8Y6U
 
 | |
1C06
 
 | | SOLUTION STRUCTURE OF RIBOSOMAL PROTEIN S4 DELTA 41, REFINED WITH DIPOLAR COUPLINGS (ENSEMBLE OF 16 STRUCTURES) | | Descriptor: | RIBOSOMAL PROTEIN S4 DELTA 41 | | Authors: | Markus, M.A, Gerstner, R.B, Draper, D.E, Torchia, D.A. | | Deposit date: | 1999-07-14 | | Release date: | 1999-09-29 | | Last modified: | 2024-04-10 | | Method: | SOLUTION NMR | | Cite: | Refining the overall structure and subdomain orientation of ribosomal protein S4 delta41 with dipolar couplings measured by NMR in uniaxial liquid crystalline phases.
J.Mol.Biol., 292, 1999
|
|
1C05
 
 | | SOLUTION STRUCTURE OF RIBOSOMAL PROTEIN S4 DELTA 41, REFINED WITH DIPOLAR COUPLINGS (MINIMIZED AVERAGE STRUCTURE) | | Descriptor: | RIBOSOMAL PROTEIN S4 DELTA 41 | | Authors: | Markus, M.A, Gerstner, R.B, Draper, D.E, Torchia, D.A. | | Deposit date: | 1999-07-14 | | Release date: | 1999-09-29 | | Last modified: | 2024-04-10 | | Method: | SOLUTION NMR | | Cite: | Refining the overall structure and subdomain orientation of ribosomal protein S4 delta41 with dipolar couplings measured by NMR in uniaxial liquid crystalline phases.
J.Mol.Biol., 292, 1999
|
|
1SP7
 
 | | Structure of the Cys-rich C-terminal domain of Hydra minicollagen | | Descriptor: | mini-collagen | | Authors: | Meier, S, Haussinger, D, Pokidysheva, E, Bachinger, H.P, Grzesiek, S. | | Deposit date: | 2004-03-16 | | Release date: | 2004-05-18 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Determination of a high-precision NMR structure of the minicollagen cysteine rich domain from Hydra and characterization of its disulfide bond formation.
Febs Lett., 569, 2004
|
|
1E08
 
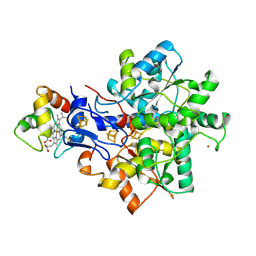 | | Structural model of the [Fe]-Hydrogenase/cytochrome c553 complex combining NMR and soft-docking | | Descriptor: | 1,3-PROPANEDITHIOL, CARBON MONOXIDE, CYANIDE ION, ... | | Authors: | Morelli, X, Czjzek, M, Hatchikian, C.E, Bornet, O, Fontecilla-Camps, J.C, Palma, N.P, Moura, J.J.G, Guerlesquin, F. | | Deposit date: | 2000-03-13 | | Release date: | 2000-08-25 | | Last modified: | 2019-11-27 | | Method: | SOLUTION NMR, THEORETICAL MODEL | | Cite: | Structural Model of the Fe-Hydrogenase/Cytochrome C553 Complex Combining Transverse Relaxation-Optimized Spectroscopy Experiments and Soft Docking Calculations.
J.Biol.Chem., 275, 2000
|
|
1UCV
 
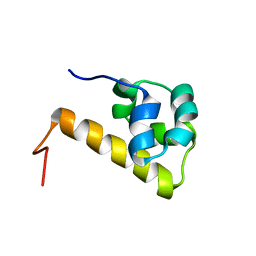 | | Sterile alpha motif (SAM) domain of ephrin type-A receptor 8 | | Descriptor: | EPHRIN TYPE-A RECEPTOR 8 | | Authors: | Goroncy, A, Kigawa, T, Koshiba, S, Kobayashi, N, Tochio, N, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-04-23 | | Release date: | 2004-05-11 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of Sterile alpha motif (SAM) domain of ephrin type-A receptor 8
To be Published
|
|
1ROT
 
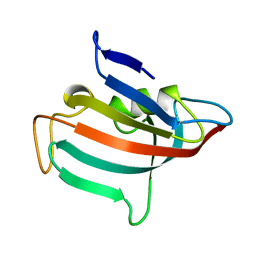 | | STRUCTURE OF FKBP59-I, THE N-TERMINAL DOMAIN OF A 59 KDA FK506-BINDING PROTEIN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | FKBP59-I | | Authors: | Craescu, C.T, Rouviere, N, Popescu, A, Cerpolini, E, Lebeau, M.-C, Baulieu, E.-E, Mispelter, J. | | Deposit date: | 1996-06-14 | | Release date: | 1996-12-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional structure of the immunophilin-like domain of FKBP59 in solution.
Biochemistry, 35, 1996
|
|
1ROU
 
