6AT0
 
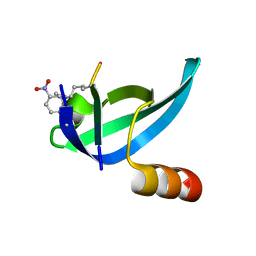 | |
4QLC
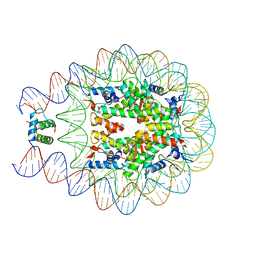 
 | | Crystal structure of chromatosome at 3.5 angstrom resolution | | Descriptor: | CITRIC ACID, DNA (167-mer), H5, ... | | Authors: | Jiang, J.S, Zhou, B.R, Xiao, T.S, Bai, Y.W. | | Deposit date: | 2014-06-11 | | Release date: | 2015-07-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.503 Å) | | Cite: | Structural Mechanisms of Nucleosome Recognition by Linker Histones.
Mol.Cell, 33 Suppl 1, 2015
|
|
6B7J
 
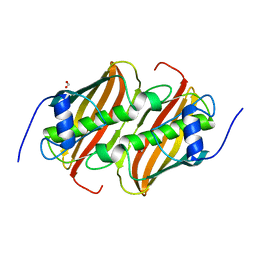 | |
4QMH
 
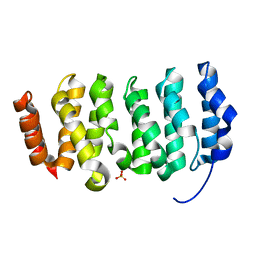 | | The XMAP215 family drives microtubule polymerization using a structurally diverse TOG array | | Descriptor: | LP04448p, SULFATE ION | | Authors: | Fox, J.C, Howard, A.E, Currie, J.D, Rogers, S.L, Slep, K.C. | | Deposit date: | 2014-06-16 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.652 Å) | | Cite: | The XMAP215 family drives microtubule polymerization using a structurally diverse TOG array.
Mol.Biol.Cell, 25, 2014
|
|
8G83
 
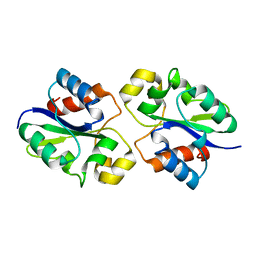 | | Structure of NAD+ consuming protein Acinetobacter baumannii TIR domain | | Descriptor: | NAD(+) hydrolase AbTIR | | Authors: | Klontz, E.H, Wang, Y, Glendening, G, Carr, J, Tsibouris, T, Buddula, S, Nallar, S, Soares, A, Snyder, G.A. | | Deposit date: | 2023-02-17 | | Release date: | 2023-10-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.03 Å) | | Cite: | The structure of NAD + consuming protein Acinetobacter baumannii TIR domain shows unique kinetics and conformations.
J.Biol.Chem., 299, 2023
|
|
4R9P
 
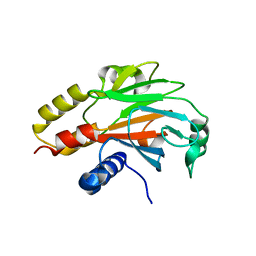 | | An Expansion to the Smad MH2-family: The structure of the N-MH2 expanded domain | | Descriptor: | RE28239p | | Authors: | Beich-Frandsen, M, Aragon, E, Llimargas, M, Benach, J, Riera, A, Macias, M.J. | | Deposit date: | 2014-09-06 | | Release date: | 2015-04-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.592 Å) | | Cite: | Structure of the N-terminal domain of the protein Expansion: an 'Expansion' to the Smad MH2
Acta Crystallogr.,Sect.D, 71, 2015
|
|
8G4P
 
