3FKW
 
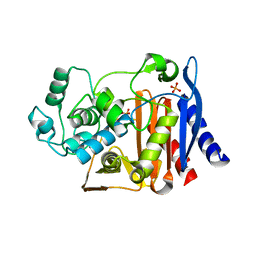 | | AmpC K67R mutant apo structure | | Descriptor: | Beta-lactamase, PHOSPHATE ION, POTASSIUM ION | | Authors: | Chen, Y, McReynolds, A, Shoichet, B.K. | | Deposit date: | 2008-12-17 | | Release date: | 2009-03-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.498 Å) | | Cite: | Re-examining the role of Lys67 in class C beta-lactamase catalysis.
Protein Sci., 18, 2009
|
|
3FKV
 
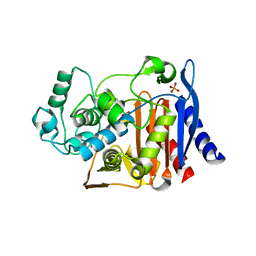 | | AmpC K67R mutant complexed with benzo(b)thiophene-2-boronic acid (bzb) | | Descriptor: | BENZO[B]THIOPHENE-2-BORONIC ACID, Beta-lactamase, PHOSPHATE ION, ... | | Authors: | Chen, Y, McReynolds, A, Shoichet, B.K. | | Deposit date: | 2008-12-17 | | Release date: | 2009-03-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.847 Å) | | Cite: | Re-examining the role of Lys67 in class C beta-lactamase catalysis.
Protein Sci., 18, 2009
|
|
3LJZ
 
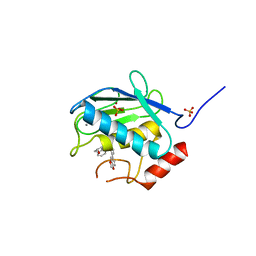 | | Crystal Structure of Human MMP-13 complexed with an Amino-2-indanol compound | | Descriptor: | (2R)-2-[4-(1,3-benzodioxol-5-yl)benzyl]-N~4~-hydroxy-N~1~-[(1S,2R)-2-hydroxy-2,3-dihydro-1H-inden-1-yl]butanediamide, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CALCIUM ION, ... | | Authors: | Shieh, H.-S, Kiefer, J.R. | | Deposit date: | 2010-01-26 | | Release date: | 2011-02-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure analysis reveals the flexibility of the ADAMTS-5 active site.
Protein Sci., 20, 2011
|
|
3LJT
 
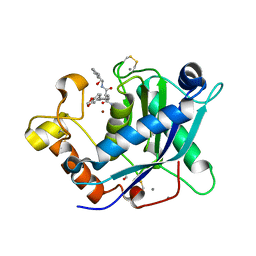 | | Crystal Structure of the Catalytic Domain of ADAMTS-5 in Complex with an Amino-2-indanol compound | | Descriptor: | (2R)-2-[4-(1,3-benzodioxol-5-yl)benzyl]-N~4~-hydroxy-N~1~-[(1S,2R)-2-hydroxy-2,3-dihydro-1H-inden-1-yl]butanediamide, 1,2-ETHANEDIOL, A disintegrin and metalloproteinase with thrombospondin motifs 5, ... | | Authors: | Shieh, H.-S, Williams, J.M, Caspers, N. | | Deposit date: | 2010-01-26 | | Release date: | 2010-03-31 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure analysis reveals the flexibility of the ADAMTS-5 active site.
Protein Sci., 20, 2011
|
|
3ONR
 | | Crystal structure of the calcium chelating immunodominant antigen, calcium dodecin (Rv0379),from Mycobacterium tuberculosis with a novel calcium-binding site | | Descriptor: | CALCIUM ION, FORMIC ACID, PLATINUM (II) ION, ... | | Authors: | Arulandu, A, Sacchettini, J.C. | | Deposit date: | 2010-08-30 | | Release date: | 2011-03-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of calcium dodecin (Rv0379), from Mycobacterium tuberculosis with a unique calcium-binding site.
Protein Sci., 20, 2011
|
|
3QPS
 
 | | Crystal structures of CmeR-bile acid complexes from Campylobacter jejuni | | Descriptor: | CHOLIC ACID, CmeR | | Authors: | Lei, H.T, Routh, M.D, Shen, Z, Su, C.C, Zhang, Q, Yu, E.W. | | Deposit date: | 2011-02-14 | | Release date: | 2011-03-09 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.351 Å) | | Cite: | Crystal structures of CmeR-bile acid complexes from Campylobacter jejuni.
Protein Sci., 20, 2011
|
|
3QQA
 
 | | Crystal structures of CmeR-bile acid complexes from Campylobacter jejuni | | Descriptor: | CmeR, TAUROCHOLIC ACID | | Authors: | Lei, H.T, Routh, M.D, Shen, Z, Su, C.-C, Zhang, Q, Yu, E.W. | | Deposit date: | 2011-02-15 | | Release date: | 2011-03-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of CmeR-bile acid complexes from Campylobacter jejuni.
Protein Sci., 20, 2011
|
|
