1QRS
 
 | |
1QRT
 
 | |
1QRU
 
 | |
1QRV
 
 | | CRYSTAL STRUCTURE OF THE COMPLEX OF HMG-D AND DNA | | Descriptor: | DNA (5'-D(*GP*CP*GP*AP*TP*AP*TP*CP*GP*C)-3'), HIGH MOBILITY GROUP PROTEIN D, SODIUM ION | | Authors: | Murphy IV, F.V, Sweet, R.M, Churchill, M.E.A. | | Deposit date: | 1999-06-15 | | Release date: | 1999-12-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The structure of a chromosomal high mobility group protein-DNA complex reveals sequence-neutral mechanisms important for non-sequence-specific DNA recognition.
EMBO J., 18, 1999
|
|
1QRW
 
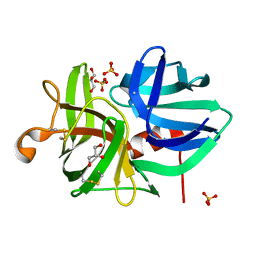 | | CRYSTAL STRUCTURE OF AN ALPHA-LYTIC PROTEASE MUTANT WITH ACCELERATED FOLDING KINETICS, R102H/G134S, PH 8 | | Descriptor: | ALHPA-LYTIC PROTEASE, GLYCEROL, SULFATE ION | | Authors: | Derman, A.I, Mau, T, Agard, D.A. | | Deposit date: | 1999-06-16 | | Release date: | 1999-06-18 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | A Genetic Screen That Targets Specifically the Folding Transition State of Alpha-Lytic Protease
To be Published, 1999
|
|
1QRX
 
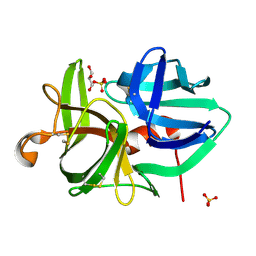 | |
1QRY
 
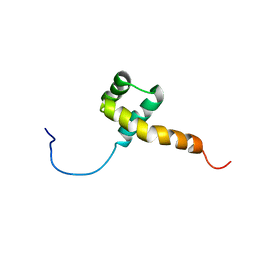 | | Homeobox protein VND (ventral nervous system defective protein) | | Descriptor: | PROTEIN (HOMEOBOX VENTRAL NERVOUS SYSTEM DEFECTIVE PROTEIN) | | Authors: | Xiang, B. | | Deposit date: | 1999-06-16 | | Release date: | 1999-07-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Distortion of the three-dimensional structure of the vnd/NK-2 homeodomain bound to DNA induced by an embryonically lethal A35T point mutation.
Biochemistry, 42, 2003
|
|
1QRZ
 
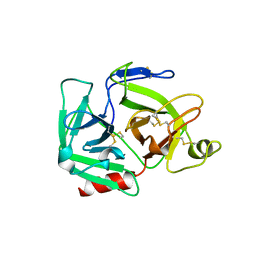 | | CATALYTIC DOMAIN OF PLASMINOGEN | | Descriptor: | PLASMINOGEN | | Authors: | Peisach, E, Wang, J, de los Santos, T, Reich, E, Ringe, D. | | Deposit date: | 1999-06-16 | | Release date: | 1999-10-14 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of the proenzyme domain of plasminogen.
Biochemistry, 38, 1999
|
|
1QS0
 
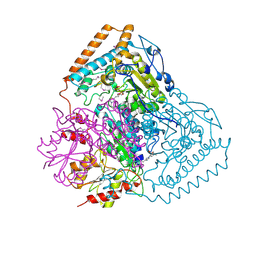 | | Crystal Structure of Pseudomonas Putida 2-oxoisovalerate Dehydrogenase (Branched-Chain Alpha-Keto Acid Dehydrogenase, E1B) | | Descriptor: | 2-OXO-4-METHYLPENTANOIC ACID, 2-OXOISOVALERATE DEHYDROGENASE ALPHA-SUBUNIT, 2-OXOISOVALERATE DEHYDROGENASE BETA-SUBUNIT, ... | | Authors: | Aevarsson, A, Seger, K, Turley, S, Sokatch, J.R, Hol, W.G.J. | | Deposit date: | 1999-06-24 | | Release date: | 1999-08-18 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of 2-oxoisovalerate and dehydrogenase and the architecture of 2-oxo acid dehydrogenase multienzyme complexes.
Nat.Struct.Biol., 6, 1999
|
|
1QS1
 
