3KNV
 
 | |
2QU5
 
 | | Crystal structure of the VEGFR2 kinase domain in complex with a benzimidazole inhibitor | | Descriptor: | 4-[[2-[[4-chloro-3-(trifluoromethyl)phenyl]amino]-3H-benzimidazol-5-yl]oxy]-N-methyl-pyridine-2-carboxamide, Vascular endothelial growth factor receptor 2 | | Authors: | Whittington, D.A, Kim, J.L, Long, A.M, Rose, P, Gu, Y, Zhao, H. | | Deposit date: | 2007-08-03 | | Release date: | 2007-09-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Design, Synthesis, and Evaluation of Orally Active Benzimidazoles and Benzoxazoles as Vascular Endothelial Growth Factor-2 Receptor Tyrosine Kinase Inhibitors.
J.Med.Chem., 50, 2007
|
|
3KOB
 
 | |
3KOR
 
 | | Crystal structure of a putative Trp repressor from Staphylococcus aureus | | Descriptor: | Possible Trp repressor | | Authors: | Lam, R, Vodsedalek, J, Lam, K, Romanov, V, Battaile, K.P, Beletskaya, I, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2009-11-13 | | Release date: | 2010-11-17 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of a putative Trp repressor from Staphylococcus aureus
To be Published
|
|
3KI5
 
 | |
3KIE
 
 | |
3KPY
 
 | | Crystal Structure of hPNMT in Complex AdoHcy and 6-Chlorooxindole | | Descriptor: | 6-chloro-1,3-dihydro-2H-indol-2-one, Phenylethanolamine N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Drinkwater, N, Martin, J.L. | | Deposit date: | 2009-11-17 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Fragment-based screening by X-ray crystallography, MS and isothermal titration calorimetry to identify PNMT (phenylethanolamine N-methyltransferase) inhibitors.
Biochem.J., 431, 2010
|
|
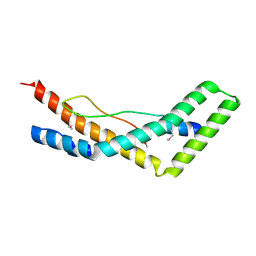
5JW9
 | |
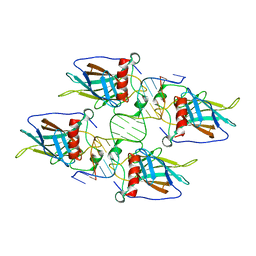
3KMD
 | |
6USZ
 
 | | Identification of the Clinical Development Candidate MRTX849, a Covalent KRASG12C Inhibitor for the Treatment of Cancer | | Descriptor: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Vigers, G.P, Smith, D.J. | | Deposit date: | 2019-10-28 | | Release date: | 2020-04-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Identification of the Clinical Development CandidateMRTX849, a Covalent KRASG12CInhibitor for the Treatment of Cancer.
J.Med.Chem., 63, 2020
|
|
3KMY
 
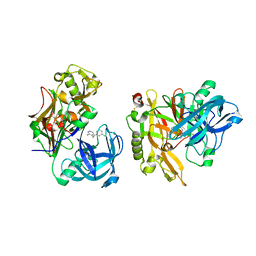 | | Structure of BACE bound to SCH12472 | | Descriptor: | 3-[2-(3-chlorophenyl)ethyl]pyridin-2-amine, Beta-secretase 1 | | Authors: | Strickland, C, Wang, Y. | | Deposit date: | 2009-11-11 | | Release date: | 2010-01-19 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Application of Fragment-Based NMR Screening, X-ray Crystallography, Structure-Based Design, and Focused Chemical Library Design to Identify Novel muM Leads for the Development of nM BACE-1 (beta-Site APP Cleaving Enzyme 1) Inhibitors.
J.Med.Chem., 53, 2010
|
|
5TZX
 
