4NFE
 
 | | Human kallikrein-related peptidase 2 in complex with benzamidine | | Descriptor: | BENZAMIDINE, Kallikrein-2, SULFATE ION | | Authors: | Skala, W, Brandstetter, H, Magdolen, V, Goettig, P. | | Deposit date: | 2013-10-31 | | Release date: | 2014-10-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-function analyses of human kallikrein-related peptidase 2 establish the 99-loop as master regulator of activity
J.Biol.Chem., 289, 2014
|
|
3BKY
 
 | | Crystal Structure of Chimeric Antibody C2H7 Fab in complex with a CD20 Peptide | | Descriptor: | B-lymphocyte antigen CD20, the Fab fragment of chimeric 2H7, heavy chain, ... | | Authors: | Du, J, Zhong, C, Ding, J. | | Deposit date: | 2007-12-07 | | Release date: | 2008-04-29 | | Last modified: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Crystal structure of chimeric antibody C2H7 Fab in complex with a CD20 peptide
Mol.Immunol., 45, 2008
|
|
3TL8
 
 | |
2Q8H
 
 | | Structure of pyruvate dehydrogenase kinase isoform 1 in complex with dichloroacetate (DCA) | | Descriptor: | DICHLORO-ACETIC ACID, POTASSIUM ION, [Pyruvate dehydrogenase [lipoamide]] kinase isozyme 1 | | Authors: | Kato, M, Li, J, Chuang, J.L, Chuang, D.T. | | Deposit date: | 2007-06-10 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Distinct Structural Mechanisms for Inhibition of Pyruvate Dehydrogenase Kinase Isoforms by AZD7545, Dichloroacetate, and Radicicol.
Structure, 15, 2007
|
|
5WSY
 
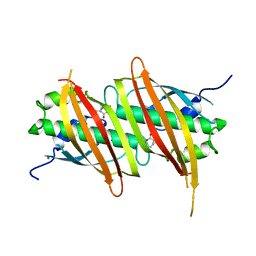 | | The complex structure of SAV606 with N-carboxymethyl-3-aminobutyrate | | Descriptor: | (3~{R})-3-(2-hydroxy-2-oxoethylamino)butanoic acid, Uncharacterized protein | | Authors: | Chisuga, T, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2016-12-08 | | Release date: | 2017-05-31 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural analysis of the dual-function thioesterase SAV606 unravels the mechanism of Michael addition of glycine to an alpha , beta-unsaturated thioester.
J. Biol. Chem., 292, 2017
|
|
3H16
 
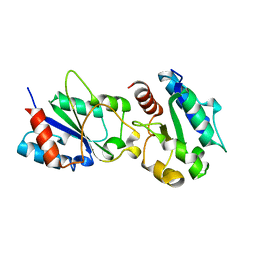 | | Crystal structure of a bacteria TIR domain, PdTIR from Paracoccus denitrificans | | Descriptor: | SULFATE ION, TIR protein | | Authors: | Chan, S.L, Low, L.Y, Santelli, E, Pascual, J. | | Deposit date: | 2009-04-11 | | Release date: | 2009-06-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Molecular Mimicry in Innate Immunity: CRYSTAL STRUCTURE OF A BACTERIAL TIR DOMAIN.
J.Biol.Chem., 284, 2009
|
|
2FSE
 
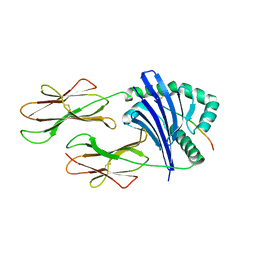 | | Crystallographic structure of a rheumatoid arthritis MHC susceptibility allele, HLA-DR1 (DRB1*0101), complexed with the immunodominant determinant of human type II collagen | | Descriptor: | Collagen alpha-1(II), H-2 class II histocompatibility antigen, E-K alpha chain, ... | | Authors: | Ivey, R.A, Rosloniec, E.F, Whittington, K.B, Kang, A.H, Park, H.W. | | Deposit date: | 2006-01-22 | | Release date: | 2006-09-19 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystallographic Structure of a Rheumatoid Arthritis MHC Susceptibility Allele, HLA-DR1 (DRB1*0101), Complexed with the Immunodominant Determinant of Human Type II Collagen.
J.Immunol., 177, 2006
|
|
6MVR
 
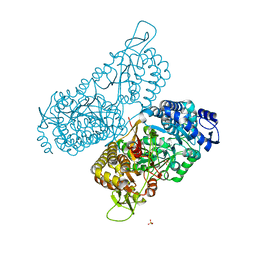 | | Structure of a bacterial ALDH16 | | Descriptor: | Aldehyde dehydrogenase, GLYCEROL, SULFATE ION | | Authors: | Tanner, J.J, Liu, L. | | Deposit date: | 2018-10-28 | | Release date: | 2018-12-26 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of Aldehyde Dehydrogenase 16 Reveals Trans-Hierarchical Structural Similarity and a New Dimer.
J. Mol. Biol., 431, 2019
|
|
2XOE
 
