6DG8
 
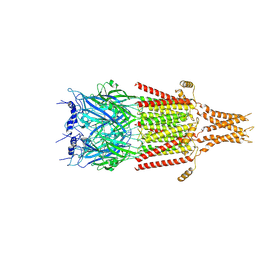 | | Full-length 5-HT3A receptor in a serotonin-bound conformation- State 2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-hydroxytryptamine receptor 3A, SEROTONIN, ... | | Authors: | Basak, S, Chakrapani, S. | | Deposit date: | 2018-05-17 | | Release date: | 2018-11-07 | | Last modified: | 2020-07-29 | | Method: | ELECTRON MICROSCOPY (3.89 Å) | | Cite: | Cryo-EM reveals two distinct serotonin-bound conformations of full-length 5-HT3Areceptor.
Nature, 563, 2018
|
|
6D3U
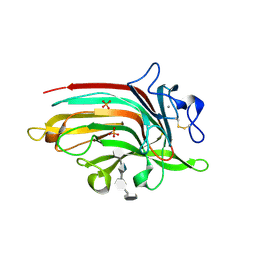 
 | | Complex structure of Ulvan lyase from Nonlaben Ulvanivorans- NLR48 | | Descriptor: | 1,2-ETHANEDIOL, 4-deoxy-alpha-L-threo-hex-4-enopyranuronic acid-(1-4)-3-O-sulfo-alpha-L-rhamnopyranose-(1-4)-beta-D-glucopyranuronic acid-(1-4)-3-O-sulfo-alpha-L-rhamnopyranose, CALCIUM ION, ... | | Authors: | Ulaganathan, T, Cygler, M. | | Deposit date: | 2018-04-16 | | Release date: | 2018-06-06 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Structural and functional characterization of PL28 family ulvan lyase NLR48 fromNonlabens ulvanivorans.
J. Biol. Chem., 293, 2018
|
|
3UJ8
 
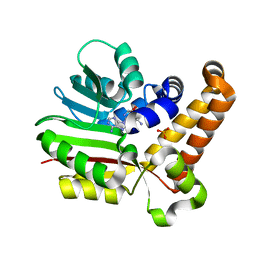 | | Phosphoethanolamine methyltransferase from Plasmodium falciparum in complex with sinefungin and PO4 | | Descriptor: | PHOSPHATE ION, Phosphoethanolamine N-methyltransferase, SINEFUNGIN | | Authors: | Lee, S.G, Kim, Y, Alpert, T.D, Nagata, A, Jez, J.M. | | Deposit date: | 2011-11-07 | | Release date: | 2011-11-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.351 Å) | | Cite: | Structure and reaction mechanism of phosphoethanolamine methyltransferase from the malaria parasite Plasmodium falciparum: an antiparasitic drug target.
J.Biol.Chem., 287, 2012
|
|
4OF3
 
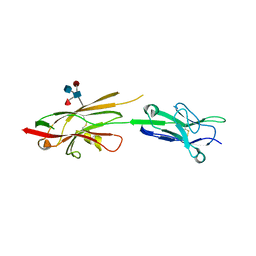 | | Crystal Structure of SYG-1 D1-D2, Glycosylated | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Protein SYG-1, isoform b, ... | | Authors: | Ozkan, E, Garcia, K.C. | | Deposit date: | 2014-01-14 | | Release date: | 2014-02-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Extracellular Architecture of the SYG-1/SYG-2 Adhesion Complex Instructs Synaptogenesis.
Cell(Cambridge,Mass.), 156, 2014
|
|
3UJC
 
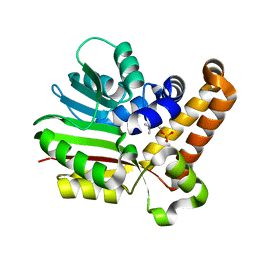 | | Phosphoethanolamine methyltransferase mutant (H132A) from Plasmodium falciparum in complex with phosphocholine | | Descriptor: | PHOSPHOCHOLINE, Phosphoethanolamine N-methyltransferase | | Authors: | Lee, S.G, Kim, Y, Alpert, T.D, Nagata, A, Jez, J.M. | | Deposit date: | 2011-11-07 | | Release date: | 2011-11-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.19 Å) | | Cite: | Structure and reaction mechanism of phosphoethanolamine methyltransferase from the malaria parasite Plasmodium falciparum: an antiparasitic drug target.
J.Biol.Chem., 287, 2012
|
|
6DG7
 
