5FYQ
 
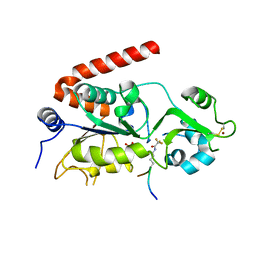 | | Sirt2 in complex with a 13-mer trifluoroacetylated Ran peptide | | Descriptor: | NAD-DEPENDENT PROTEIN DEACETYLASE SIRTUIN-2, RAN AA 31-43, SULFATE ION, ... | | Authors: | Knyphausen, P, de Boor, S, Scislowski, L, Extra, A, Baldus, L, Schacherl, M, Baumann, U, Neundorf, I, Lammers, M. | | Deposit date: | 2016-03-09 | | Release date: | 2016-05-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Insights Into Lysine-Deacetylation of Natively Folded Substrate Proteins by Sirtuins.
J.Biol.Chem., 291, 2016
|
|
8QCH
 
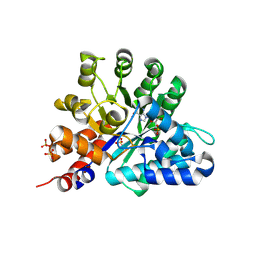 | | Human Adenosine deaminase-like protein in complex with compound AT8001 | | Descriptor: | Adenosine deaminase-like protein, DI(HYDROXYETHYL)ETHER, SODIUM ION, ... | | Authors: | Zimberger, C, canard, B, Ferron, F. | | Deposit date: | 2023-08-26 | | Release date: | 2024-07-31 | | Last modified: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (2.442 Å) | | Cite: | The activation cascade of the broad-spectrum antiviral bemnifosbuvir characterized at atomic resolution.
Plos Biol., 22, 2024
|
|
6MBY
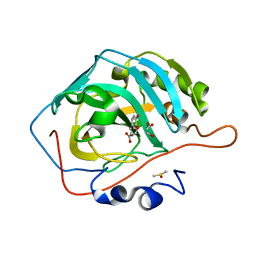 
 | | Carbonic Anhydrase II in complex with Ferulic Acid | | Descriptor: | 3-(4-HYDROXY-3-METHOXYPHENYL)-2-PROPENOIC ACID, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Lomelino, C.L, McKenna, R. | | Deposit date: | 2018-08-30 | | Release date: | 2019-03-13 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.496 Å) | | Cite: | Carbonic anhydrase II in complex with carboxylic acid-based inhibitors.
Acta Crystallogr F Struct Biol Commun, 75, 2019
|
|
8TOZ
 
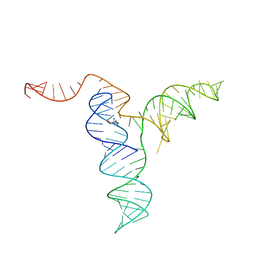 | |
7UI3
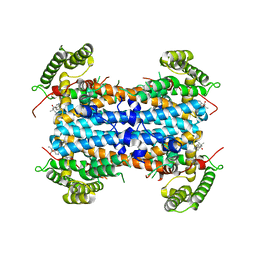 
 | |
7V1W
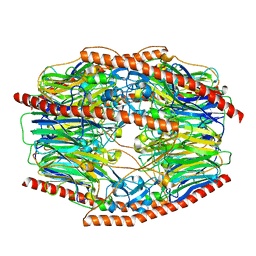 
 | | Difructose dianhydride I synthase/hydrolase (alphaFFase1) from Bifidobacterium dentium in complex with beta-D-arabinofuranose | | Descriptor: | CALCIUM ION, Difructose dianhydride I synthase/hydrolase (alphaFFase1), beta-D-arabinofuranose | | Authors: | Kashima, T, Arakawa, T, Yamada, C, Fujita, K, Fushinobu, S. | | Deposit date: | 2021-08-06 | | Release date: | 2021-11-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Identification of difructose dianhydride I synthase/hydrolase from an oral bacterium establishes a novel glycoside hydrolase family.
J.Biol.Chem., 297, 2021
|
|
5GGG
 