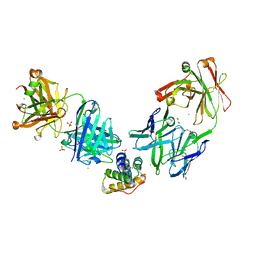 | | Crystal structure of the peanut allergen Ara h 2 bound by two neutralizing antibodies 13T1 and 13T5 | | Descriptor: | 1,2-ETHANEDIOL, 13T1 Fab light chain, 13T5 Fab heavy chain, ... | | Authors: | Pedersen, L.C, Mueller, G.A, Min, J. | | Deposit date: | 2023-02-10 | | Release date: | 2023-12-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Design of an Ara h 2 hypoallergen from conformational epitopes.
Clin Exp Allergy, 54, 2024
|
|
6BN1
 
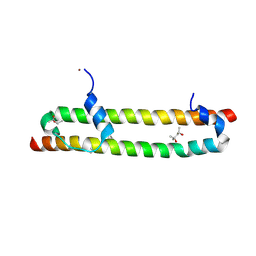 | | Salvador Hippo SARAH domain complex | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, NICKEL (II) ION, Scaffold protein salvador, ... | | Authors: | Cairns, L, Kavran, J.M. | | Deposit date: | 2017-11-15 | | Release date: | 2018-03-14 | | Last modified: | 2018-04-25 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Salvador has an extended SARAH domain that mediates binding to Hippo kinase.
J. Biol. Chem., 293, 2018
|
|
4QUC
 
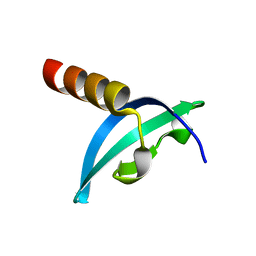 | | Crystal structure of chromodomain of Rhino | | Descriptor: | RE36324p | | Authors: | Li, S, Patel, D.J. | | Deposit date: | 2014-07-10 | | Release date: | 2014-08-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.502 Å) | | Cite: | Transgenerationally inherited piRNAs trigger piRNA biogenesis by changing the chromatin of piRNA clusters and inducing precursor processing.
Genes Dev., 28, 2014
|
|
4QUF
 
 | | crystal structure of chromodomain of Rhino with H3K9me3 | | Descriptor: | H3(1-15)K9me3 peptide, RE36324p | | Authors: | Li, S, Patel, D.J. | | Deposit date: | 2014-07-10 | | Release date: | 2014-08-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.501 Å) | | Cite: | Transgenerationally inherited piRNAs trigger piRNA biogenesis by changing the chromatin of piRNA clusters and inducing precursor processing.
Genes Dev., 28, 2014
|
|
6BQM
 
 | | Secreted serine protease VesC from Vibrio cholerae | | Descriptor: | serine protease VesC | | Authors: | Park, Y.J, Korotkov, K.V, Delarosa, J.R, Turley, S, DiMaio, F, Hol, W.G.J. | | Deposit date: | 2017-11-28 | | Release date: | 2018-11-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Suppressor Mutations in Type II Secretion Mutants of Vibrio cholerae: Inactivation of the VesC Protease.
Msphere, 5, 2020
|
|
8H7J
 
 | | The crystal structure of CD163 SRCR5-9 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Scavenger receptor cysteine-rich type 1 protein M130, ... | | Authors: | Luo, Z.P, Li, R, Ma, H.F. | | Deposit date: | 2022-10-20 | | Release date: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of CD163 SRCR5-9
To Be Published
|
|
6TUU
 
 | | Leishmania infantum Rad51 surrogate LiRadA10 in complex with 5,6,7,8-tetrahydro-2-naphthoic acid | | Descriptor: | 5,6,7,8-tetrahydronaphthalene-2-carboxylic acid, CHLORIDE ION, DNA repair and recombination protein RadA, ... | | Authors: | Pantelejevs, T, Hyvonen, M. | | Deposit date: | 2020-01-08 | | Release date: | 2021-01-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Development of dedicated crystallographic systems for structure-guided drug discovery
To Be Published
|
|
4NWO
 