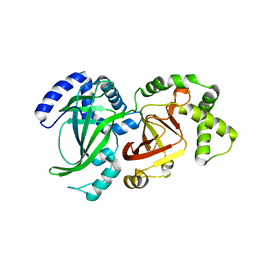 | | CRYSTAL STRUCTURE OF VEGETATIVE INSECTICIDAL PROTEIN2 (VIP2) | | Descriptor: | ADP-RIBOSYLTRANSFERASE | | Authors: | Han, S, Craig, J.A, Putnam, C.D, Carozzi, N.B, Tainer, J.A. | | Deposit date: | 1999-06-25 | | Release date: | 1999-12-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Evolution and mechanism from structures of an ADP-ribosylating toxin and NAD complex.
Nat.Struct.Biol., 6, 1999
|
|
1QS2
 
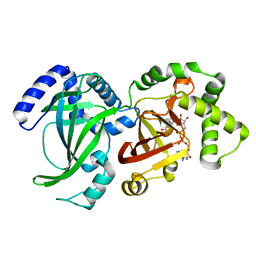 | | CRYSTAL STRUCTURE OF VIP2 WITH NAD | | Descriptor: | ADP-RIBOSYLTRANSFERASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Han, S, Craig, J.A, Putnam, C.D, Carozzi, N.B, Tainer, J.A. | | Deposit date: | 1999-06-25 | | Release date: | 1999-12-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Evolution and mechanism from structures of an ADP-ribosylating toxin and NAD complex.
Nat.Struct.Biol., 6, 1999
|
|
1QS3
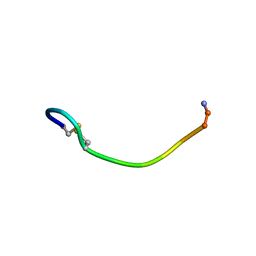 
 | | NMR SOLUTION CONFORMATION OF AN ANTITOXIC ANALOG OF ALPHA-CONOTOXIN GI | | Descriptor: | DES-GLU1-[CYS3ALA]-DES-CYS13-ALPHA CONOTOXIN GI | | Authors: | Mok, K.H, Han, K.-H. | | Deposit date: | 1999-06-25 | | Release date: | 1999-10-06 | | Last modified: | 2014-03-12 | | Method: | SOLUTION NMR | | Cite: | NMR solution conformation of an antitoxic analogue of alpha-conotoxin GI: identification of a common nicotinic acetylcholine receptor alpha 1-subunit binding surface for small ligands and alpha-conotoxins.
Biochemistry, 38, 1999
|
|
1QS4
 
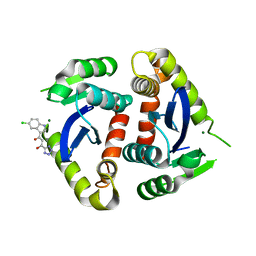 | | Core domain of HIV-1 integrase complexed with Mg++ and 1-(5-chloroindol-3-yl)-3-hydroxy-3-(2H-tetrazol-5-yl)-propenone | | Descriptor: | 1-(5-CHLOROINDOL-3-YL)-3-HYDROXY-3-(2H-TETRAZOL-5-YL)-PROPENONE, MAGNESIUM ION, PROTEIN (HIV-1 INTEGRASE (E.C.2.7.7.49)) | | Authors: | Goldgur, Y, Craigie, R, Fujiwara, T, Yoshinaga, T, Davies, D.R. | | Deposit date: | 1999-06-25 | | Release date: | 1999-11-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of the HIV-1 integrase catalytic domain complexed with an inhibitor: a platform for antiviral drug design.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
1QS5
 