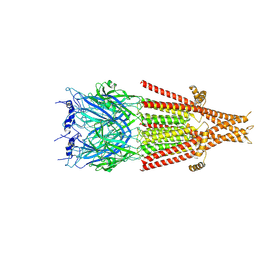 | | Full-length 5-HT3A receptor in a serotonin-bound conformation- State 1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-hydroxytryptamine receptor 3A, SEROTONIN, ... | | Authors: | Basak, S, Chakrapani, S. | | Deposit date: | 2018-05-17 | | Release date: | 2018-11-07 | | Last modified: | 2020-07-29 | | Method: | ELECTRON MICROSCOPY (3.32 Å) | | Cite: | Cryo-EM reveals two distinct serotonin-bound conformations of full-length 5-HT3Areceptor.
Nature, 563, 2018
|
|
8DWR
 
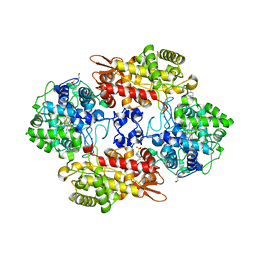 | | Crystal structure of the L333V variant of catalase-peroxidase from Mycobacterium tuberculosis | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Catalase-peroxidase, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Diaz-Vilchis, A, Uribe-Vazquez, B, Avila-Linares, A, Rudino-Pinera, E, Soberon, X. | | Deposit date: | 2022-08-01 | | Release date: | 2023-08-02 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Characterization of a catalase-peroxidase variant (L333V-KatG) identified in an INH-resistant Mycobacterium tuberculosis clinical isolate.
Biochem Biophys Rep, 37, 2024
|
|
6LAW
 
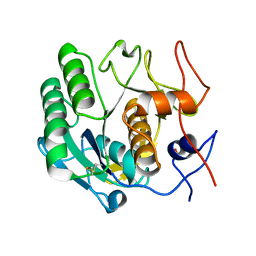 | |
4V9H
 
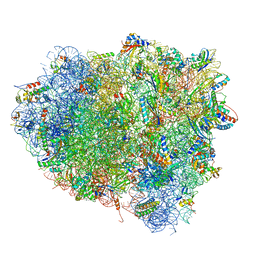 | | Crystal structure of the ribosome bound to elongation factor G in the guanosine triphosphatase state | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Tourigny, D.S, Fernandez, I.S, Kelley, A.C, Ramakrishnan, V. | | Deposit date: | 2013-03-25 | | Release date: | 2014-07-09 | | Last modified: | 2014-12-10 | | Method: | X-RAY DIFFRACTION (2.857 Å) | | Cite: | Elongation factor G bound to the ribosome in an intermediate state of translocation.
Science, 340, 2013
|
|
4V9R
 
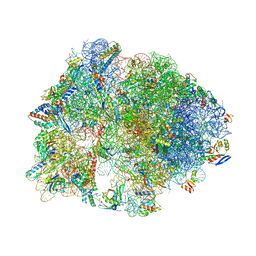 | | Crystal structure of antibiotic DITYROMYCIN bound to 70S ribosome | | Descriptor: | 16S Ribosomal RNA, 23S Ribosomal RNA, 30S Ribosomal Protein S10, ... | | Authors: | Bulkley, D.P, Brandi, L, Polikanov, Y.S, Fabbretti, A, O'Connor, M, Gualerzi, C.O, Steitz, T.A. | | Deposit date: | 2013-12-05 | | Release date: | 2014-07-09 | | Last modified: | 2014-12-10 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The antibiotics dityromycin and GE82832 bind protein S12 and block EF-G-catalyzed translocation.
Cell Rep, 6, 2014
|
|
7LWH
 
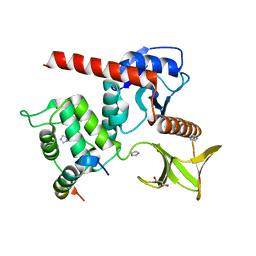 | | Human neurofibromin 2/merlin residues 1-339 in complex with LATS1 | | Descriptor: | GLYCEROL, IMIDAZOLE, Merlin, ... | | Authors: | Primi, M.C, Rangarajan, E.S, Izard, T. | | Deposit date: | 2021-03-01 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.606 Å) | | Cite: | Conformational flexibility determines the Nf2/merlin tumor suppressor functions.
Matrix Biol Plus, 12, 2021
|
|
7LMI
 