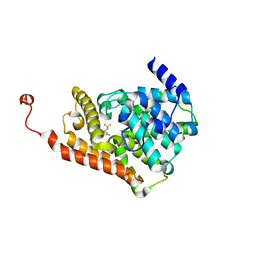 | | CRYSTAL STRUCTURE OF HUMAN PHOSPHODIESTERASE 2A IN COMPLEX WITH 1-[(3-chloro-4-fluorophenyl)carbonyl]-3,3-difluoro-5-{5-methyl-[1,2,4]triazolo[1,5-a]pyrimidin-7-yl}piperidine | | Descriptor: | (3-chloro-4-fluorophenyl)[(5S)-3,3-difluoro-5-(5-methyl[1,2,4]triazolo[1,5-a]pyrimidin-7-yl)piperidin-1-yl]methanone, MAGNESIUM ION, ZINC ION, ... | | Authors: | Xu, R, Aertgeerts, K. | | Deposit date: | 2016-11-22 | | Release date: | 2017-02-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Design and Synthesis of Novel and Selective Phosphodiesterase 2 (PDE2a) Inhibitors for the Treatment of Memory Disorders.
J. Med. Chem., 60, 2017
|
|
3KE1
 | |
3KNU
 
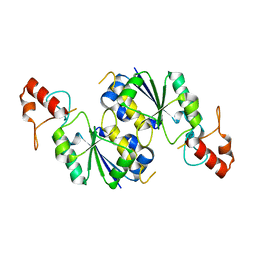 | |
2QJR
 
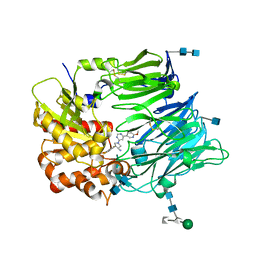 | | dipepdyl peptidase IV in complex with inhibitor PZF | | Descriptor: | (3R,4S)-1-[6-(6-METHOXYPYRIDIN-3-YL)PYRIMIDIN-4-YL]-4-(2,4,5-TRIFLUOROPHENYL)PYRROLIDIN-3-AMINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Shenping, L. | | Deposit date: | 2007-07-09 | | Release date: | 2008-07-29 | | Last modified: | 2022-04-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | (3R,4S)-4-(2,4,5-Trifluorophenyl)-pyrrolidin-3-ylamine inhibitors of dipeptidyl peptidase IV: synthesis, in vitro, in vivo, and X-ray crystallographic characterization.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
3KO8
 
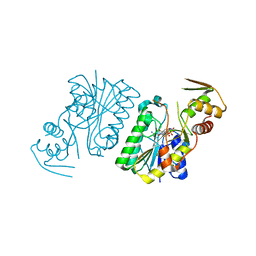 | | Crystal Structure of UDP-galactose 4-epimerase | | Descriptor: | NAD-dependent epimerase/dehydratase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, PHOSPHATE ION | | Authors: | Sakuraba, H, Kawai, T, Yoneda, K, Ohshima, T. | | Deposit date: | 2009-11-13 | | Release date: | 2010-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of UDP-galactose 4-epimerase from the hyperthermophilic archaeon Pyrobaculum calidifontis
Arch.Biochem.Biophys., 512, 2011
|
|
5K0J
 
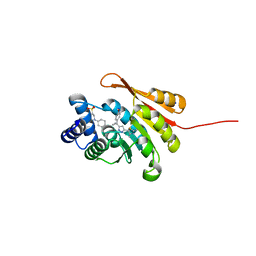 | | Crystal Structure of COMT in complex with 5-[5-[1-(4-methoxyphenyl)cyclopropyl]-1H-pyrazol-3-yl]-2,4-dimethyl-1,3-thiazole | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, 5-{3-[1-(4-methoxyphenyl)cyclopropyl]-1H-pyrazol-5-yl}-2,4-dimethyl-1,3-thiazole, Catechol O-methyltransferase, ... | | Authors: | Ehler, A, Rodriguez-Sarmiento, R.M, Rudolph, M.G. | | Deposit date: | 2016-05-17 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Design of Potent and Druglike Nonphenolic Inhibitors for Catechol O-Methyltransferase Derived from a Fragment Screening Approach Targeting the S-Adenosyl-l-methionine Pocket.
J. Med. Chem., 59, 2016
|
|
5K03
 | | Crystal Structure of COMT in complex with 2,6-dimethyl-3-(1H-pyrazol-3-yl)imidazo[1,2-a]pyridine | | Descriptor: | 2,6-dimethyl-3-(1H-pyrazol-5-yl)imidazo[1,2-a]pyridine, Catechol O-methyltransferase, POTASSIUM ION | | Authors: | Ehler, A, Rodriguez-Sarmiento, R.M, Rudolph, M.G. | | Deposit date: | 2016-05-17 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Design of Potent and Druglike Nonphenolic Inhibitors for Catechol O-Methyltransferase Derived from a Fragment Screening Approach Targeting the S-Adenosyl-l-methionine Pocket.
J. Med. Chem., 59, 2016
|
|
3KPH
 | |
3KF2
 