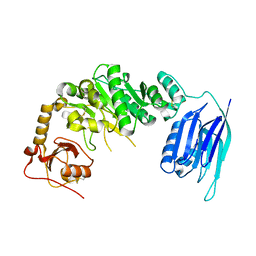 | |
7M3R
 
 | |
7P3G
 
 | | EED in complex with compound 4 | | Descriptor: | N-(2,3-dihydro-1-benzofuran-4-ylmethyl)-8-(4-methylsulfonylphenyl)-[1,2,4]triazolo[4,3-c]pyrimidin-5-amine, N5-[(5-fluoranyl-2,3-dihydro-1-benzofuran-4-yl)methyl]-N8-methyl-N8-(1-methylpyrazol-3-yl)-[1,2,4]triazolo[4,3-c]pyrimidine-5,8-diamine, Polycomb protein EED | | Authors: | Read, J.A. | | Deposit date: | 2021-07-07 | | Release date: | 2021-12-08 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Diverse, Potent, and Efficacious Inhibitors That Target the EED Subunit of the Polycomb Repressive Complex 2 Methyltransferase.
J.Med.Chem., 64, 2021
|
|
5GGN
 
 | | Crystal structure of N-terminal domain of human protein O-mannose beta-1,2-N-acetylglucosaminyltransferase in complex with GlcNAc-beta-pNP | | Descriptor: | 4-nitrophenyl 2-acetamido-2-deoxy-beta-D-glucopyranoside, Protein O-linked-mannose beta-1,2-N-acetylglucosaminyltransferase 1 | | Authors: | Kuwabara, N, Senda, T, Kato, R. | | Deposit date: | 2016-06-16 | | Release date: | 2016-08-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.211 Å) | | Cite: | Carbohydrate-binding domain of the POMGnT1 stem region modulates O-mannosylation sites of alpha-dystroglycan
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
8QIY
 
 | | Structure of Mycobacterium abscessus Phosphopantetheine adenylyltransferase in complex with inhibitor | | Descriptor: | 1-(2-aminophenyl)-5-(trifluoromethyl)pyrazole-4-carboxylic acid, Phosphopantetheine adenylyltransferase | | Authors: | Thomas, S.E, McCarthy, W.J, Coyne, A.G, Blundell, T.L. | | Deposit date: | 2023-09-12 | | Release date: | 2024-07-24 | | Last modified: | 2024-09-11 | | Method: | X-RAY DIFFRACTION (1.5149 Å) | | Cite: | A Fragment-Based Competitive 19 F LB-NMR Platform For Hotspot-Directed Ligand Profiling.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
7V1X
 
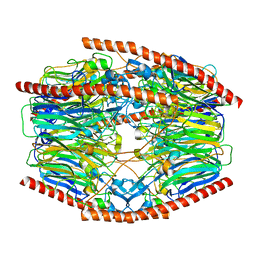 | | Difructose dianhydride I synthase/hydrolase (alphaFFase1) from Bifidobacterium dentium in complex with beta-D-fructofuranose | | Descriptor: | CALCIUM ION, Difructose dianhydride I synthase/hydrolase, beta-D-fructofuranose | | Authors: | Kashima, T, Arakawa, T, Yamada, C, Fujita, K, Fushinobu, S. | | Deposit date: | 2021-08-06 | | Release date: | 2021-11-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Identification of difructose dianhydride I synthase/hydrolase from an oral bacterium establishes a novel glycoside hydrolase family.
J.Biol.Chem., 297, 2021
|
|
5BXD
 