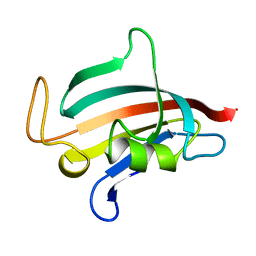 | | STRUCTURE OF FKBP59-I, THE N-TERMINAL DOMAIN OF A 59 KDA FK506-BINDING PROTEIN, NMR, 22 STRUCTURES | | Descriptor: | FKBP59-I | | Authors: | Craescu, C.T, Rouviere, N, Popescu, A, Cerpolini, E, Lebeau, M.-C, Baulieu, E.-E, Mispelter, J. | | Deposit date: | 1996-06-14 | | Release date: | 1996-12-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional structure of the immunophilin-like domain of FKBP59 in solution.
Biochemistry, 35, 1996
|
|
1TVC
 
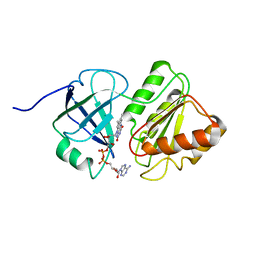 | | FAD and NADH binding domain of methane monooxygenase reductase from Methylococcus capsulatus (Bath) | | Descriptor: | DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, METHANE MONOOXYGENASE COMPONENT C | | Authors: | Chatwood, L.L, Mueller, J, Gross, J.D, Wagner, G, Lippard, S.J. | | Deposit date: | 2004-06-29 | | Release date: | 2004-10-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Flavin Domain from Soluble Methane Monooxygenase Reductase from Methylococcus capsulatus (Bath)
Biochemistry, 43, 2004
|
|
1UG1
 
 | | SH3 domain of Hypothetical protein BAA76854.1 | | Descriptor: | KIAA1010 protein | | Authors: | Nagata, T, Muto, Y, Kamewari, Y, Shirouzu, M, Terada, T, Kigawa, T, Inoue, M, Yabuki, T, Aoki, M, Seki, E, Matsuda, T, Hirota, H, Yoshida, M, Kobayashi, N, Tanaka, A, Osanai, T, Matsuo, Y, Ohara, O, Nagase, T, Kikuno, R, Nakayama, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-06-11 | | Release date: | 2003-12-11 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR structure of SH3 domain of Hypothetical protein BAA76854.1
To be Published
|
|
1MYF
 
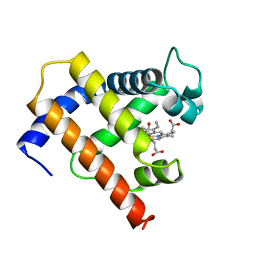 | | SOLUTION STRUCTURE OF CARBONMONOXY MYOGLOBIN DETERMINED FROM NMR DISTANCE AND CHEMICAL SHIFT CONSTRAINTS | | Descriptor: | CARBON MONOXIDE, MYOGLOBIN, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Osapay, K, Theriault, Y, Wright, P.E, Case, D.A. | | Deposit date: | 1994-12-02 | | Release date: | 1995-02-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of carbonmonoxy myoglobin determined from nuclear magnetic resonance distance and chemical shift constraints.
J.Mol.Biol., 244, 1994
|
|
1JHI
 
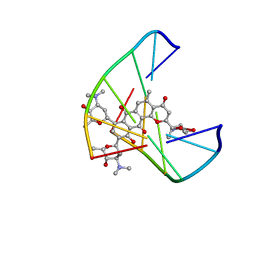 | | Solution Structure of a Hedamycin-DNA complex | | Descriptor: | 5'-D(*AP*CP*CP*(HEH)GP*GP*T)-3', HEDAMYCIN | | Authors: | Owen, E.A, Burley, G.A, Carver, J.A, Wickham, G, Keniry, M.A. | | Deposit date: | 2001-06-27 | | Release date: | 2003-07-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural investigation of the hedamycin:d(ACCGGT)2 complex by NMR and restrained molecular dynamics.
Biochem.Biophys.Res.Commun., 290, 2002
|
|
1JRJ
 
