2FLD
 
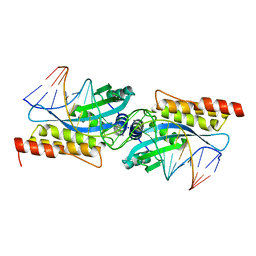 | | I-MsoI Re-Designed for Altered DNA Cleavage Specificity | | Descriptor: | 5'-D(*CP*GP*GP*AP*AP*CP*GP*GP*TP*CP*TP*CP*AP*CP*GP*AP*CP*CP*TP*TP*CP*TP*GP*C)-3', 5'-D(*GP*CP*AP*GP*AP*AP*GP*GP*TP*CP*GP*TP*GP*AP*GP*AP*CP*CP*GP*TP*TP*CP*CP*G)-3', CALCIUM ION, ... | | Authors: | Ashworth, J, Duarte, C.M, Havranek, J.J, Sussman, D, Monnat, R.J, Stoddard, B.L, Baker, D. | | Deposit date: | 2006-01-05 | | Release date: | 2006-06-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Computational redesign of endonuclease DNA binding and cleavage specificity.
Nature, 441, 2006
|
|
4N83
 
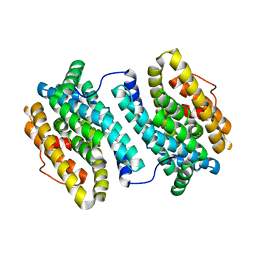 | |
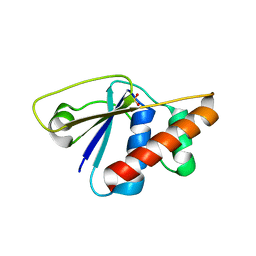
3RH0
 
 | |
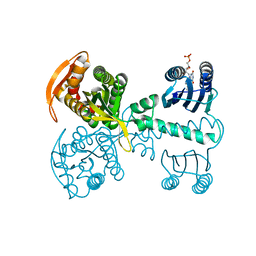
5X4T
 
 | |
3RJS
 
 | |
3ADI
 
 | |
5X4V
 
 | | Roseoflavin substituted OaPAC | | Descriptor: | 1-deoxy-1-[8-(dimethylamino)-7-methyl-2,4-dioxo-3,4-dihydrobenzo[g]pteridin-10(2H)-yl]-D-ribitol, Photoactivated adenylyl cyclase | | Authors: | Ohki, M, Park, S.-Y. | | Deposit date: | 2017-02-14 | | Release date: | 2017-07-26 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular mechanism of photoactivation of a light-regulated adenylate cyclase.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
3RNM
 
 | | The crystal structure of the subunit binding of human dihydrolipoamide transacylase (E2b) bound to human dihydrolipoamide dehydrogenase (E3) | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, BETA-MERCAPTOETHANOL, Dihydrolipoyl dehydrogenase, ... | | Authors: | Brautigam, C.A, Wynn, R.M, Chuang, J.C, Young, B.B, Chuang, D.T. | | Deposit date: | 2011-04-22 | | Release date: | 2011-05-04 | | Last modified: | 2011-07-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural and Thermodynamic Basis for Weak Interactions between Dihydrolipoamide Dehydrogenase and Subunit-binding Domain of the Branched-chain {alpha}-Ketoacid Dehydrogenase Complex.
J.Biol.Chem., 286, 2011
|
|
3ADL
 
 | |
3R3J
 
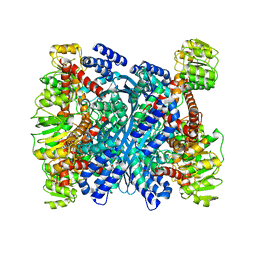 | | Kinetic and structural characterization of Plasmodium falciparum glutamate dehydrogenase 2 | | Descriptor: | Glutamate dehydrogenase | | Authors: | Zocher, K, Fritz-Wolf, K, Kehr, S, Rahlfs, S, Becker, K. | | Deposit date: | 2011-03-16 | | Release date: | 2012-03-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Biochemical and structural characterization of Plasmodium falciparum glutamate dehydrogenase 2.
Mol.Biochem.Parasitol., 183, 2012
|
|
2G9H
 