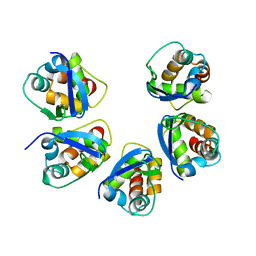 | |
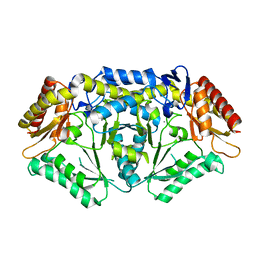
5TI8
 
 | |
8QLL
 
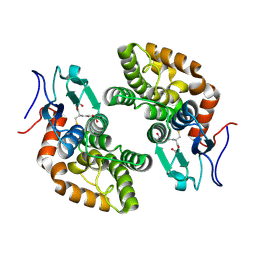 | |
5GHG
 
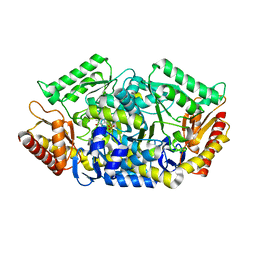 | | Transaminase W58L with SMBA | | Descriptor: | 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, Aminotransferase class-III | | Authors: | Kim, J, Park, J. | | Deposit date: | 2016-06-20 | | Release date: | 2017-05-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Active Site Engineering of omega-Transaminase Guided by Docking Orientation Analysis and Virtual Activity Screening
Acs Catalysis, 7, 2017
|
|
8TTA
 
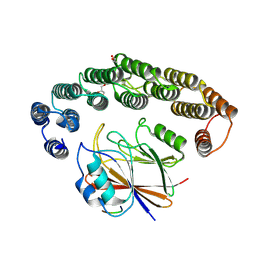 | | Structure of retromer VPS29-VPS35 (483-796) complexed with Fam21A repeat 21 (1328-1341) | | Descriptor: | ACETATE ION, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Chen, K.-E, Guo, Q, Collins, B.M. | | Deposit date: | 2023-08-13 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (3.46 Å) | | Cite: | Structural basis for coupling of the WASH subunit FAM21 with the endosomal SNX27-Retromer complex.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
7M3T
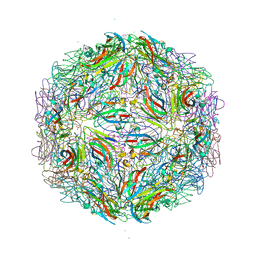 
 | |
6MA4
 
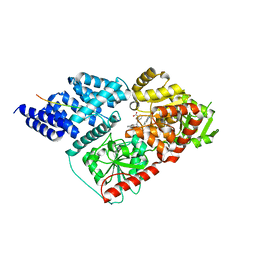 | | Crystal structure of human O-GlcNAc transferase bound to a peptide from HCF-1 pro-repeat 2 (11-26) and inhibitor 3a | | Descriptor: | 5-{2-[(1R)-2-{(carboxymethyl)[(thiophen-2-yl)methyl]amino}-2-oxo-1-{[(2-oxo-1,2-dihydroquinolin-6-yl)sulfonyl]amino}ethyl]phenoxy}pentanoic acid, Host Cell Factor 1 peptide, UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase 110 kDa subunit | | Authors: | Martin, S.E.S, Lazarus, M.B, Walker, S. | | Deposit date: | 2018-08-25 | | Release date: | 2018-10-17 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-Based Evolution of Low Nanomolar O-GlcNAc Transferase Inhibitors.
J. Am. Chem. Soc., 140, 2018
|
|
5TJ3
 
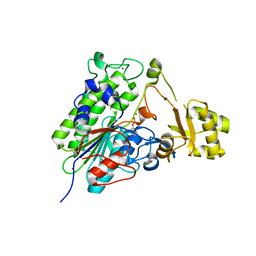 | | Crystal structure of wild type alkaline phosphatase PafA to 1.7A resolution | | Descriptor: | Alkaline phosphatase PafA, ZINC ION | | Authors: | Lyubimov, A.Y, Sunden, F, Ressl, S, Herschlag, D. | | Deposit date: | 2016-10-03 | | Release date: | 2016-11-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Mechanistic and Evolutionary Insights from Comparative Enzymology of Phosphomonoesterases and Phosphodiesterases across the Alkaline Phosphatase Superfamily.
J.Am.Chem.Soc., 138, 2016
|
|
5TJD
 