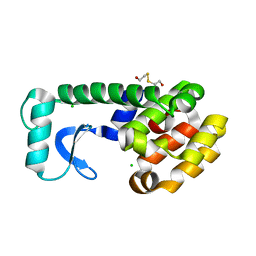 | |
1QS7
 
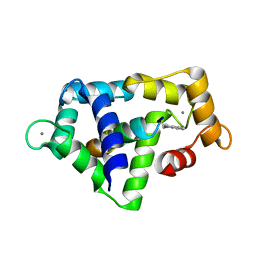 | |
1QS8
 
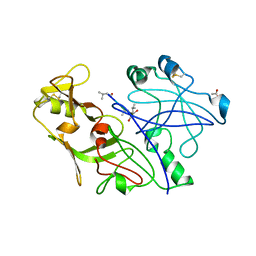 | | Crystal structure of the P. vivax aspartic proteinase plasmepsin complexed with the inhibitor pepstatin A | | Descriptor: | ACETATE ION, PEPSTATIN A, PLASMEPSIN | | Authors: | Khazanovich Bernstein, N, Cherney, M.M, Yowell, C.A, Dame, J.B, James, M.N.G. | | Deposit date: | 1999-06-25 | | Release date: | 1999-07-07 | | Last modified: | 2013-02-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into the activation of P. vivax plasmepsin.
J.Mol.Biol., 329, 2003
|
|
1QS9
 
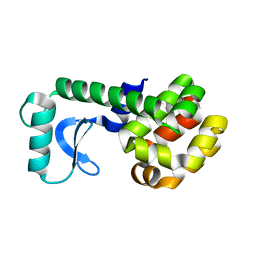 | |
1QSA
 
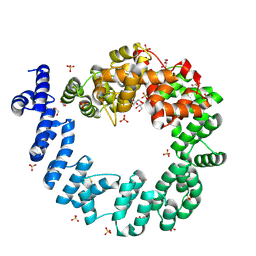 | | CRYSTAL STRUCTURE OF THE 70 KDA SOLUBLE LYTIC TRANSGLYCOSYLASE SLT70 FROM ESCHERICHIA COLI AT 1.65 ANGSTROMS RESOLUTION | | Descriptor: | ACETATE ION, GLYCEROL, PROTEIN (SOLUBLE LYTIC TRANSGLYCOSYLASE SLT70), ... | | Authors: | van Asselt, E.J, Thunnissen, A.-M.W.H, Dijkstra, B.W. | | Deposit date: | 1999-06-20 | | Release date: | 1999-09-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | High resolution crystal structures of the Escherichia coli lytic transglycosylase Slt70 and its complex with a peptidoglycan fragment.
J.Mol.Biol., 291, 1999
|
|
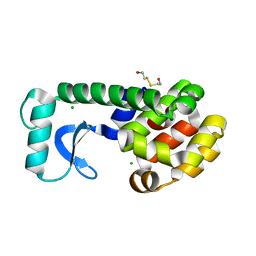
1QSB
 
 | |
1QSC
 
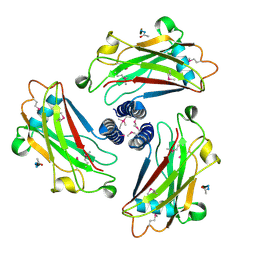 | | CRYSTAL STRUCTURE OF THE TRAF DOMAIN OF TRAF2 IN A COMPLEX WITH A PEPTIDE FROM THE CD40 RECEPTOR | | Descriptor: | CD40 RECEPTOR, TNF RECEPTOR ASSOCIATED FACTOR 2 | | Authors: | McWhirter, S.M, Pullen, S.S, Holton, J.M, Crute, J.J, Kehry, M.R, Alber, T. | | Deposit date: | 1999-06-20 | | Release date: | 1999-08-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystallographic analysis of CD40 recognition and signaling by human TRAF2.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
1QSD
 
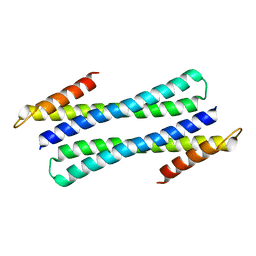 | | RBL2P, BETA-TUBULIN BINDING POST-CHAPERONIN COFACTOR | | Descriptor: | PROTEIN (BETA-TUBULIN BINDING POST-CHAPERONIN COFACTOR) | | Authors: | Steinbacher, S. | | Deposit date: | 1999-06-21 | | Release date: | 1999-12-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the post-chaperonin beta-tubulin binding cofactor Rbl2p
Nat.Struct.Biol., 6, 1999
|
|
1QSE
 
