2QTP
 
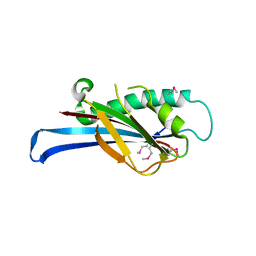 | |
4HF7
 
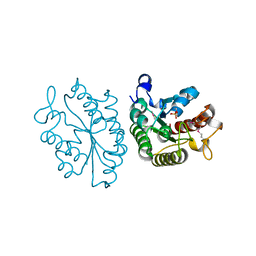 | |
2QZC
 
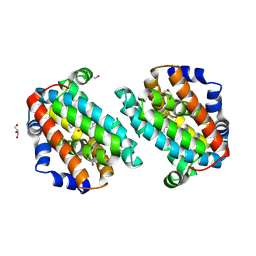 | |
4GOT
 
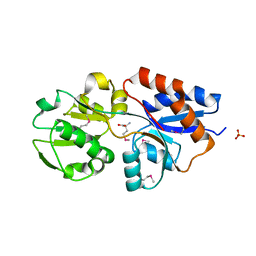 | |
2R4I
 
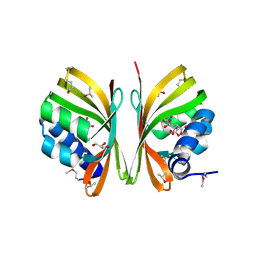 | |
2QHP
 
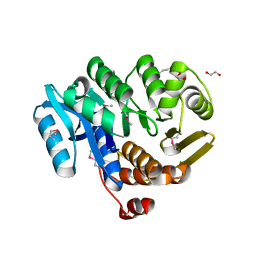 | |
2QJV
 
 | |
4GOQ
 
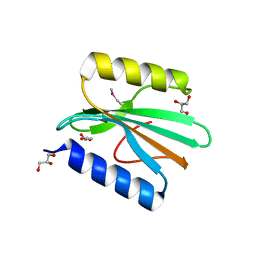 | |
4JPQ
 
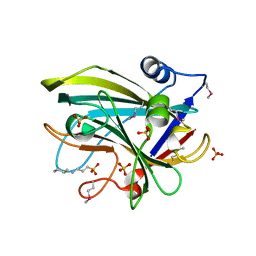 | |
4JM1
 
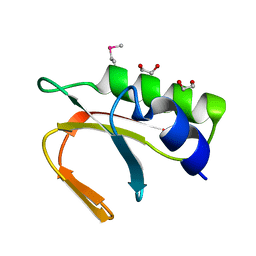 | |
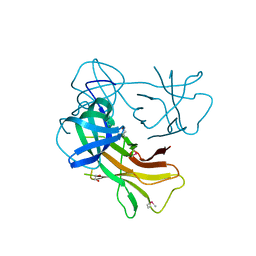
4J8Q
 
 | |
4JX0
 
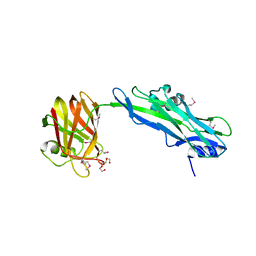 | |
3V4K
 
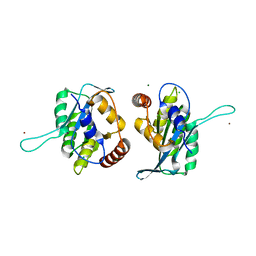 | | First-In-Class Small Molecule Inhibitors of the Single-strand DNA Cytosine Deaminase APOBEC3G | | Descriptor: | CHLORIDE ION, DNA dC->dU-editing enzyme APOBEC-3G, MAGNESIUM ION, ... | | Authors: | Shandilya, S.M.D, Ali, A, Schiffer, C.A. | | Deposit date: | 2011-12-15 | | Release date: | 2012-01-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | First-In-Class Small Molecule Inhibitors of the Single-Strand DNA Cytosine Deaminase APOBEC3G.
Acs Chem.Biol., 7, 2012
|
|
3V57
 
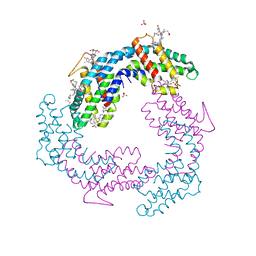 | |
4O83
 
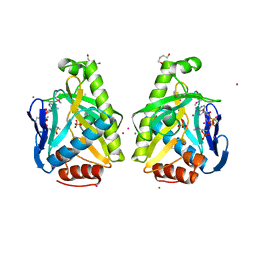 | |
3UPC
 
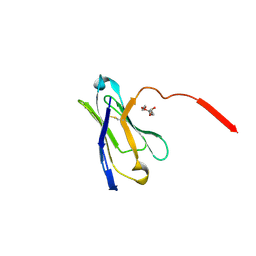 | | A general strategy for the generation of human antibody variable domains with increased aggregation resistance | | Descriptor: | DI(HYDROXYETHYL)ETHER, PENTAETHYLENE GLYCOL, TETRAETHYLENE GLYCOL, ... | | Authors: | Dudgeon, K, Rouet, R, Kokmeijer, I, Langley, D.B, Christ, D. | | Deposit date: | 2011-11-17 | | Release date: | 2012-06-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | General strategy for the generation of human antibody variable domains with increased aggregation resistance
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3UVH
 