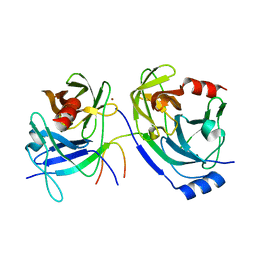 | | The HCV NS3/NS4A protease apo structure | | Descriptor: | 19-mer peptide from Genome polyprotein, Polyprotein, ZINC ION | | Authors: | Lindberg, J.D, Nystrom, S, Cummings, M.D. | | Deposit date: | 2009-10-27 | | Release date: | 2010-03-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Induced-Fit Binding of the Macrocyclic Noncovalent Inhibitor TMC435 to its HCV NS3/NS4A Protease Target
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
3KGS
 
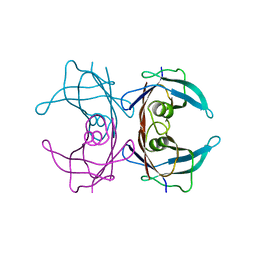 | | V30M mutant human transthyretin (TTR) (apoV30M) pH 7.5 | | Descriptor: | Transthyretin | | Authors: | Trivella, D.B, Polikarpov, I. | | Deposit date: | 2009-10-29 | | Release date: | 2010-03-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Conformational differences between the wild type and V30M mutant transthyretin modulate its binding to genistein: implications to tetramer stability and ligand-binding.
J.Struct.Biol., 170, 2010
|
|
3KPQ
 
 | | Crystal Structure of HLA B*4405 in complex with EEYLKAWTF, a mimotope | | Descriptor: | Beta-2-microglobulin, EEYLKAWTF, mimotope peptide, ... | | Authors: | Macdonald, W.A, Chen, Z, Gras, S, Archbold, J.K, Tynan, F.E, Clements, C.S, Bharadwaj, M, Kjer-Nielsen, L, Saunders, P.M, Wilce, M.C, Crawford, F, Stadinsky, B, Jackson, D, Brooks, A.G, Purcell, A.W, Kappler, J.W, Burrows, S.R, Rossjohn, J, McCluskey, J. | | Deposit date: | 2009-11-16 | | Release date: | 2009-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | T cell allorecognition via molecular mimicry.
Immunity, 31, 2009
|
|
3KQ5
 
 | | Crystal structure of an uncharacterized protein from Coxiella burnetii | | Descriptor: | Hypothetical cytosolic protein | | Authors: | Bonanno, J.B, Freeman, M, Bain, K.T, Chang, S, Ozyurt, S, Wasserman, S, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-11-17 | | Release date: | 2009-12-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of an uncharacterized protein from Coxiella burnetii
To be Published
|
|
3KGY
 
 | |
5K0N
 
 | | Crystal Structure of COMT in complex with 4-[5-[1-(4-methoxyphenyl)cyclopropyl]-1H-pyrazol-3-yl]-1,3-dimethylpyrazole | | Descriptor: | 1,2-ETHANEDIOL, 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, 5-[1-(4-methoxyphenyl)cyclopropyl]-1',3'-dimethyl-1'H,2H-3,4'-bipyrazole, ... | | Authors: | Ehler, A, Rodriguez-Sarmiento, R.M. | | Deposit date: | 2016-05-17 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Design of Potent and Druglike Nonphenolic Inhibitors for Catechol O-Methyltransferase Derived from a Fragment Screening Approach Targeting the S-Adenosyl-l-methionine Pocket.
J. Med. Chem., 59, 2016
|
|