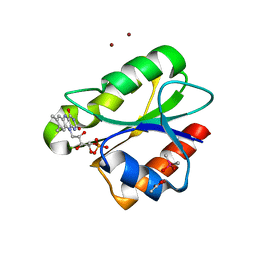 | | Crystal structure of flavoprotein NrdI from Bacillus anthracis in the semiquinone form | | Descriptor: | ACETATE ION, CACODYLATE ION, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Johansson, R, Sprenger, J, Torrents, E, Sahlin, M, Sjoberg, B.M, Logan, D.T. | | Deposit date: | 2010-08-14 | | Release date: | 2010-08-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | High Resolution Crystal Structures of Nrdi in the Oxidised and Reduced States: An Unusual Flavodoxin
FEBS J., 277, 2010
|
|
3BM2
 
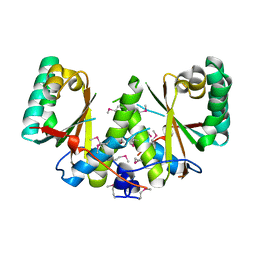 | |
3BW6
 | | Crystal structure of the longin domain of yeast Ykt6 | | Descriptor: | SULFATE ION, Synaptobrevin homolog YKT6 | | Authors: | Pylypenko, O, Schonichen, A, Ludwig, D, Ungermann, C, Goody, R.S, Rak, A, Geyer, M. | | Deposit date: | 2008-01-08 | | Release date: | 2008-04-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Farnesylation of the SNARE protein Ykt6 increases its stability and helical folding.
J.Mol.Biol., 377, 2008
|
|
8OLT
 
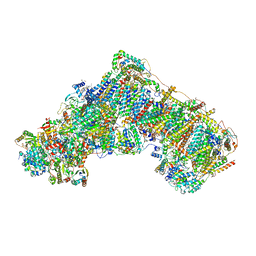 | | Mitochondrial complex I from Mus musculus in the active state bound with piericidin A | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, Acyl carrier protein, ... | | Authors: | Grba, D.N, Chung, I, Bridges, H.R, Agip, A.N.A, Hirst, J. | | Deposit date: | 2023-03-30 | | Release date: | 2023-08-09 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Investigation of hydrated channels and proton pathways in a high-resolution cryo-EM structure of mammalian complex I.
Sci Adv, 9, 2023
|
|
8OM1
 
 | | Mitochondrial complex I from Mus musculus in the active state | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Grba, D.N, Chung, I, Bridges, H.R, Agip, A.N.A, Hirst, J. | | Deposit date: | 2023-03-31 | | Release date: | 2023-08-09 | | Last modified: | 2023-08-16 | | Method: | ELECTRON MICROSCOPY (2.39 Å) | | Cite: | Investigation of hydrated channels and proton pathways in a high-resolution cryo-EM structure of mammalian complex I.
Sci Adv, 9, 2023
|
|
3BXH
 
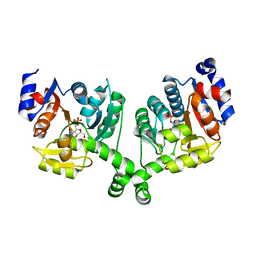 | |
2GN8
 
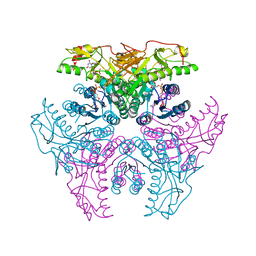 | | Crystal structure of UDP-GlcNAc inverting 4,6-dehydratase in complex with NADP and UDP | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, UDP-GlcNAc C6 dehydratase, ... | | Authors: | Ishiyama, N, Creuzenet, C, Lam, J.S, Berghuis, A.M. | | Deposit date: | 2006-04-09 | | Release date: | 2006-05-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Studies of FlaA1 from Helicobacter pylori Reveal the Mechanism for Inverting 4,6-Dehydratase Activity.
J.Biol.Chem., 281, 2006
|
|
4ZEW
 
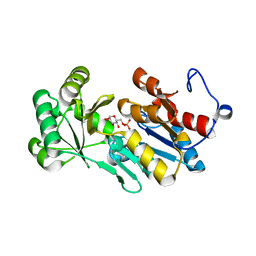 | | Crystal structure of PfHAD1 in complex with glucose-6-phosphate | | Descriptor: | 6-O-phosphono-alpha-D-glucopyranose, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Park, J, Tolia, N.H. | | Deposit date: | 2015-04-20 | | Release date: | 2015-09-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Cap-domain closure enables diverse substrate recognition by the C2-type haloacid dehalogenase-like sugar phosphatase Plasmodium falciparum HAD1.
Acta Crystallogr. D Biol. Crystallogr., 71, 2015
|
|
2O5N
 