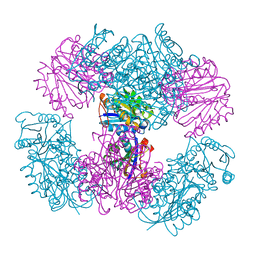 | | Computationally Designed Two-Component Self-Assembling Tetrahedral Cage T33-15 | | Descriptor: | CALCIUM ION, Chorismate mutase AroH, Molybdenum cofactor biosynthesis protein MogA | | Authors: | McNamara, D.E, King, N.P, Bale, J.B, Sheffler, W, Baker, D, Yeates, T.O. | | Deposit date: | 2013-12-06 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Accurate design of co-assembling multi-component protein nanomaterials.
Nature, 510, 2014
|
|
4TS6
 | |
4RKH
 
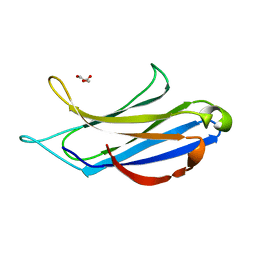
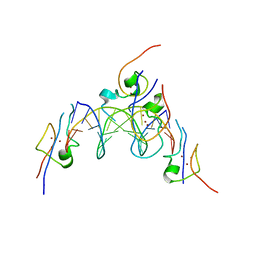 | | Structure of the MSL2 CXC domain bound with a specific MRE sequence | | Descriptor: | DNA (5'-D(*AP*TP*CP*CP*AP*TP*CP*TP*CP*GP*CP*TP*CP*AP*T)-3'), DNA (5'-D(*AP*TP*GP*AP*GP*CP*GP*AP*GP*AP*TP*GP*GP*AP*T)-3'), E3 ubiquitin-protein ligase msl-2, ... | | Authors: | Zheng, S, Ye, K. | | Deposit date: | 2014-10-13 | | Release date: | 2015-01-21 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis of X chromosome DNA recognition by the MSL2 CXC domain during Drosophila dosage compensation.
Genes Dev., 28, 2014
|
|
4OF8
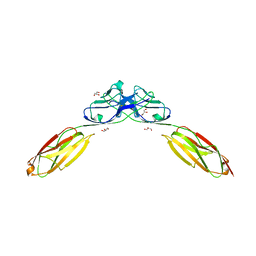 
 | | Crystal Structure of Rst D1-D2 | | Descriptor: | GLYCEROL, Irregular chiasm C-roughest protein, SODIUM ION | | Authors: | Ozkan, E, Garcia, K.C. | | Deposit date: | 2014-01-14 | | Release date: | 2014-02-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Extracellular Architecture of the SYG-1/SYG-2 Adhesion Complex Instructs Synaptogenesis.
Cell(Cambridge,Mass.), 156, 2014
|
|
8HLS
 
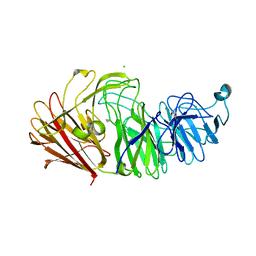 | | Crystal structure of the sialidase Am1547 from A.muciniphila | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, Sialidase domain-containing protein | | Authors: | Rui, B, Xinyue, T. | | Deposit date: | 2022-11-30 | | Release date: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Structural analysis of a novel sialidase from Akkermansia muciniphila reveals a distinct catalytic motif in GH33 family
To Be Published
|
|
4RWT
 
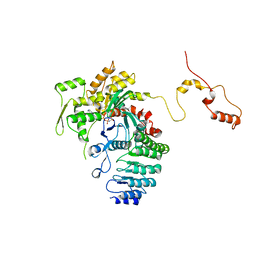 | | Structure of actin-Lmod complex | | Descriptor: | Actin-5C, Leiomodin-2, MAGNESIUM ION, ... | | Authors: | Chen, X, Ni, F, Wang, Q. | | Deposit date: | 2014-12-05 | | Release date: | 2015-10-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | Mechanisms of leiomodin 2-mediated regulation of actin filament in muscle cells.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4MAP
 