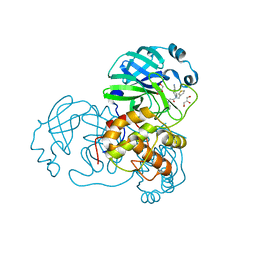 | | SARS-CoV-1 3CLPro in complex with N-(4-(1H-pyrazol-4-yl)phenyl)-2-(1H-benzo[d][1,2,3]triazol-1-yl)-N-(thiophen-3-ylmethyl)acetamide | | Descriptor: | 2-(benzotriazol-1-yl)-~{N}-[4-(1~{H}-pyrazol-4-yl)phenyl]-~{N}-(thiophen-3-ylmethyl)ethanamide, 3C-like proteinase, GLYCEROL | | Authors: | Arya, T, Goins, C.M, Macdonald, J.D, Stauffer, S.R. | | Deposit date: | 2021-02-05 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.707 Å) | | Cite: | Structure-Based Optimization of ML300-Derived, Noncovalent Inhibitors Targeting the Severe Acute Respiratory Syndrome Coronavirus 3CL Protease (SARS-CoV-2 3CL pro ).
J.Med.Chem., 65, 2022
|
|
7LDG
 
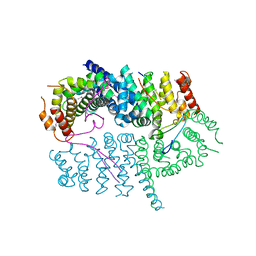 | | Crystal structure of the MEILB2-BRCA2 complex | | Descriptor: | Breast cancer type 2 susceptibility protein, Heat shock factor 2-binding protein | | Authors: | Nandakumar, J, Pendlebury, D.F. | | Deposit date: | 2021-01-13 | | Release date: | 2021-08-11 | | Last modified: | 2021-08-25 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Structure of a meiosis-specific complex central to BRCA2 localization at recombination sites.
Nat.Struct.Mol.Biol., 28, 2021
|
|
7LMD
 
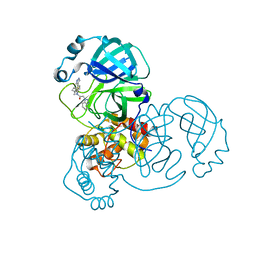 | | SARS-CoV-2 3CLPro in complex with 2-(benzotriazol-1-yl)-N-[4-(1H-pyrazol-4-yl)phenyl]-N-(3-thienylmethyl)acetamide | | Descriptor: | 2-(benzotriazol-1-yl)-~{N}-[4-(1~{H}-pyrazol-4-yl)phenyl]-~{N}-(thiophen-3-ylmethyl)ethanamide, 3C-like proteinase | | Authors: | Goins, C.M, Arya, T, Macdonald, J.D, Stauffer, S.R. | | Deposit date: | 2021-02-05 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structure-Based Optimization of ML300-Derived, Noncovalent Inhibitors Targeting the Severe Acute Respiratory Syndrome Coronavirus 3CL Protease (SARS-CoV-2 3CL pro ).
J.Med.Chem., 65, 2022
|
|
7LGM
 
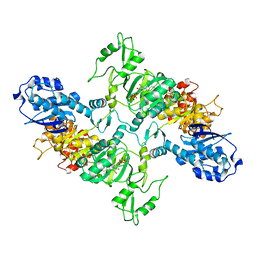 | | Cyanophycin synthetase from A. baylyi DSM587 with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Cyanophycin synthase | | Authors: | Sharon, I, Haque, A.S, Lahiri, I, Leschziner, A, Schmeing, T.M. | | Deposit date: | 2021-01-20 | | Release date: | 2021-08-18 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Structures and function of the amino acid polymerase cyanophycin synthetase.
Nat.Chem.Biol., 17, 2021
|
|
7LRG
 
 | |
7LG5
 
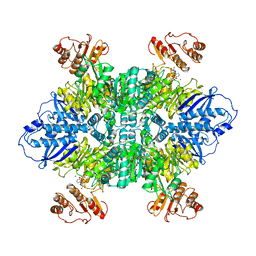 | |
7LME
 
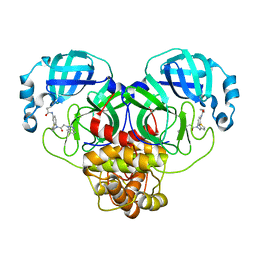 | | SARS-CoV-2 3CLPro in complex with N-[4-[[2-(benzotriazol-1-yl)acetyl]-(3-thienylmethyl)amino]phenyl]cyclopropanecarboxamide | | Descriptor: | 3C-like proteinase, ~{N}-[4-[2-(benzotriazol-1-yl)ethanoyl-(thiophen-3-ylmethyl)amino]phenyl]cyclopropanecarboxamide | | Authors: | Goins, C.M, Arya, T, Macdonald, J.D, Stauffer, S.R. | | Deposit date: | 2021-02-05 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-Based Optimization of ML300-Derived, Noncovalent Inhibitors Targeting the Severe Acute Respiratory Syndrome Coronavirus 3CL Protease (SARS-CoV-2 3CL pro ).
J.Med.Chem., 65, 2022
|
|
7LHP
 