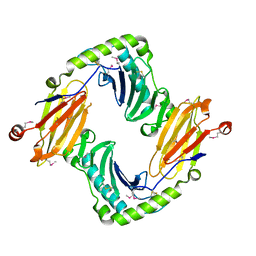 | | Crystal structure of a Viral Glycoprotein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, MuHV1gpm153, ... | | Authors: | Mans, J, Natarajan, K, Robinson, H, Margulies, D.H. | | Deposit date: | 2006-12-06 | | Release date: | 2007-09-25 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Cellular Expression and Crystal Structure of the Murine Cytomegalovirus Major Histocompatibility Complex Class I-like Glycoprotein, m153.
J.Biol.Chem., 282, 2007
|
|
3SMC
 
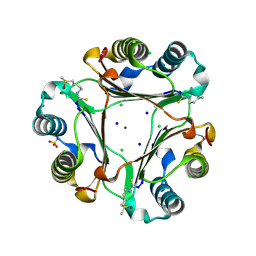 | |
2Q6V
 
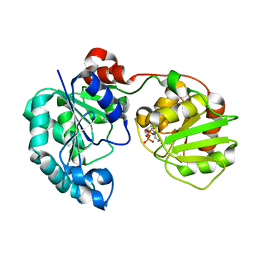 | | Crystal Structure of GumK in complex with UDP | | Descriptor: | Glucuronosyltransferase GumK, URIDINE-5'-DIPHOSPHATE | | Authors: | Barreras, M. | | Deposit date: | 2007-06-05 | | Release date: | 2008-06-10 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase.
J.Biol.Chem., 283, 2008
|
|
3FMZ
 | |
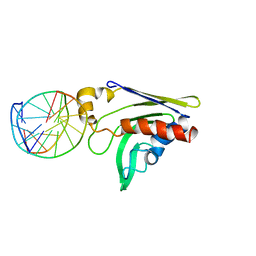
2GKD
 
 | |
4O2A
 
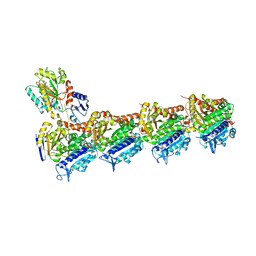 | | Tubulin-BAL27862 complex | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 3-[(4-{1-[2-(4-aminophenyl)-2-oxoethyl]-1H-benzimidazol-2-yl}-1,2,5-oxadiazol-3-yl)amino]propanenitrile, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Prota, A.E, Franck, D, Bachmann, F, Bargsten, K, Buey, R.M, Pohlmann, J, Reinelt, S, Lane, H, Steinmetz, M.O. | | Deposit date: | 2013-12-17 | | Release date: | 2014-03-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Novel Microtubule-Destabilizing Drug BAL27862 Binds to the Colchicine Site of Tubulin with Distinct Effects on Microtubule Organization.
J.Mol.Biol., 426, 2014
|
|
3F45
 
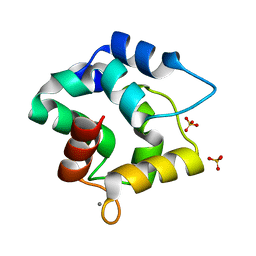 | | Structure of the R75A mutant of rat alpha-Parvalbumin | | Descriptor: | CALCIUM ION, Parvalbumin alpha, SULFATE ION | | Authors: | Hoh, F, Padilla, A. | | Deposit date: | 2008-10-31 | | Release date: | 2009-07-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Removing the invariant salt bridge of parvalbumin increases flexibility in the AB-loop structure
Acta Crystallogr.,Sect.D, 65, 2009
|
|
2HEZ
 
 | | Bifidobacterium longum bile salt hydrolase | | Descriptor: | Bile salt hydrolase, SULFATE ION | | Authors: | Suresh, C.G, Kumar, R.S, Brannigan, J.A. | | Deposit date: | 2006-06-22 | | Release date: | 2006-09-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and Functional Analysis of a Conjugated Bile Salt Hydrolase from Bifidobacterium longum Reveals an Evolutionary Relationship with Penicillin V Acylase.
J.Biol.Chem., 281, 2006
|
|
2Q8F
 
 | | Structure of pyruvate dehydrogenase kinase isoform 1 | | Descriptor: | POTASSIUM ION, [Pyruvate dehydrogenase [lipoamide]] kinase isozyme 1 | | Authors: | Kato, M, Li, J, Chuang, J.L, Chuang, D.T. | | Deposit date: | 2007-06-10 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Distinct Structural Mechanisms for Inhibition of Pyruvate Dehydrogenase Kinase Isoforms by AZD7545, Dichloroacetate, and Radicicol.
Structure, 15, 2007
|
|