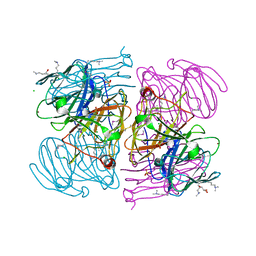
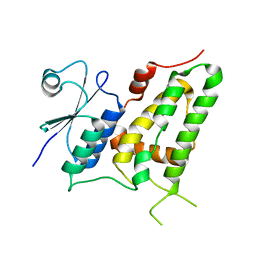 | | Crystal Structure Analysis of E81M mutant of human CLIC1 | | Descriptor: | Chloride intracellular channel protein 1 | | Authors: | Fanucchi, S, Achilonu, I.A, Fernandes, M.A, Legg-E'Silva, D.A, Stoychev, S.H, Dirr, H.W. | | Deposit date: | 2011-11-30 | | Release date: | 2011-12-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal Structure Analysis of E81M mutant of human CLIC1
To be Published
|
|
4O7L
 
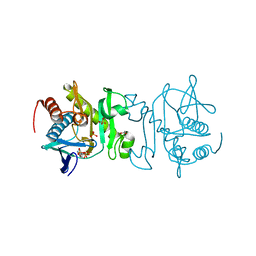 | |
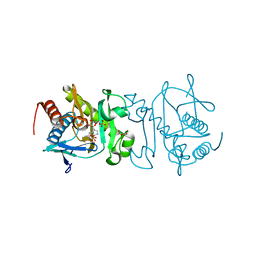
4O7Z
 
 | |
8DWM
 
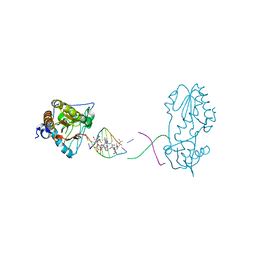 | | Host-guest complex of bleomycin A2 fully bound to CTTAGTTATAACTAAG | | Descriptor: | BLEOMYCIN A2, COBALT (III) ION, DNA (5'-D(*CP*TP*TP*AP*GP*TP*TP*A)-3'), ... | | Authors: | Georgiadis, M.M. | | Deposit date: | 2022-08-01 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Two distinct rotations of bithiazole DNA intercalation revealed by direct comparison of crystal structures of Co(III)•bleomycin A 2 and B 2 bound to duplex 5'-TAGTT sites.
Bioorg.Med.Chem., 77, 2023
|
|
4O7Y
 
 | |
4OKA
 
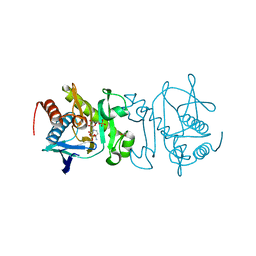
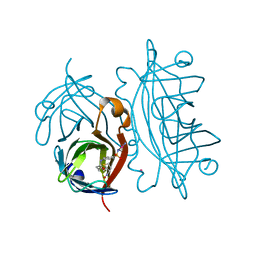 | | Structural-, Kinetic- and Docking Studies of Artificial Imine Reductases Based on the Biotin-Streptavidin Technology: An Induced Lock-and-Key Hypothesis | | Descriptor: | IRIDIUM ION, Streptavidin, [N-(4-{[2-(amino-kappaN)ethyl]sulfamoyl-kappaN}phenyl)-5-(2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl)pentanamidato]iridium(III) | | Authors: | Schirmer, T, Heinisch, T. | | Deposit date: | 2014-01-22 | | Release date: | 2014-11-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.505 Å) | | Cite: | Structural, Kinetic, and Docking Studies of Artificial Imine Reductases Based on Biotin-Streptavidin Technology: An Induced Lock-and-Key Hypothesis
J.Am.Chem.Soc., 136, 2014
|
|
4OJI
 
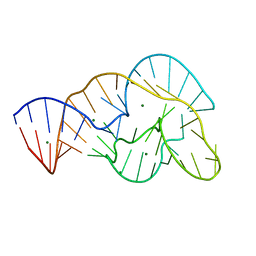 | | Crystal Structure of Twister Ribozyme | | Descriptor: | MAGNESIUM ION, RNA (52-MER) | | Authors: | Liu, Y, Wilson, T.J, McPhee, S.A, Lilley, D.M.J. | | Deposit date: | 2014-01-21 | | Release date: | 2014-07-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure and mechanistic investigation of the twister ribozyme.
Nat.Chem.Biol., 10, 2014
|
|
3VHR
 
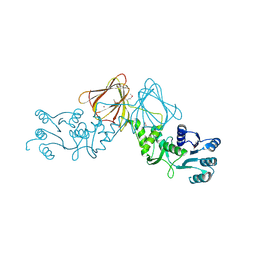 | | Crystal Structure of capsular polysaccharide assembling protein CapF from Staphylococcus aureus in space group C2221 | | Descriptor: | Capsular polysaccharide synthesis enzyme Cap5F, FORMIC ACID, ZINC ION | | Authors: | Miyafusa, T, Yoshikazu, T, Caaveiro, J.M.M, Tsumoto, K. | | Deposit date: | 2011-09-01 | | Release date: | 2012-02-15 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the enzyme CapF of Staphylococcus aureus reveals a unique architecture composed of two functional domains.
Biochem.J., 443, 2012
|
|
3VOL
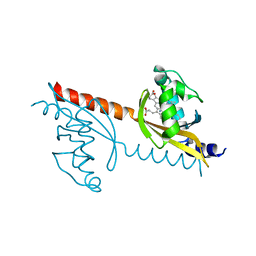 
 | | X-ray Crystal Structure of PAS-HAMP Aer2 in the CN-bound Form | | Descriptor: | Aerotaxis transducer Aer2, CYANIDE ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Sawai, H, Sugimoto, H, Shiro, Y, Aono, S. | | Deposit date: | 2012-01-27 | | Release date: | 2012-05-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | Structural basis for oxygen sensing and signal transduction of the heme-based sensor protein Aer2 from Pseudomonas aeruginosa
Chem.Commun.(Camb.), 48, 2012
|
|