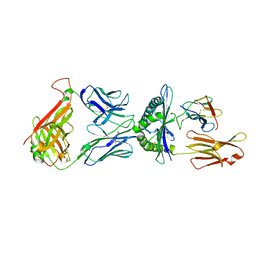 | | STRUCTURE OF HUMAN A6-TCR BOUND TO HLA-A2 COMPLEXED WITH ALTERED HTLV-1 TAX PEPTIDE V7R | | Descriptor: | PROTEIN (MHC class I HLA-A), PROTEIN (beta-2 microglobulin), PROTEIN (human T-Cell receptor), ... | | Authors: | Ding, Y.H, Baker, B.M, Garboczi, D.N, Biddison, W.E, Wiley, D.C. | | Deposit date: | 1999-06-21 | | Release date: | 1999-12-21 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Four A6-TCR/peptide/HLA-A2 structures that generate very different T cell signals are nearly identical.
Immunity, 11, 1999
|
|
1QSF
 
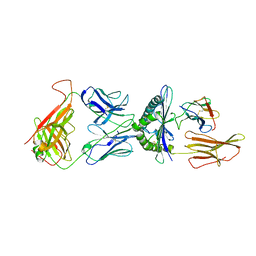 | | STRUCTURE OF A6-TCR BOUND TO HLA-A2 COMPLEXED WITH ALTERED HTLV-1 TAX PEPTIDE Y8A | | Descriptor: | BETA-2 MICROGLOBULIN, HUMAN T-CELL RECEPTOR, MHC CLASS I HLA-A, ... | | Authors: | Ding, Y.H, Baker, B.M, Garboczi, D.N, Biddison, W.E, Wiley, D.C. | | Deposit date: | 1999-06-21 | | Release date: | 1999-12-21 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Four A6-TCR/peptide/HLA-A2 structures that generate very different T cell signals are nearly identical.
Immunity, 11, 1999
|
|
1QSG
 
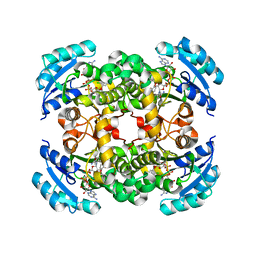 | | CRYSTAL STRUCTURE OF ENOYL REDUCTASE INHIBITION BY TRICLOSAN | | Descriptor: | ENOYL-[ACYL-CARRIER-PROTEIN] REDUCTASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, TRICLOSAN, ... | | Authors: | Stewart, M.J, Parikh, S, Xiao, G, Tonge, P.J, Kisker, C. | | Deposit date: | 1999-06-21 | | Release date: | 1999-07-21 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural basis and mechanism of enoyl reductase inhibition by triclosan.
J.Mol.Biol., 290, 1999
|
|
1QSH
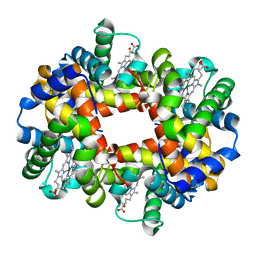 
 | | MAGNESIUM(II)-AND ZINC(II)-PROTOPORPHYRIN IX'S STABILIZE THE LOWEST OXYGEN AFFINITY STATE OF HUMAN HEMOGLOBIN EVEN MORE STRONGLY THAN DEOXYHEME | | Descriptor: | PROTEIN (HEMOGLOBIN ALPHA CHAIN), PROTEIN (HEMOGLOBIN BETA CHAIN), PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Miyazaki, G, Morimoto, H, Yun, K.-M, Park, S.-Y, Nakagawa, A, Minagawa, H, Shibayama, N. | | Deposit date: | 1999-06-22 | | Release date: | 1999-07-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Magnesium(II) and zinc(II)-protoporphyrin IX's stabilize the lowest oxygen affinity state of human hemoglobin even more strongly than deoxyheme.
J.Mol.Biol., 292, 1999
|
|
