date: | 2011-05-04 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A novel calcium-binding site of von Willebrand factor A2 domain regulates its cleavage by ADAMTS13
Blood, 117, 2011
|
|
9JHY
 
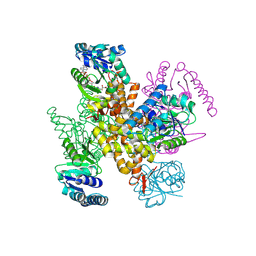 | | 3-Hydroxybutyryl-CoA dehydrogenase mutant (S117A) with acetoacetyl CoA | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain protein, ACETOACETYL-COENZYME A | | Authors: | Yang, J.W, Jeon, H.J, Park, S.H, Jang, S.H, Park, J.A, Kim, S.H, Hwang, K.Y. | | Deposit date: | 2024-09-10 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Insights and Catalytic Mechanism of 3-Hydroxybutyryl-CoA Dehydrogenase from Faecalibacterium Prausnitzii A2-165.
Int J Mol Sci, 25, 2024
|
|
9JI0
 
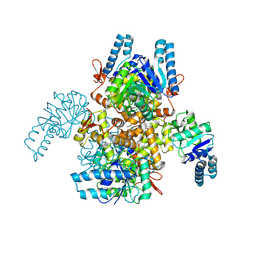 | | 3-Hydroxybutyryl-CoA dehydrogenase | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain protein | | Authors: | Yang, J.W, Jeon, H.J, Park, S.H, Jang, S.H, Park, J.A, Kim, S.H, Hwang, K.Y. | | Deposit date: | 2024-09-10 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Structural Insights and Catalytic Mechanism of 3-Hydroxybutyryl-CoA Dehydrogenase from Faecalibacterium Prausnitzii A2-165.
Int J Mol Sci, 25, 2024
|
|
9JHE
 
 | | 3-hydroxybutyryl-CoA dehydrogenase with NAD | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain protein, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Park, J.A, Yang, J.W, Park, S.H, Kim, S.H, Hwang, K.Y. | | Deposit date: | 2024-09-09 | | Release date: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural Insights and Catalytic Mechanism of 3-Hydroxybutyryl-CoA Dehydrogenase from Faecalibacterium Prausnitzii A2-165.
Int J Mol Sci, 25, 2024
|
|
9JHZ
 
 | | 3-Hydroxybutyryl-CoA dehydrogenase mutant(S117A) with acetoacetyl CoA and NAD | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain protein, ACETOACETYL-COENZYME A, ... | | Authors: | Yang, J.W, Jeon, H.J, Park, S.H, Kim, S.H, Hwang, K.Y. | | Deposit date: | 2024-09-10 | | Release date: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Insights and Catalytic Mechanism of 3-Hydroxybutyryl-CoA Dehydrogenase from Faecalibacterium Prausnitzii A2-165.
Int J Mol Sci, 25, 2024
|
|
7N6E
 
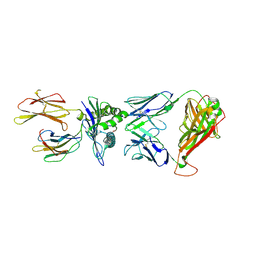 | | TCR peptide HLA-A2 complex | | Descriptor: | Beta-2-microglobulin, MHC class I antigen, Spike protein S1, ... | | Authors: | Chaurasia, P, Rossjohn, J, Petersen, J. | | Deposit date: | 2021-06-08 | | Release date: | 2021-07-28 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural basis of biased T cell receptor recognition of an immunodominant HLA-A2 epitope of the SARS-CoV-2 spike protein.
J.Biol.Chem., 297, 2021
|
|
4E5X
 
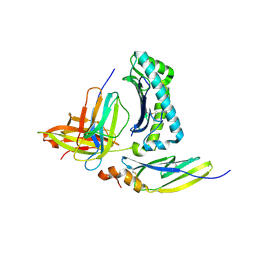 | |
1TG1
 
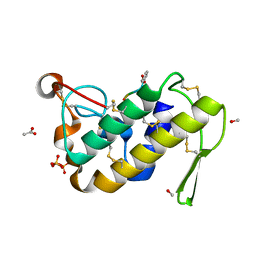 | | Crystal Structure of the complex formed between russells viper phospholipase A2 and a designed peptide inhibitor PHQ-Leu-Val-Arg-Tyr at 1.2A resolution | | Descriptor: | ACETIC ACID, METHANOL, Phospholipase A2, ... | | Authors: | Singh, N, Kaur, P, Somvanshi, R.K, Sharma, S, Dey, S, Perbandt, M, Betzel, C, Singh, T.P. | | Deposit date: | 2004-05-28 | | Release date: | 2004-06-08 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Crystal Structure of the complex formed between russells viper phospholipase A2 and a designed peptide inhibitor Cbz-dehydro-Leu-Val-Arg-Tyr at 1.2A resolution
To be Published
|
|
1SZ8
 
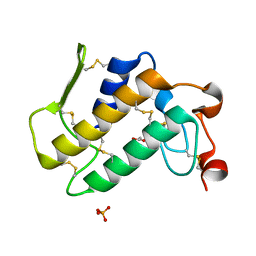 | | Crystal Structure of an Acidic Phospholipase A2 from Naja Naja Sagittifera at 1.5 A resolution | | Descriptor: | ACETIC ACID, CALCIUM ION, PHOSPHATE ION, ... | | Authors: | Singh, R.K, Sharma, S, Jabeen, T, Kaur, P, Singh, T.P. | | Deposit date: | 2004-04-05 | | Release date: | 2004-04-20 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of an Acidic Phospholipase A2 from Naja Naja Sagittifera at 1.5 A Resolution
To be Published
|
|
1TDV
 
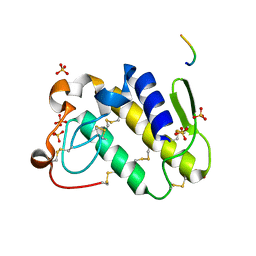 | | Non-specific binding to phospholipase A2:Crystal structure of the complex of PLA2 with a designed peptide Tyr-Trp-Ala-Ala-Ala-Ala at 1.7A resolution | | Descriptor: | Phospholipase A2 VRV-PL-VIIIa, SULFATE ION, YWAAAA | | Authors: | Singh, N, Jabeen, T, Ethayathulla, A.S, Somvanshi, R.K, Sharma, S, Dey, S, Perbandt, M, Betzel, C, Singh, T.P. | | Deposit date: | 2004-05-24 | | Release date: | 2004-06-08 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Non-specific binding to phospholipase A2:Crystal structure of the complex of PLA2 with a designed peptide Tyr-Trp-Ala-Ala-Ala-Ala at 1.7A resolution
to be published
|
|
1TK4
 
 | | Crystal structure of russells viper phospholipase A2 in complex with a specifically designed tetrapeptide Ala-Ile-Arg-Ser at 1.1 A resolution | | Descriptor: | Phospholipase A2 VRV-PL-VIIIa, SULFATE ION, Tetrapeptide Ala-Ile-Arg-Ser | | Authors: | Singh, N, Bilgrami, S, Somvanshi, R.K, Sharma, S, Dey, S, Perbandt, M, Betzel, C, Kaur, P, Singh, T.P. | | Deposit date: | 2004-06-08 | | Release date: | 2004-06-22 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal structure of russells viper phospholipase A2 with a specifically designed tetrapeptide Ala-Ile-Arg-Ser at 1.1 A resolution
TO BE PUBLISHED
|
|
