7MQ1
 
 | | C9A Streptococcus pneumoniae CstR in the reduced state, space group C2 | | Descriptor: | CHLORIDE ION, Copper-sensing transcriptional repressor csoR, GLYCEROL, ... | | Authors: | Fakhoury, J.N, Gonzalez-Gutierrez, G, Giedroc, D.P. | | Deposit date: | 2021-05-05 | | Release date: | 2022-03-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Functional asymmetry and chemical reactivity of CsoR family persulfide sensors.
Nucleic Acids Res., 49, 2021
|
|
5GGP
 
 | | Crystal structure of N-terminal domain of human protein O-mannose beta-1,2-N-acetylglucosaminyltransferase in complex with GlcNAc-beta1,2-Man-peptide | | Descriptor: | 10-mer Peptide from Dystroglycan, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose, Protein O-linked-mannose beta-1,2-N-acetylglucosaminyltransferase 1, ... | | Authors: | Kuwabara, N, Senda, T, Kato, R. | | Deposit date: | 2016-06-16 | | Release date: | 2016-08-10 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.599 Å) | | Cite: | Carbohydrate-binding domain of the POMGnT1 stem region modulates O-mannosylation sites of alpha-dystroglycan
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
8U1A
 
 | |
7UK6
 
 | |
8QDI
 
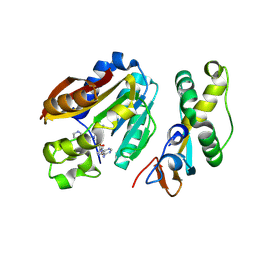 | | compound 1b bound KMT9 crystal structure | | Descriptor: | (2~{S})-4-[[(2~{R},3~{S},4~{R},5~{R})-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methyl-[2-[(2~{R})-pyrrolidin-2-yl]ethyl]amino]-2-azanyl-butanoic acid, Methyltransferase N6AMT1, Multifunctional methyltransferase subunit TRM112-like protein | | Authors: | Sheng, W, Eric, M, Roland, S. | | Deposit date: | 2023-08-29 | | Release date: | 2024-09-11 | | Method: | X-RAY DIFFRACTION (1.467 Å) | | Cite: | compound 1a bound KMT9 crystal structure
To Be Published
|
|
8JRC
 
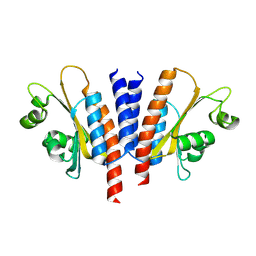 | |
7UK8
 
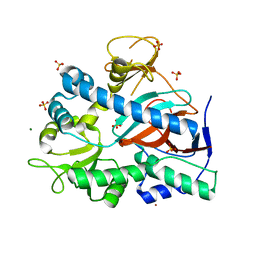 | | Apo form of YgiC from Escherichia coli K-12 | | Descriptor: | MAGNESIUM ION, NICKEL (II) ION, Putative acid--amine ligase YgiC, ... | | Authors: | Pederick, J.L, Bruning, J.B. | | Deposit date: | 2022-03-31 | | Release date: | 2023-02-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Escherichia coli YgiC and YjfC Possess Peptide─Spermidine Ligase Activity.
Biochemistry, 62, 2023
|
|
7OL7
 
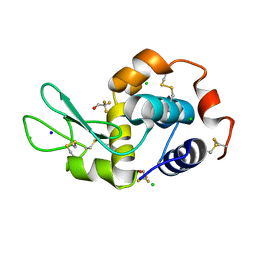 | |
8TX9
 
 | |
5BYT
 
 | |
5TZQ
 
 | | Crystal Structure of FPV039:Bmf BH3 complex | | Descriptor: | 1,2-ETHANEDIOL, Bcl-2-like protein FPV039, Bcl-2-modifying factor | | Authors: | Anasir, M.I, Kvansakul, M. | | Deposit date: | 2016-11-22 | | Release date: | 2017-05-03 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis of apoptosis inhibition by the fowlpox virus protein FPV039.
J. Biol. Chem., 292, 2017
|
|
8JT4
 
 | |
7OL8
 | |
6LTD
 
 | |
7UVA
 
 | | Crystal structure of KDM2A histone demethylase catalytic domain in complex with an H3C36 peptide modified by UNC8015 | | Descriptor: | FE (III) ION, Histone H3.2, Lysine-specific demethylase 2A, ... | | Authors: | Budziszewski, G.R, Azzam, D.N, Spangler, C.J, Skrajna, A, Foley, C.A, James, L.I, Frye, S.V, McGinty, R.K. | | Deposit date: | 2022-04-29 | | Release date: | 2023-02-22 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structural basis of paralog-specific KDM2A/B nucleosome recognition.
Nat.Chem.Biol., 19, 2023
|
|
7Y4H
 