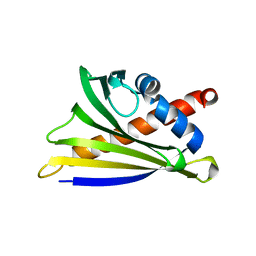 | | Crystal structure of Ara h 8 purified with heating | | Descriptor: | Ara h 8 allergen, SODIUM ION | | Authors: | Offermann, L.R, Hurlburt, B.K, Majorek, K.A, McBride, J.K, Maleki, S.J, Chruszcz, M. | | Deposit date: | 2013-08-16 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure and Function of the Peanut Panallergen Ara h 8.
J.Biol.Chem., 288, 2013
|
|
4M9B
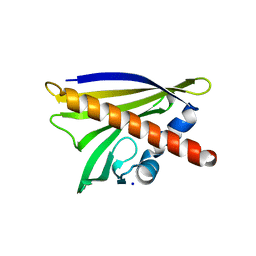 
 | | Crystal structure of Apo Ara h 8 | | Descriptor: | Ara h 8 allergen, SODIUM ION | | Authors: | Offermann, L.R, Hurlburt, B.K, Majorek, K.A, McBride, J.K, Maleki, S.J, Chruszcz, M. | | Deposit date: | 2013-08-14 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure and Function of the Peanut Panallergen Ara h 8.
J.Biol.Chem., 288, 2013
|
|
4RKG
 
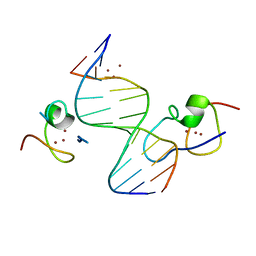 | | Structure of the MSL2 CXC domain bound with a non-specific (GC)6 DNA | | Descriptor: | DNA (5'-D(*GP*CP*GP*CP*GP*CP*GP*CP*GP*CP*GP*C)-3'), E3 ubiquitin-protein ligase msl-2, ZINC ION | | Authors: | Zheng, S, Ye, K. | | Deposit date: | 2014-10-13 | | Release date: | 2015-01-21 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis of X chromosome DNA recognition by the MSL2 CXC domain during Drosophila dosage compensation.
Genes Dev., 28, 2014
|
|
4RV0
 
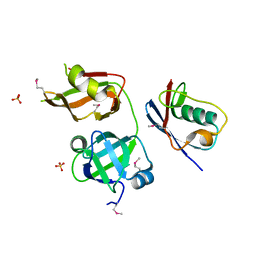 | | Crystal structure of TN complex | | Descriptor: | Nuclear protein localization protein 4 homolog, SULFATE ION, Transitional endoplasmic reticulum ATPase TER94 | | Authors: | Hao, Q, Jiao, S, Shi, Z.B, Zhou, Z.C. | | Deposit date: | 2014-11-23 | | Release date: | 2015-11-25 | | Method: | X-RAY DIFFRACTION (1.998 Å) | | Cite: | Crystal structure of TN complex
To be Published
|
|
4U77
 
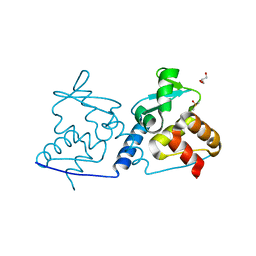 | | BTB domain from Drosophila CP190 | | Descriptor: | Centrosome-associated zinc finger protein CP190, DIMETHYL SULFOXIDE, GLYCEROL | | Authors: | Allemand, F, Cohen-Gonsaud, M, Labesse, G, Nollmann, M. | | Deposit date: | 2014-07-30 | | Release date: | 2014-08-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Chromatin Insulator Factors Involved in Long-Range DNA Interactions and Their Role in the Folding of the Drosophila Genome.
Plos Genet., 10, 2014
|
|
4LW3
 
 | |