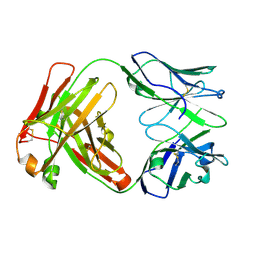 | | Computer-based rational design of improved functionality for antibody catalysts toward organophosphorus compounds | | Descriptor: | FAB A.17 L47K mutant HEAVY CHAIN, FAB A.17 L47K mutant Light Chain | | Authors: | Chatziefthimiou, S, Stepanova, A, Smirnov, I, Gabibov, A, Wilmanns, M. | | Deposit date: | 2016-10-04 | | Release date: | 2018-05-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Multiscale computation delivers organophosphorus reactivity and stereoselectivity to immunoglobulin scavengers.
Proc.Natl.Acad.Sci.USA, 2020
|
|
7M2V
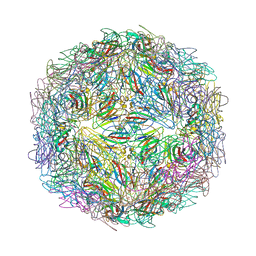 
 | |
7V1V
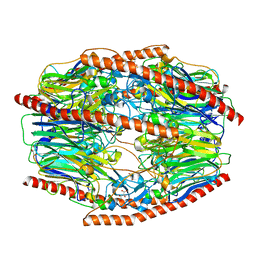 
 | | Difructose dianhydride I synthase/hydrolase (alphaFFase1) from Bifidobacterium dentium, ligand-free form | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCIUM ION, D(-)-TARTARIC ACID, ... | | Authors: | Kashima, T, Arakawa, T, Yamada, C, Fujita, K, Fushinobu, S. | | Deposit date: | 2021-08-06 | | Release date: | 2021-11-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Identification of difructose dianhydride I synthase/hydrolase from an oral bacterium establishes a novel glycoside hydrolase family.
J.Biol.Chem., 297, 2021
|
|
8TQF
 
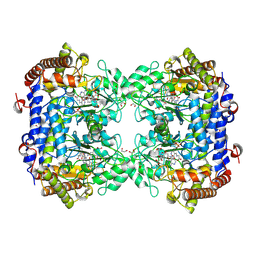 | | Crystal structure of Soybean SHMT8 in complex with PLP-glycine and diglutamylated 5-formyltetrahydrofolate | | Descriptor: | 1,2-ETHANEDIOL, N-GLYCINE-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YL-METHANE], N-[4-({[(6R)-2-amino-5-formyl-4-hydroxy-5,6,7,8-tetrahydropteridin-6-yl]methyl}amino)benzoyl]-L-gamma-glutamyl-L-glutamic acid, ... | | Authors: | Owuocha, L.F, Beamer, L.J. | | Deposit date: | 2023-08-07 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural insights into high affinity binding of polyglutamylated tetrahydrofolate to soybean serine hydroxymethyltransferase
To Be Published
|
|
8TTU
 
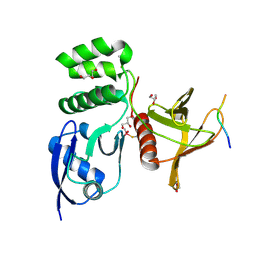 | | Structure of SNX27 FERM complexed with Fam21A repeat 19 (1261 - 1274) | | Descriptor: | DI(HYDROXYETHYL)ETHER, Fam21A repeat 19 peptide, GLYCEROL, ... | | Authors: | Guo, Q, Chen, K.-E, Collins, B.M. | | Deposit date: | 2023-08-15 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Structural basis for coupling of the WASH subunit FAM21 with the endosomal SNX27-Retromer complex.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