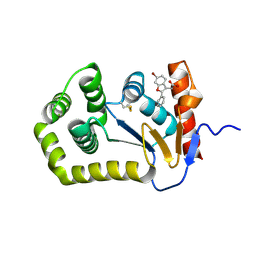 | |
7LHV
 
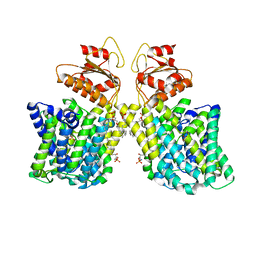 | | Structure of Arabidopsis thaliana sulfate transporter AtSULTR4;1 | | Descriptor: | (2S,3R,4E)-2-amino-3-hydroxyoctadec-4-en-1-yl dihydrogen phosphate, 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, SULFATE ION, ... | | Authors: | Wang, L, Chen, K, Zhou, M. | | Deposit date: | 2021-01-26 | | Release date: | 2021-08-11 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.75 Å) | | Cite: | Structure and function of an Arabidopsis thaliana sulfate transporter.
Nat Commun, 12, 2021
|
|
7LKQ
 
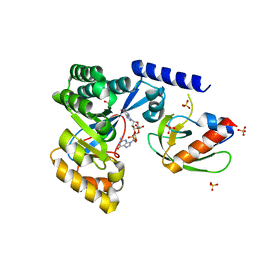 | | The PilZ(delta107-117)-FimX(GGDEF-EAL) complex from Xanthomonas citri | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), FimX(GGDEF-EAL), MAGNESIUM ION, ... | | Authors: | Llontop, E.E, Guzzo, C.R, Farah, C.S. | | Deposit date: | 2021-02-02 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | The PilB-PilZ-FimX regulatory complex of the Type IV pilus from Xanthomonas citri.
Plos Pathog., 17, 2021
|
|
7LKO
 
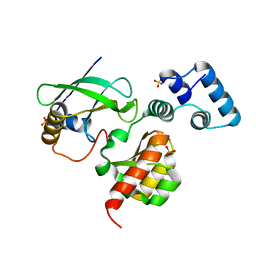 | |
7LMH
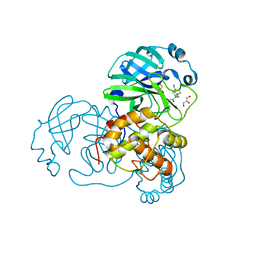 
 | | SARS-CoV-1 3CLPro in complex with 2-(1H-benzo[d][1,2,3]triazol-1-yl)-N-(4-(pyridin-3-yl)phenyl)-N-(thiophen-3-ylmethyl)acetamide | | Descriptor: | 2-(benzotriazol-1-yl)-~{N}-(4-pyridin-3-ylphenyl)-~{N}-(thiophen-3-ylmethyl)ethanamide, 3C-like proteinase, GLYCEROL | | Authors: | Arya, T, Goins, C.M, Macdonald, J.D, Stauffer, S.R. | | Deposit date: | 2021-02-05 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure-Based Optimization of ML300-Derived, Noncovalent Inhibitors Targeting the Severe Acute Respiratory Syndrome Coronavirus 3CL Protease (SARS-CoV-2 3CL pro ).
J.Med.Chem., 65, 2022
|
|
7LMJ
 
 | | SARS-CoV-1 3CLPro in complex with 2-(1H-benzo[d][1,2,3]triazol-1-yl)-N-(3-chlorobenzyl)-N-(4-(2-oxo-1,2-dihydropyridin-3-yl)phenyl)acetamide | | Descriptor: | 2-(benzotriazol-1-yl)-~{N}-[(3-chlorophenyl)methyl]-~{N}-[4-(2-oxidanylidene-1~{H}-pyridin-3-yl)phenyl]ethanamide, 3C-like proteinase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Arya, T, Goins, C.M, Macdonald, J.D, Stauffer, S.R. | | Deposit date: | 2021-02-05 | | Release date: | 2021-08-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.686 Å) | | Cite: | Structure-Based Optimization of ML300-Derived, Noncovalent Inhibitors Targeting the Severe Acute Respiratory Syndrome Coronavirus 3CL Protease (SARS-CoV-2 3CL pro ).
J.Med.Chem., 65, 2022
|
|
7LKN
 
 | |
