3QBN
 
 | |
3T13
 
 | | Crystal structure of Staphylococcal nuclease variant Delta+PHS A69G at cryogenic temperature | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, CALCIUM ION, PHOSPHATE ION, ... | | Authors: | Doctrow, B.M, Schlessman, J.L, Garcia-Moreno E, B, Heroux, A. | | Deposit date: | 2011-07-21 | | Release date: | 2011-08-03 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Staphylococcal nuclease variant Delta+PHS A69G at cryogenic temperature
To be Published
|
|
3T40
 
 | | Crystal Structure of Mycobacterium tuberculosis Indole Glycerol Phosphate Synthase (IGPS) complex with N-2-Carboxyphenyl Glycine (CPG) | | Descriptor: | 2-[(carboxymethyl)amino]benzoic acid, Indole-3-glycerol phosphate synthase, SULFATE ION | | Authors: | Reddy, M.C.M, Bruning, J.B, Thurman, C, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2011-07-25 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.752 Å) | | Cite: | Structural Insights and Inhibition of Mycobacterium tuberculosis Indole Glycerol Phosphate Synthase (IGPS): an Essential Enzyme for Tryptophan Biosynthesis
To be Published
|
|
3T45
 
 | | Crystal structure of bacteriorhodopsin mutant A215T, a phototaxis signaling mutant at 3.0 A resolution | | Descriptor: | 1-[2,6,10.14-TETRAMETHYL-HEXADECAN-16-YL]-2-[2,10,14-TRIMETHYLHEXADECAN-16-YL]GLYCEROL, Bacteriorhodopsin (GROUND STATE), RETINAL | | Authors: | Ozorowski, G, Luecke, H. | | Deposit date: | 2011-07-25 | | Release date: | 2011-12-21 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | A transporter converted into a sensor, a phototaxis signaling mutant of bacteriorhodopsin at 3.0 angstrom.
J.Mol.Biol., 415, 2012
|
|
3T4A
 
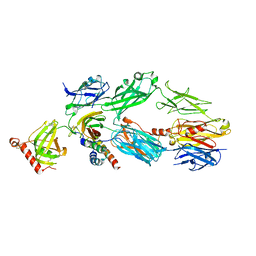 | | Structure of a truncated form of Staphylococcal Complement Inhibitor B bound to human C3c at 3.4 Angstrom resolution | | Descriptor: | Complement C3 beta chain, Complement C3c alpha' chain fragment 1, Complement C3c alpha' chain fragment 2, ... | | Authors: | Garcia, B.L, Geisbrecht, B.V, Summers, B.J. | | Deposit date: | 2011-07-25 | | Release date: | 2011-11-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Diversity in the C3b Convertase Contact Residues and Tertiary Structures of the Staphylococcal Complement Inhibitor (SCIN) Protein Family.
J.Biol.Chem., 287, 2012
|
|
3T7J
 
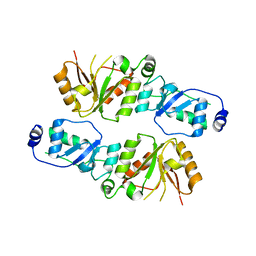 | | Crystal structure of Rtt107p (residues 820-1070) | | Descriptor: | Regulator of Ty1 transposition protein 107 | | Authors: | Li, X, Li, F, Wu, J, Shi, Y. | | Deposit date: | 2011-07-30 | | Release date: | 2012-02-15 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.042 Å) | | Cite: | Structure of C-terminal Tandem BRCT Repeats of Rtt107 Protein Reveals Critical Role in Interaction with Phosphorylated Histone H2A during DNA Damage Repair
J.Biol.Chem., 287, 2012
|
|
3T8I
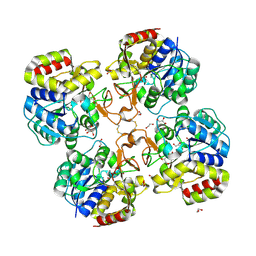 
 | | Structural analysis of thermostable S. solfataricus purine-specific nucleoside hydrolase | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Minici, C, Cacciapuoti, G, De Leo, E, Porcelli, M, Degano, M. | | Deposit date: | 2011-08-01 | | Release date: | 2012-05-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | New Determinants in the Catalytic Mechanism of Nucleoside Hydrolases from the Structures of Two Isozymes from Sulfolobus solfataricus.
Biochemistry, 51, 2012
|
|
3T8N
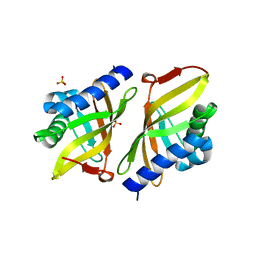 
 | | Crystal structure of ketosteroid isomerase Y16AD103A from Pseudomonas putida | | Descriptor: | SULFATE ION, Steroid Delta-isomerase, {[-(BIS-CARBOXYMETHYL-AMINO)-ETHYL]-CARBOXYMETHYL-AMINO}-ACETIC ACID | | Authors: | Gonzalez, A, Tsai, Y, Schwans, J, Sunden, F, Herschlag, D. | | Deposit date: | 2011-08-01 | | Release date: | 2011-11-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Evaluating the catalytic contribution from the oxyanion hole in ketosteroid isomerase.
J.Am.Chem.Soc., 133, 2011
|
|
3TBH
 