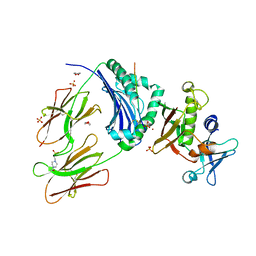 | | Crystal Structure of Staphylococcal Enterotoxin I (SEI) in Complex with a Human MHC class II Molecule | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, HLA class II histocompatibility antigen, ... | | Authors: | Fernandez, M.M, Guan, R, Malchiodi, E.L, Mariuzza, R.A. | | Deposit date: | 2006-03-06 | | Release date: | 2006-07-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of staphylococcal enterotoxin I (SEI) in complex with a human major histocompatibility complex class II molecule.
J.Biol.Chem., 281, 2006
|
|
4NWZ
 
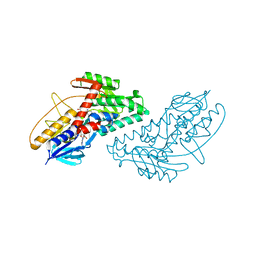 | | Structure of bacterial type II NADH dehydrogenase from Caldalkalibacillus thermarum at 2.5A resolution | | Descriptor: | FAD-dependent pyridine nucleotide-disulfide oxidoreductase, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Nakatani, Y, Heikal, A, Lott, J.S, Sazanov, L.A, Baker, E.N, Cook, G.M. | | Deposit date: | 2013-12-07 | | Release date: | 2014-02-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the bacterial type II NADH dehydrogenase: a monotopic membrane protein with an essential role in energy generation.
Mol.Microbiol., 91, 2014
|
|
3S34
 
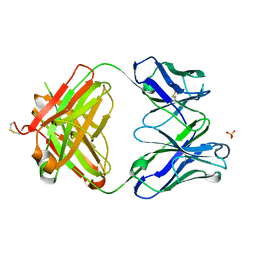 | | Structure of the 1121B Fab fragment | | Descriptor: | 1121B Fab heavy chain, 1121B Fab light chain, PHOSPHATE ION | | Authors: | Franklin, M.C. | | Deposit date: | 2011-05-17 | | Release date: | 2011-08-24 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Structural Basis for the Function of Two Anti-VEGF Receptor 2 Antibodies.
Structure, 19, 2011
|
|
3CMW
 
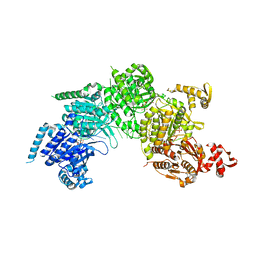 | |
3CMX
 
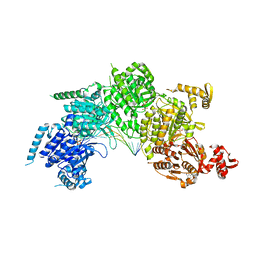 | | Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA (5'-D(*DAP*DAP*DAP*DAP*DAP*DAP*DAP*DAP*DAP*DAP*DAP*DA)-3'), DNA (5'-D(*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DTP*DT)-3'), ... | | Authors: | Pavletich, N.P. | | Deposit date: | 2008-03-24 | | Release date: | 2008-05-20 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures.
Nature, 453, 2008
|
|
3H1T
 