 | |
5U6F
 
 | |
7OL6
 
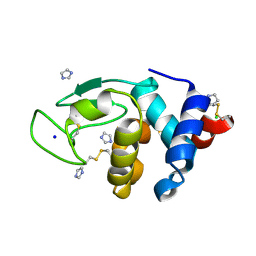 | |
7UI4
 
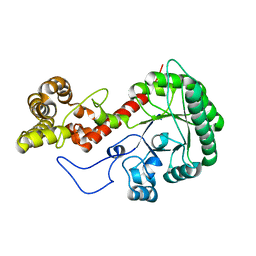 | |
8QDG
 
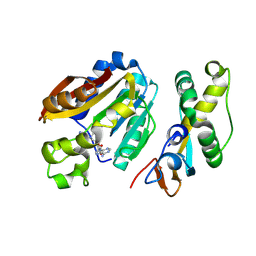 | | compound 1a bound KMT9 crystal structure | | Descriptor: | (2~{S})-4-[[(2~{R},3~{S},4~{R},5~{R})-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methyl-[(3~{S})-pyrrolidin-3-yl]amino]-2-azanyl-butanoic acid, Methyltransferase N6AMT1, Multifunctional methyltransferase subunit TRM112-like protein | | Authors: | Sheng, W, Eric, M, Roland, S. | | Deposit date: | 2023-08-29 | | Release date: | 2024-09-11 | | Method: | X-RAY DIFFRACTION (1.392 Å) | | Cite: | compound 1a bound KMT9 crystal structure
To Be Published
|
|
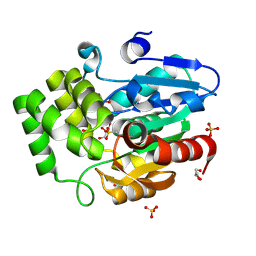
8U2L
 
 | |
5GHF
 
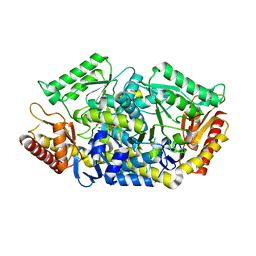 | | Transaminase with L-ala | | Descriptor: | 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, Aminotransferase class-III | | Authors: | Kim, J, Park, J. | | Deposit date: | 2016-06-19 | | Release date: | 2017-05-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Active Site Engineering of omega-Transaminase Guided by Docking Orientation Analysis and Virtual Activity Screening
Acs Catalysis, 7, 2017
|
|
8U33
 
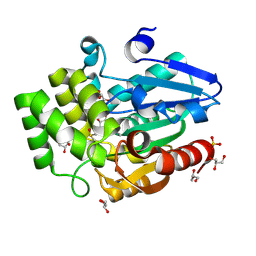 | |
7OMO
 
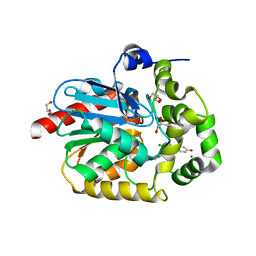 | |
7YD9
 
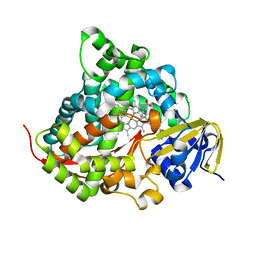 | | Crystal structure of the P450 BM3 heme domain mutant F87G/T268V/A184V/A328V in complex with N-imidazolyl-hexanoyl-L-phenylalanine,methylbenzene and hydroxylamine | | Descriptor: | (2S)-2-(6-imidazol-1-ylhexanoylamino)-3-phenyl-propanoic acid, Bifunctional cytochrome P450/NADPH--P450 reductase, HYDROXYAMINE, ... | | Authors: | Dong, S, Chen, J, Jiang, Y, Cong, Z, Feng, Y. | | Deposit date: | 2022-07-04 | | Release date: | 2023-05-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.748 Å) | | Cite: | Regiodivergent and Enantioselective Hydroxylation of C-H bonds by Synergistic Use of Protein Engineering and Exogenous Dual-Functional Small Molecules.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