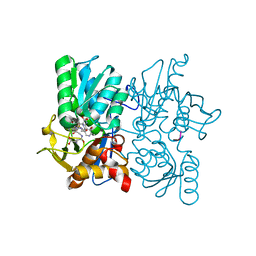 | |
3TCV
 
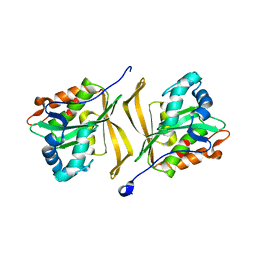 | |
3TDE
 
 | |
3TE8
 
 | |
3SQB
 | | Structure of the major type 1 pilus subunit FimA bound to the FimC chaperone | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Chaperone protein fimC, ... | | Authors: | Scharer, M.A, Eidam, O, Grutter, M.G, Glockshuber, R, Capitani, G. | | Deposit date: | 2011-07-05 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Quality control of disulfide bond formation in pilus subunits by the chaperone FimC.
Nat.Chem.Biol., 8, 2012
|
|
3STK
 | |
3SW5
 
 | |
3SWX
 
 | |
3TE2
 
 | | Crystal structure of HSC K16S | | Descriptor: | TETRAETHYLENE GLYCOL, formate/nitrite transporter, octyl beta-D-glucopyranoside | | Authors: | Czyzewski, B.K, Wang, D.-N. | | Deposit date: | 2011-08-11 | | Release date: | 2012-03-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Identification and characterization of a bacterial hydrosulphide ion channel.
Nature, 483, 2012
|
|
3SXH
 
 | | Crystal structure of Staphylococcal nuclease variant Delta+PHS I92AL103A at cryogenic temperature | | Descriptor: | CALCIUM ION, THYMIDINE-3',5'-DIPHOSPHATE, Thermonuclease | | Authors: | Caro, J.A, Clark, I, Schlessman, J.L, Heroux, A, Garcia-Moreno E, B. | | Deposit date: | 2011-07-14 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure of Staphylococcal nuclease variant Delta+PHS I92AL103A at cryogenic temperature
To be Published
|
|
3T0L
 
 | |
3T28
 | | TMAO-grown trypsin (bovine)-previously unreported tetragonal crystal form | | Descriptor: | BENZAMIDINE, CALCIUM ION, Cationic trypsin, ... | | Authors: | Cahn, J, Venkat, M, Saxby, S, Juers, D. | | Deposit date: | 2011-07-22 | | Release date: | 2011-12-28 | | Last modified: | 2018-03-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The use of trimethylamine N-oxide as a primary precipitating agent and related methylamine osmolytes as cryoprotective agents for macromolecular crystallography.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3SOE
 
 | | Crystal Structure of the 3rd PDZ domain of the human Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 3 (MAGI3) | | Descriptor: | 1,2-ETHANEDIOL, Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 3 | | Authors: | Ivarsson, Y, Filippakopoulos, P, Picaud, S, Vollmar, M, von Delft, F, Arrowsmith, C.H, Weigelt, J, Edwards, A, Bountra, C, Knapp, S, Zimmermann, P, Structural Genomics Consortium (SGC) | | Deposit date: | 2011-06-30 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structure of the 3rd PDZ domain of the human Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 3 (MAGI3)
To be Published
|
|
3SOI
 
 | | Crystallographic structure of Bacillus licheniformis beta-lactamase W210F/W229F/W251F at 1.73 angstrom resolution | | Descriptor: | Beta-lactamase, CITRIC ACID | | Authors: | Acierno, J.P, Capaldi, S, Risso, V.A, Monaco, H.L, Ermacora, M.R. | | Deposit date: | 2011-06-30 | | Release date: | 2011-12-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.729 Å) | | Cite: | X-ray evidence of a native state with increased compactness populated by tryptophan-less B. licheniformis beta-lactamase.
Protein Sci., 21, 2012
|
|
3SPX
 
 | | Crystal structure of O-Acetyl Serine Sulfhydrylase from Leishmania donovani | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, O-acetyl serine sulfhydrylase, ... | | Authors: | Raj, I, Gourinath, S. | | Deposit date: | 2011-07-04 | | Release date: | 2012-07-04 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | The narrow active-site cleft of O-acetylserine sulfhydrylase from Leishmania donovani allows complex formation with serine acetyltransferases with a range of C-terminal sequences
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3SQD
 
 | |
3SQF
 
 | | Crystal structure of monomeric M-PMV retroviral protease | | Descriptor: | Protease | | Authors: | Jaskolski, M, Kazmierczyk, M, Gilski, M, Krzywda, S, Pichova, I, Zabranska, H, Khatib, F, DiMaio, F, Cooper, S, Thompson, J, Popovic, Z, Baker, D, Group, Foldit Contenders | | Deposit date: | 2011-07-05 | | Release date: | 2011-09-21 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6324 Å) | | Cite: | Crystal structure of a monomeric retroviral protease solved by protein folding game players.
Nat.Struct.Mol.Biol., 18, 2011
|
|