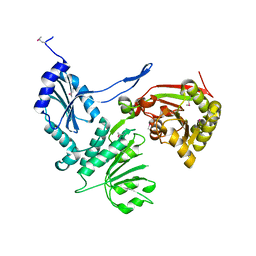 | |
3CQF
 
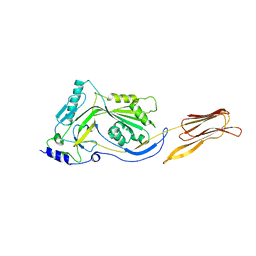 | | Crystal structure of anthrolysin O (ALO) | | Descriptor: | Thiol-activated cytolysin | | Authors: | Bourdeau, R.W, Malito, E, Tang, W.J. | | Deposit date: | 2008-04-02 | | Release date: | 2009-03-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Cellular Functions and X-ray Structure of Anthrolysin O, a Cholesterol-dependent Cytolysin Secreted by Bacillus anthracis
J.Biol.Chem., 284, 2009
|
|
2GBA
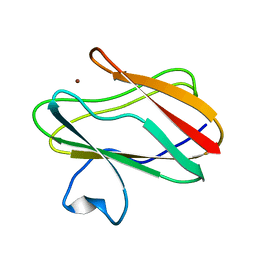 
 | | Reduced Cu(I) form at pH 4 of P52G mutant of amicyanin | | Descriptor: | COPPER (I) ION, amicyanin | | Authors: | Ma, J.K, Carrell, C.J, Mathews, F.S, Davidson, V.L. | | Deposit date: | 2006-03-10 | | Release date: | 2006-08-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (0.92 Å) | | Cite: | Site-Directed Mutagenesis of Proline 52 To Glycine in Amicyanin Converts a True Electron Transfer Reaction into One that Is Conformationally Gated.
Biochemistry, 45, 2006
|
|
3D3V
 
 | | The complex between TCR A6 and human Class I MHC HLA-A2 with the modified HTLV-1 TAX (Y5(3,4-difluoroPhenylalanine)) peptide | | Descriptor: | A6 TCR alpha chain, A6 TCR beta chain, Beta-2-microglobulin, ... | | Authors: | Borbulevych, O.Y, Clemens, J.R, Baker, B.M. | | Deposit date: | 2008-05-12 | | Release date: | 2009-06-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Fluorine substitutions in an antigenic peptide selectively modulate T-cell receptor binding in a minimally perturbing manner.
Biochem.J., 423, 2009
|
|
2QJH
 
 | |
5YX2
 
 | |
2QMC
 
 | |
3HG1
 
 | | Germline-governed recognition of a cancer epitope by an immunodominant human T cell receptor | | Descriptor: | Beta-2-microglobulin, CANCER/MART-1, GLYCEROL, ... | | Authors: | Cole, D.K, Yuan, F, Rizkallah, P.J, Miles, J.J, Gostick, E, Price, D.A, Gao, G.F, Jakobsen, B.K, Sewell, A.K. | | Deposit date: | 2009-05-13 | | Release date: | 2009-07-28 | | Last modified: | 2014-04-09 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Germ line-governed recognition of a cancer epitope by an immunodominant human T-cell receptor.
J.Biol.Chem., 284, 2009
|
|
2ERP
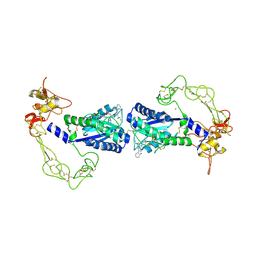 
 | | Crystal structure of vascular apoptosis-inducing protein-1(inhibitor-bound form) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-(N-HYDROXYCARBOXAMIDO)-2-ISOBUTYLPROPANOYL-TRP-METHYLAMIDE, CALCIUM ION, ... | | Authors: | Takeda, S, Igarashi, T, Araki, S. | | Deposit date: | 2005-10-25 | | Release date: | 2006-06-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Crystal structures of VAP1 reveal ADAMs' MDC domain architecture and its unique C-shaped scaffold
Embo J., 25, 2006
|
|
3TQ0
 
 | | Crystal structure of Leishmania major dihydroorotate dehydrogenase in complex with fumarate | | Descriptor: | Dihydroorotate dehydrogenase, FLAVIN MONONUCLEOTIDE, FUMARIC ACID, ... | | Authors: | Cordeiro, A.T, Feliciano, P.R, Nonato, M.C. | | Deposit date: | 2011-09-08 | | Release date: | 2012-05-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of dihydroorotate dehydrogenase from Leishmania major.
Biochimie, 94, 2012
|
|