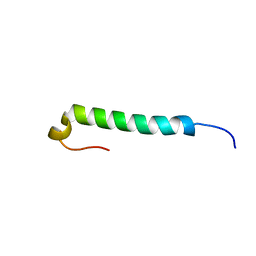 | |
1SNJ
 
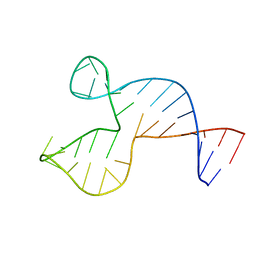 | | Solution structure of the DNA three-way junction with the A/C-stacked conformation | | Descriptor: | 36-MER | | Authors: | Wu, B, Girard, F, van Buuren, B, Schleucher, J, Tessari, M, Wijmenga, S. | | Deposit date: | 2004-03-11 | | Release date: | 2005-04-05 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Global structure of a DNA three-way junction by solution NMR: towards prediction of 3H fold.
Nucleic Acids Res., 32, 2004
|
|
1S24
 | | Rubredoxin domain II from Pseudomonas oleovorans | | Descriptor: | CADMIUM ION, Rubredoxin 2 | | Authors: | Perry, A, Tambyrajah, W, Grossmann, J.G, Lian, L.Y, Scrutton, N.S. | | Deposit date: | 2004-01-08 | | Release date: | 2004-05-04 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the two-iron rubredoxin of Pseudomonas oleovorans determined by NMR spectroscopy and solution X-ray scattering and interactions with rubredoxin reductase.
Biochemistry, 43, 2004
|
|
6SZF
 
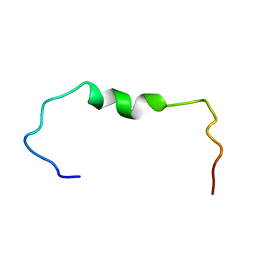 | | Solution structure of the amyloid beta-peptide (1-42) | | Descriptor: | Amyloid-beta precursor protein | | Authors: | Grimaldi, M, Santoro, A, Stillitano, I, Buonocore, M, D'Ursi, A.M. | | Deposit date: | 2019-10-02 | | Release date: | 2020-10-14 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Exploring the Early Stages of the Amyloid A beta (1-42) Peptide Aggregation Process: An NMR Study.
Pharmaceuticals, 14, 2021
|
|
6TWR
 
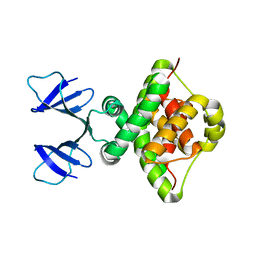 | |
3IHZ
 
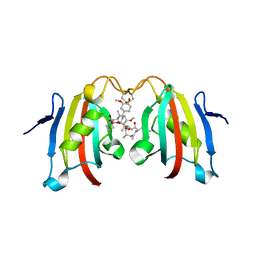 | | Crystal structure of the FK506 binding domain of Plasmodium vivax FKBP35 in complex with FK506 | | Descriptor: | 70 kDa peptidylprolyl isomerase, putative, 8-DEETHYL-8-[BUT-3-ENYL]-ASCOMYCIN | | Authors: | Qureshi, I.A, Alag, R, Yoon, H.S, Lescar, J. | | Deposit date: | 2009-07-31 | | Release date: | 2010-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | NMR and crystallographic structures of the FK506 binding domain of human malarial parasite Plasmodium vivax FKBP35
Protein Sci., 19, 2010
|
|
1ULP
 
 | |
1UUB
 | |
1UUA
 | |
7CAP
 
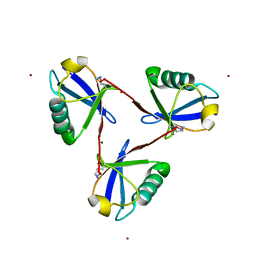 | | Cyclic Lys48-linked triubiquitin | | Descriptor: | Ubiquitin, ZINC ION | | Authors: | Hiranyakorn, M, Yanaka, S, Satoh, T, Wilasri, T, Jityuti, B, Yagi-Utsumi, T, Kato, K. | | Deposit date: | 2020-06-09 | | Release date: | 2020-08-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | NMR Characterization of Conformational Interconversions of Lys48-Linked Ubiquitin Chains.
Int J Mol Sci, 21, 2020
|
|
1NP9
 
 | |
6YE5
 
 | | Structure of ribosomal binding factor A RbfA of Staphylococcus aureus bacterium by NMR | | Descriptor: | Ribosome-binding factor A | | Authors: | Blokhin, D.S, Usachev, K.S, Bikmullin, A.G, Nurullina, L, Garaeva, N, Validov, S, Klochkov, V, Aganov, A, Khusainov, I, Yusupov, M. | | Deposit date: | 2020-03-24 | | Release date: | 2021-03-31 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structure of ribosomal binding factor A RbfA of Staphylococcus aureus bacterium by NMR
To Be Published
|
|
1JUA
 | |
